
Tracing the Impact of a Long-Lost Relative on Modern Bread Wheat
Genetic detective work has uncovered an obscure ancestor of modern bread wheat, in a finding similar to uncovering a famous long-lost relative through DNA analysis in humans.
In a study which appears in Nature Biotechnology researchers sequenced the DNA from 242 unique accessions of Aegilops tauschii gathered over decades from across its native range — from Turkey to Central Asia.
Population genome analysis led by Dr. Kumar Gaurav from the John Innes Centre revealed the existence of a distinct lineage of Aegilops tauschii restricted to present-day Georgia, in the Caucuses region – some 500 kilometers from the Fertile Crescent where wheat was first cultivated – an area stretching across modern-day Iraq, Syria, Lebanon, Palestine, Israel, Jordan, and Egypt.
First author of the study in Nature Biotechnology, Dr. Kumar Gaurav said, “The discovery of this previously unknown contribution to the bread wheat genome is akin to discovering the introgression of Neanderthal DNA into the out of Africa human genome.
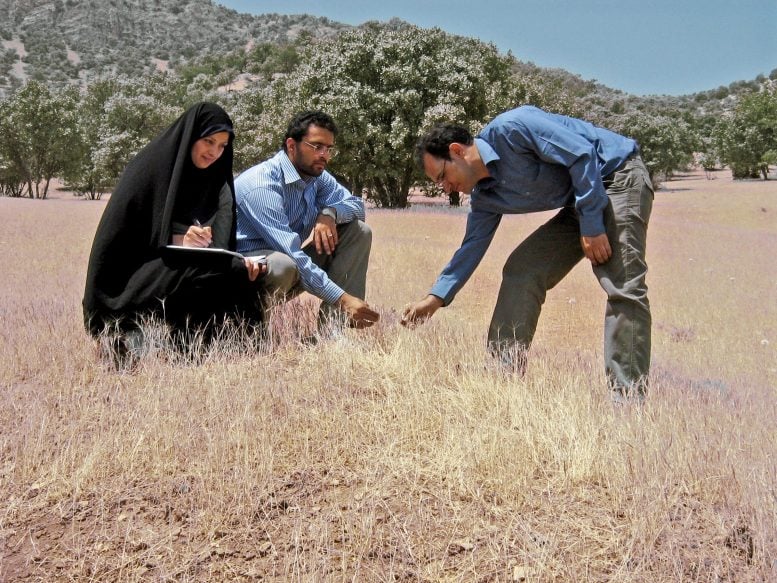

Researchers on a wild wheat relatives foraging trip in the central Zagros mountains in western Iran. Credit: Ali Mehrabi
“It is most likely to have occurred through a hybridization outside the Fertile Crescent. This group of Georgian accessions form a distinct lineage that contributed to the wheat genome by leaving a footprint in the DNA.”
The discovery comes via a major international collaboration to improve crops by exploring useful genetic diversity in Aegilops tauschii, a wild relative of bread wheat. The Open Wild Wheat Consortium brought together 38 research groups and researchers from 17 countries.
Further research by Dr. Jesse Poland’s group at Kansas State University was published in a companion paper in Communications Biology and shows that the ancestral Aegilops tauschii DNA found in modern bread wheat includes the gene that gives superior strength and elasticity to dough.
Dr. Poland said, “We were amazed to discover that this lineage has provided the best-known gene for superior dough quality.”
The researchers speculate that the newly discovered lineage may have been more geographically widespread in the past, and that it may have become separated as a refugium population during the last ice-age.
Reflecting on all that has come together to make this work possible, Dr Brande Wulff, corresponding author of the study, remarked, “Fifty or sixty years ago at a time when we barely understood DNA, my scientific forebears were traversing the Zagros mountains in the middle east and Syria and Iraq. They were collecting seeds, perhaps having an inkling that one day these could be used for improving wheat. Now we are so close to unlocking that potential, and for me that is extraordinarily exciting.”
