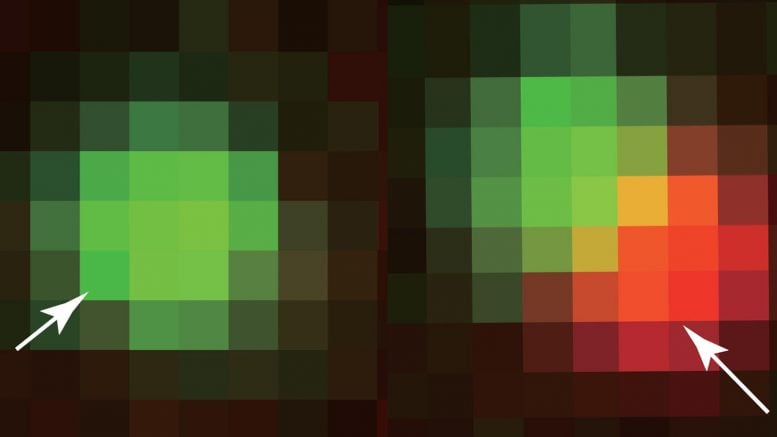

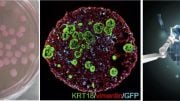
Princeton researchers have developed a framework to engineer the protein droplets that organize crucial functions inside a cell. Key to their findings was the theory of bubble formation, adapted from classical materials science. In the above, an engineered protein fragment (green) “seeds” the formation of a protein droplet (red), the basis for higher-order organelles such as the nucleolus. The new work marks a seismic shift in scientists’ ability to manipulate cells. Credit: Image courtesy of the researchers
Theory of Bubbles Lifts Cell Biology Into a New, More Quantitative Era
A study published on September 22, 2021, in the journal Nature details how an established physics theory governing bubble and droplet formation led to a new understanding of the principles organizing the contents of living cells. The work marks a seismic shift in researchers’ ability both to understand and control the complex soft materials within our cells.
“This approach is common in materials science, but we’ve adapted it to do something unprecedented in cells,” said principal investigator Clifford Brangwynne, the June K. Wu ’92 Professor in Engineering and director of the Princeton Bioengineering Initiative.
The current work follows Brangwynne’s discovery more than a decade ago that cellular proteins organize into liquid structures inside the cell. That insight gave rise to a new field of study examining how parts of cells form much like oil drops coalescing in water. Scientists have puzzled ever since over the exact details of how those structures assemble. But it’s a hard thing to measure the squishy dynamics of individual molecules inside a cell, where mysterious, overlapping processes roil chaotically as minute structures form and dissolve a thousand times per second.
Postdoctoral researcher Shunsuke Shimobayashi had studied soft matter physics at the Kyoto University and wondered whether his background working on organic compounds called lipids might illuminate anything interesting about the problem. If protein molecules condense out of their surroundings the way oil separates from water, maybe the math that described the first steps in that process, called nucleation, would prove useful in proteins as well.
Shimobayashi turned to classical nucleation theory, a pillar of materials science. Its equations had powered some of the most profound technological transformations of the 20th century, from the climate models that first revealed global warming to the fertilizers that helped lift billions of people out of starvation.
He was also keenly aware of a critical distinction: those equations describe simple, inanimate systems, but the inside of a cell is in turmoil. “It’s a much more complex material environment for biomolecules,” Shimobayashi said. To tackle this complexity, the team expanded to include theorists Pierre Ronceray, a former fellow at the Princeton Center for Theoretical Science, and Mikko Haataja, professor of mechanical and aerospace engineering whose prior theoretical and computational work with the Brangwynne lab had led to key insights in related studies. The researchers stripped the theory down to its two most important parameters, adapting it to try to understand how the process might work in cells. Then to test the theory, Shimobayashi turned to an advanced protein tool developed in Brangwynne’s lab in 2018 that provided an ideal, simplified system that mimics how the process occurs naturally in cells. Putting them together, the results came as something of a shock.
When Shimobayashi tried to induce the droplets to seed instantaneously, the system failed. But when he seeded the droplets more slowly, they nucleated at precisely defined locations, in a way that lined up perfectly with his adapted theory. He had predicted how, where, and when the protein droplets formed with what Brangwynne called “remarkable accuracy.”
The team next turned back to the messy complexity of native cell structures, now collaborating with another Brangwynne lab postdoc, David Sanders, an expert on internal cell structures called stress granules. When they accounted for all the processes that act on protein concentrations, they found that the theory worked just as well for stress granules and other condensates. They had quantified the molecule-by-molecule assembly of proteins into the complex liquid structures that regulate life’s most basic routines. Not only do these structures look and act like oil in water, Shimobayashi said, they also form droplets in the same basic nucleation patterns, clustering around minute variations in their environment at rates that can be predicted with the same quantitative precision as other kinds of materials.
With that predictive power comes an accelerated engineering capacity, according to Brangwynne. He believes quantifying biomolecular processes and developing predictive models in the mold of physics will lead to a world in which we no longer watch passively as our loved ones succumb to diseases like Alzheimer’s.
“We first have to understand how it works, with quantitative mathematical frameworks that are the bedrock of society’s engineering marvels. And then we can take the next steps, to manipulate biological systems with greater control,” Brangwynne said. “We need to be able to turn the knobs.”
Reference: “Nucleation landscape of biomolecular condensates” by Shunsuke F. Shimobayashi, Pierre Ronceray, David W. Sanders, Mikko P. Haataja and Clifford P. Brangwynne, 22 September 2021, Nature.
DOI: 10.1038/s41586-021-03905-5
This work, titled “Nucleation landscape of biomolecular condensates,” was supported in part by the Howard Hughes Medical Institute, a Focused Research Team Award from Princeton’s School of Engineering and Applied Science, the National Institutes of Health and the Princeton Center for Complex Materials., Air Force Office of Scientific Research, and the Princeton Center for Complex Materials.







Be the first to comment on "Framework for Engineering Living Cells Inspired by the Math That Describes How Bubbles Form in a Glass of Champagne"