
Researchers have pinpointed genes linked to plant survival in the extreme conditions of the Atacama Desert in Chile.
Evolutionary genomics approach identifies genes that enable plants to live in the Atacama Desert, offering clues for engineering more resilient crops to face climate change.
An international team of researchers has identified genes associated with plant survival in one of the harshest environments on Earth: the Atacama Desert in Chile. Their findings, published in Proceedings of the National Academy of Sciences (PNAS), may help scientists breed resilient crops that can thrive in increasingly drier climates.
“In an era of accelerated climate change, it is critical to uncover the genetic basis to improve crop production and resilience under dry and nutrient-poor conditions,” said Gloria Coruzzi, Carroll & Milton Petrie Professor in the New York University (NYU) Department of Biology and Center for Genomics and Systems Biology, who co-led the study with Rodrigo Gutiérrez.
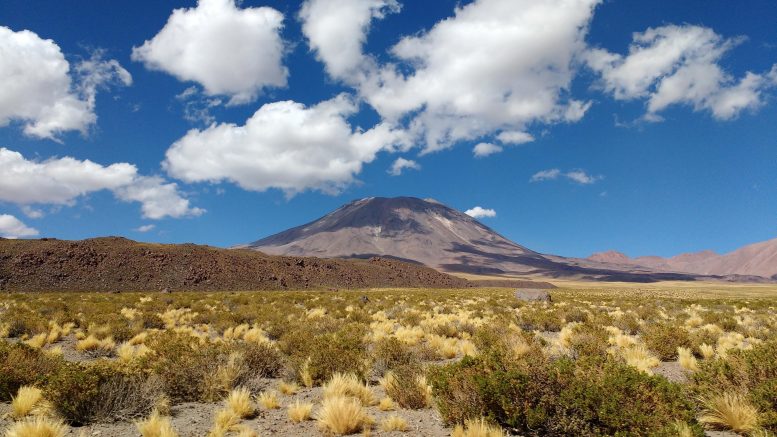

The Chilean research team established an unparalleled “natural laboratory” in the Atacama Desert in northern Chile, one of the driest and harshest environments on Earth. Credit: Melissa Aguilar
The study was an international collaboration among botanists, microbiologists, ecologists, evolutionary and genomic scientists. This unique combination of expertise enabled the team to identify the plants, associated microbes, and genes that enable the Atacama plants to adapt to and flourish in extreme desert conditions, which could ultimately help to enhance crop growth and reduce food insecurity.
“Our study of plants in the Atacama Desert is directly relevant to regions around the world that are becoming increasingly arid, with factors such as drought, extreme temperatures, and salt in water and soil posing a significant threat to global food production,” said Gutiérrez, professor in the Department of Molecular Genetics and Microbiology at Pontificia Universidad Católica de Chile.
Establishing a “natural laboratory” in one of Earth’s driest places
The Atacama Desert in northern Chile, sandwiched between the Pacific Ocean and Andes Mountains, is the driest place on the planet (excluding the poles). Yet dozens of plants grow there, including grasses, annuals, and perennial shrubs. In addition to limited water, plants in the Atacama must cope with high altitude, low availability of nutrients in the soil, and extremely high radiation from sunlight.
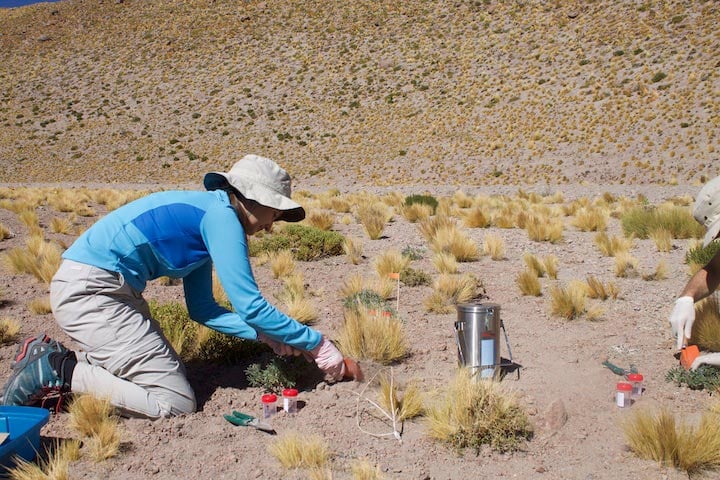