Deciphering Wheat’s Complex Genome
Modern “hexaploid” wheat, is a complex genetic combination of different grasses with a huge genetic code, split into A, B and D sub-genomes. Hexaploid wheat accounts for 95 percent of all cultivated wheat. Hexaploid means that the DNA contains six sets of chromosomes — three pairs of each.
Through a combination of natural hybridizations and human cultivation, Aegilops tauschii provided the D-genome to modern wheat. The D-genome added the properties for making dough, and enabled bread wheat to flourish in different climates and soils.
The origin of modern hexaploid bread wheat has long been the subject of intense scrutiny with archeological and genetic evidence suggesting that the first wheat was cultivated 10,000 years ago in the Fertile Crescent.
Domestication, while increasing yield and increasing agronomic performance, came at the cost of a pronounced genetic bottleneck eroding genetic diversity for protective traits to be found in Aegilops tauschii such as disease resistance and heat tolerance.
Analysis performed by Dr. Gaurav and the research team revealed that just 25% of the genetic diversity present in Aegilops tauschii made it into hexaploid wheat. To explore this diversity in the wild gene pool, they used a technique called association mapping to discover new candidate genes for disease and pest resistance, yield and environmental resilience.
Dr. Sanu Arora, who had earlier led a study to clone disease resistance genes from Aegilops tauschii said, “Previously we were restricted to exploring a very small subset of the genome for disease resistance, but in the current study, we have generated data and techniques to undertake an unbiased exploration of the species diversity”.
Further experiments demonstrated the transfer of candidate genes for a subset of these traits into wheat using genetic transformation and conventional crossing — facilitated by a library of synthetic wheats — specially bred material which incorporates Aegilops tauschii genomes.
This publicly available library of synthetic wheats captures 70 percent of the diversity present across all three known Aegilops tauschii lineages, enabling researchers to assess traits rapidly in a background of hexaploid wheats.
“Our study provides an end-to-end pipeline for rapid and systematic exploration of the Aegilops tauschii gene pool for improving modern bread wheat,” says Dr. Wulff.
“High molecular weight glutenin gene diversity in Aegilops tauschii demonstrates unique origin of superior wheat quality,” appears in Communications Biology.
Reference: “Population genomic analysis of Aegilops tauschii identifies targets for bread wheat improvement” by Kumar Gaurav, Sanu Arora, Paula Silva, Javier Sánchez-Martín, Richard Horsnell, Liangliang Gao, Gurcharn S. Brar, Victoria Widrig, W. John Raupp, Narinder Singh, Shuangye Wu, Sandip M. Kale, Catherine Chinoy, Paul Nicholson, Jesús Quiroz-Chávez, James Simmonds, Sadiye Hayta, Mark A. Smedley, Wendy Harwood, Suzannah Pearce, David Gilbert, Ngonidzashe Kangara, Catherine Gardener, Macarena Forner-Martínez, Jiaqian Liu, Guotai Yu, Scott A. Boden, Attilio Pascucci, Sreya Ghosh, Amber N. Hafeez, Tom O’Hara, Joshua Waites, Jitender Cheema, Burkhard Steuernagel, Mehran Patpour, Annemarie Fejer Justesen, Shuyu Liu, Jackie C. Rudd, Raz Avni, Amir Sharon, Barbara Steiner, Rizky Pasthika Kirana, Hermann Buerstmayr, Ali A. Mehrabi, Firuza Y. Nasyrova, Noam Chayut, Oadi Matny, Brian J. Steffenson, Nitika Sandhu, Parveen Chhuneja, Evans Lagudah, Ahmed F. Elkot, Simon Tyrrell, Xingdong Bian, Robert P. Davey, Martin Simonsen, Leif Schauser, Vijay K. Tiwari, H. Randy Kutcher, Pierre Hucl, Aili Li, Deng-Cai Liu, Long Mao, Steven Xu, Gina Brown-Guedira, Justin Faris, Jan Dvorak, Ming-Cheng Luo, Ksenia Krasileva, Thomas Lux, Susanne Artmeier, Klaus F. X. Mayer, Cristobal Uauy, Martin Mascher, Alison R. Bentley, Beat Keller, Jesse Poland and Brande B. H. Wulff, 1 November 2021, Nature Biotechnology.
