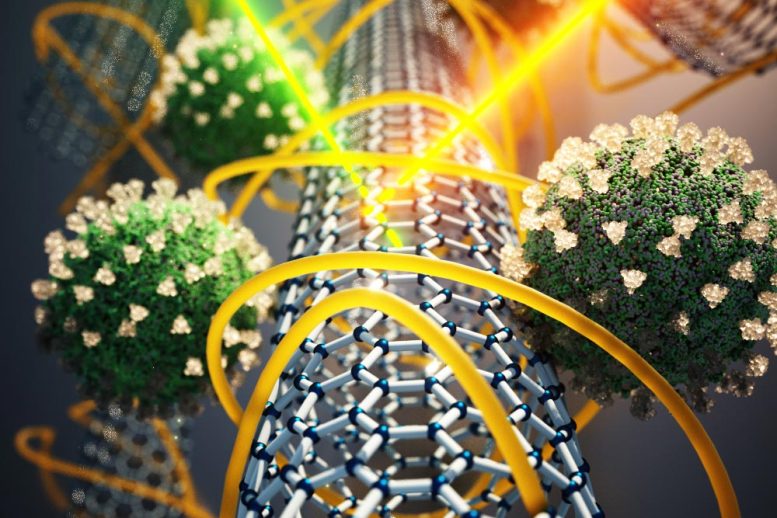

Using specialized carbon nanotubes, MIT engineers have designed a novel sensor that can detect SARS-CoV-2 without any antibody, giving a result within minutes. Credit: Courtesy of the researchers
The technology could be developed as a rapid diagnostic for COVID-19 or other emerging pathogens.
Using specialized carbon nanotubes, MIT engineers have designed a novel sensor that can detect SARS-CoV-2 without any antibodies, giving a result within minutes. Their new sensor is based on technology that can quickly generate rapid and accurate diagnostics, not just for COVID-19 but for future pandemics, the researchers say.
“A rapid test means that you can open up travel much earlier in a future pandemic. You can screen people getting off of an airplane and determine whether they should quarantine or not. You could similarly screen people entering their workplace and so forth,” says Michael Strano, the Carbon P. Dubbs Professor of Chemical Engineering at MIT and the senior author of the study. “We do not yet have technology that can develop and deploy such sensors fast enough to prevent economic loss.”
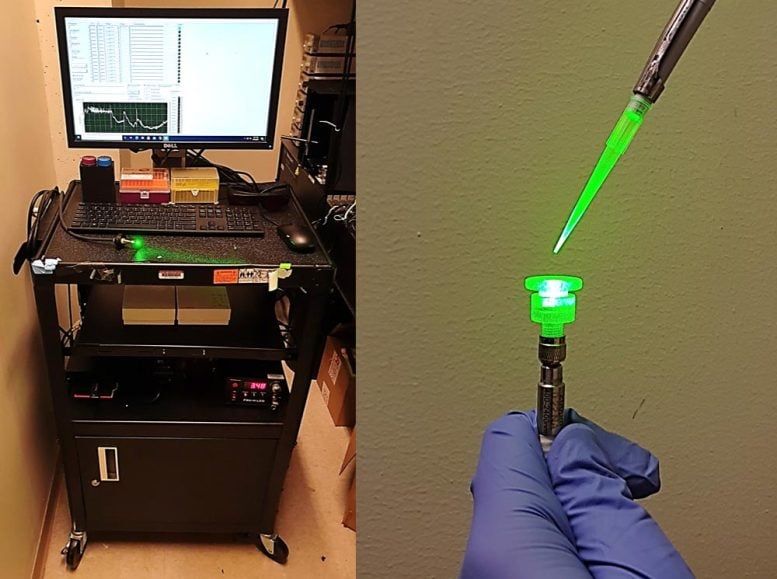

The researchers incorporated their sensor into a prototype with a fiber optic tip that can detect changes in fluorescence in the test sample. Credit: MIT
The diagnostic is based on carbon nanotube sensor technology that Strano’s lab has previously developed. Once the researchers began working on a COVID-19 sensor, it took them just 10 days to identify a modified carbon nanotube capable of selectively detecting the viral proteins they were looking for, and then test it and incorporate it into a working prototype. This approach also eliminates the need for antibodies or other reagents that are time-consuming to generate, purify, and make widely available.