Gabriela Carrasco, an undergraduate researcher at the time, is identifying, labeling, collecting, and freezing plant samples in the Atacama Desert. These samples then traveled 1,000 miles, kept under dry ice to be processed for RNA extractions in Rodrigo Gutiérrez’s lab in Santiago de Chile. The species Carrasco is collecting here are Jarava frigida and Lupinus oreophilus. Credit: Melissa Aguilar
The Chilean research team established an unparalleled “natural laboratory” in the Atacama Desert over a 10-year period, in which they collected and characterized the climate, soil, and plants at 22 sites in different vegetational areas and elevations (every 100 meters of altitude) along the Talabre-Lejía Transect. Measuring a variety of factors, they recorded temperatures that fluctuated more than 50 degrees from day to night, very high radiation levels, soil that was largely sand and lacked nutrients, and minimal rain, with most annual rain falling over a few days.
Using genomics to explore the evolution of resilient plants
The Chilean researchers brought the plant and soil samples—preserved in liquid nitrogen—1,000 miles back to the lab to sequence the genes expressed in the 32 dominant plant species in the Atacama and assess the plant-associated soil microbes based on DNA sequences. They found that some plant species developed growth-promoting bacteria near their roots, an adaptive strategy to optimize the intake of nitrogen—a nutrient critical for plant growth—in the nitrogen-poor soils of the Atacama.
To identify the genes whose protein sequences were adapted in the Atacama species, the researchers at NYU next conducted an analysis using an approach called phylogenomics, which aims to reconstruct evolutionary history using genomic data. In consultation with colleagues at the New York Botanical Garden, they compared the genomes of the 32 Atacama plants with 32 non-adapted but genetically similar “sister” species, as well as several model species.
“The goal was to use this evolutionary tree based on genome sequences to identify the changes in amino acid sequences encoded in the genes that support the evolution of the Atacama plant adaptation to desert conditions,” said Coruzzi.
“This computationally intense genomic analysis involved comparing 1,686,950 protein sequences across more than 70 species. We used the resulting super-matrix of 8,599,764 amino acids for phylogenomic reconstruction of the evolutionary history of the Atacama species,” said Gil Eshel, who conducted this analysis using the High Performance Computing Cluster at NYU.
The study identified 265 candidate genes whose protein sequence changes were selected by evolutionary forces across multiple Atacama species. These adaptive mutations occurred in genes that could underlie plant adaptation to the desert conditions, including genes involved in response to light and photosynthesis, which may enable plants to adapt to the extreme high-light radiation in the Atacama. Similarly, the researchers uncovered genes involved in the regulation of stress response, salt, detoxification, and metal ions, which could be related to the adaptation of these Atacama plants to their stressful, nutrient-poor environment.
What we can learn from this “genetic goldmine”
The majority of scientific knowledge of plant stress responses and tolerance has been generated through traditional lab-based studies using a few model species. While beneficial, such molecular studies likely miss the ecological context in which plants have evolved.
“By studying an ecosystem in its natural environment, we were able to identify adaptive genes and molecular processes among species facing a common harsh environment,” said Viviana Araus of the Pontificia Universidad Católica de Chile in Gutierrez’ lab and a former postdoctoral associate at NYU’s Center for Genomics and Systems Biology.
“Most of the plant species we characterized in this research have not been studied before. As some Atacama plants are closely related to staple crops, including grains, legumes, and potatoes, the candidate genes we identified represent a genetic goldmine to engineer more resilient crops, a necessity given the increased desertification of our planet,” said Gutiérrez.
In addition to Gutiérrez and Araus, their collaborators in Chile included Claudio Latorre of the Pontificia Universidad Católica de Chile and Mauricio González of the Universidad de Chile. Coruzzi and Eshel at NYU worked on the phylogenomic pipeline and analysis with collaborators in the U.S., including Kranthi Varala of Purdue University, Dennis Stevenson of the New York Botanical Garden, Rob DeSalle of the American Museum of Natural History, as well as members of their research teams.
Reference: “Plant ecological genomics at the limits of life in the Atacama Desert” by Gil Eshel, Viviana Araus, Soledad Undurraga, Daniela C. Soto, Carol Moraga, Alejandro Montecinos, Tomás Moyano, Jonathan Maldonado, Francisca P. Díaz, Kranthi Varala, Chase W. Nelson, Orlando Contreras-López, Henrietta Pal-Gabor, Tatiana Kraiser, Gabriela Carrasco-Puga, Ricardo Nilo-Poyanco, Charles M. Zegar, Ariel Orellana, Martín Montecino, Alejandro Maass, Miguel L. Allende, Robert DeSalle, Dennis W. Stevenson, Mauricio González, Claudio Latorre, Gloria M. Coruzzi and Rodrigo A. Gutiérrez, 1 November 2021, Proceedings of the National Academy of Sciences.
DOI: 10.1073/pnas.2101177118
This work was supported by Fondo de Desarrollo de Areas Prioritarias (FONDAP) Center for Genome Regulation (15090007) in Chile, and in the U.S. by the Zegar Family Foundation (A160051), and by a Department of Energy Biological and Environmental Research grant (DE-SC0014377).







cool