DOI: 10.1038/s41587-021-01058-4







Very Interesting research.
Some thoughts for consideration in feeding the worlds expected population of 10 billion with grains.
Wheat is a popular dietary choice. As is rice! These two variety of Grass constitute the staple diet of human beings today.
For other four legged creatures green grass with more cellulose and hay appear to be popular feeds. Grazing by Sheep which gives mother earth a really close shave needs to be controlled as the land it leaves behind is not able to grow fresh green grass on the grazed land. 😁
Probably the same with camels (single humped or double humped). Not sure.
Other four legged creatures permit recycling of Grazing pastures. That was an aside.
I would strongly recommend lawn mowers which don’t give a clean shave and instead of these four legged milk and milk products gift of mother nature being treated with utter disrespect and cruelty for their meat, they should be treated as the sustainers of life.
The evolution of modern day wheat grown and used in many drier parts of the 🌎Globe is good detective work and it’s properties deserves a 👍👍 thumbs up.
Now coming to the crux of the article. ⬇️⬇️⬇️
1. Figure out how to include the daily nutrients and minerals required by humans so it can be made an integral part of the Wheat grown , so the wheat flour used as a bread 🍞 or bread equivalent like roti in South Asia and Mediterranean bread or Nan in the middle east, provides these recommend requirements, during the meal which consists primarily of roti / dal (lentils) and A vegetable 🥦🥕🌽 which provides the protein necessary. Ensuring hunger is banished using such technology should be given number one priority.
2. Soil conservation and ensuring fertility of Soil using known methodologies like crop rotation etc. and ensuring the appropriate conditions for Soil health is a critical requirement.
3. Rice and it’s numerous varieties is the second major crop which is consumed globally. Though it is a water intensive crop, it’s energy content is multiple times of the wheat crops. So, the amount of rice required to provide the energy requirements of human cells is probably much lower. So it’s consumption should be limited to provide daily energy requirements . Of course plant based lentils and dals should be a regular part of the neal. Was wondering if similar studies on the Genetic pool of Rice has been done?
The Daily necessary Recommended requirements of trace minerals and FDA should be included in the crop stage itself.
Diversity is critical for survival of the planet and just as in business and work place, the ensurance of variety and diversity will result in better resilience and survivability. The wild strains of rice and wheat have evolved and survived and probably have hardier genes / genetic characteristics.
4. There are many other grains and fruits🍍🍎🍓🍇 and vegetables which are highly nutritious and addition of nuts and dried fruits 🍍🍎🍓🍇 can more than fulfill human needs. Time for Chefs and cooks to step up and show their creative talent without using the flesh of dead animals in the diet. The same tastes can be made far more cheaply without fear of infection by unfriendly pathogens, bacteria and viruses.
Views expressed are personal and not binding on anyone.
Good Comments…I was would like to add that besides the toxicity of chemical controls, fungal mycotoxins are waiting solution. We have been at this long enough now to stop making people sick when it is both within our power and ability to not only make the changes… but to simply and effectively detect the tasteless odorless poisons Some people are more sensitive, having reactions and disease that doctors seems to misunderstand. I have spent the past 7 years sick from these and I was told that it was just something that happens to some people. Make the adjustment?? Cancer, chronic fatigue, fibromyalgia, autism, nervous disorders, dementia the list goes on…and on. I’m outraged. Just start reading about fungal infections. Browse mycotoxins. I found out inadvertently when what I was told was a bone infection actually bloomed.and produced spores. If I’d not been more afraid of the local medics cutting off more body parts o guess I would never have known. Speaking to medical professionals really opens the eyes. The information is out there. It’s not new. Yet, they do not know and are not prepared to take action. Arizona is on point with the Coccidia mycosis. A fine point made for no till agriculture I might add. The regulations for feeding the mycotoxins to farm animals are tighter than controls for our bread? We can definitely do better.
You left out the #1 grain crop. corn.