MIT postdoc Sooyeon Cho and graduate student Xiaojia Jin are the lead authors of the paper, which appears today (October 26, 2021) in the journal Analytical Chemistry. Other authors include MIT graduate students Sungyun Yang and Jianqiao Cui, and postdoc Xun Gong.
Molecular recognition
Several years ago, Strano’s lab developed a novel approach to designing sensors for a variety of molecules. Their technique relies on carbon nanotubes — hollow, nanometer-thick cylinders made of carbon that naturally fluoresce when exposed to laser light. They have shown that by wrapping such tubes in different polymers, they can create sensors that respond to specific target molecules by chemically recognizing them.
Their approach, known as Corona Phase Molecular Recognition (CoPhMoRe), takes advantage of a phenomenon that occurs when certain types of polymers bind to a nanoparticle. Known as amphiphilic polymers, these molecules have hydrophobic regions that latch onto the tubes like anchors and hydrophilic regions that form a series of loops extending away from the tubes.
Those loops form a layer called a corona surrounding the nanotube. Depending on the arrangement of the loops, different types of target molecules can wedge into the spaces between the loops, and this binding of the target alters the intensity or peak wavelength of fluorescence produced by the carbon nanotube.
Earlier this year, Strano and InnoTech Precision Medicine, a Boston-based diagnostics developer, received a National Institutes of Health grant to create a CoPhMoRe sensor for SARS-CoV-2 proteins. Researchers in Strano’s lab had already developed strategies that allow them to predict which amphiphilic polymers will interact best with a particular target molecule, so they were able to quickly generate a set of 11 strong candidates for SARS-CoV-2.
Within about 10 days of starting the project, the researchers had identified accurate sensors for both the nucleocapsid and the spike protein of the SARS-CoV-2 virus. During that time, they also were able to incorporate the sensors into a prototype device with a fiber optic tip that can detect fluorescence changes of the biofluid sample in real time. This eliminates the need to send the sample to a lab, which is required for the gold-standard PCR diagnostic test for COVID-19.
This device produces a result within about five minutes, and can detect concentrations as low as 2.4 picograms of viral protein per milliliter of sample. In more recent experiments done after this paper was submitted, the researchers have achieved a limit of detection lower than the rapid tests that are now commercially available.
The researchers also showed that the device could detect the SARS-CoV-2 nucleocapsid protein (but not the spike protein) when it was dissolved in saliva. Detecting viral proteins in saliva is usually difficult because saliva contains sticky carbohydrate and digestive enzyme molecules that interfere with protein detection, which is why most COVID-19 diagnostics require nasal swabs.
“This sensor shows the highest range of limit of detection, response time, and saliva compatibility even without any antibody and receptor design,” Cho says. “It is a unique feature of this type of molecular recognition scheme that rapid design and testing is possible, unhindered by the development time and supply chain requirements of a conventional antibody or enzymatic receptor.”
Quick response
The speed with which the researchers were able to develop a working prototype suggests that this approach could prove useful for developing diagnostics more quickly during future pandemics, Strano says.
“We’re able to go from someone handing us viral markers to a working fiber optic sensor in an extremely short amount of time,” he says.
Sensors that rely on antibodies to detect viral proteins, which form the basis of many of the rapid COVID-19 tests now available, take much longer to develop because the process of designing the right protein antibody is so time-consuming.
The researchers have filed for a patent on the technology in hopes that it could be commercialized for use as a COVID-19 diagnostic. Strano also hopes to further develop the technology so that it could be deployed quickly in response to future pandemics.
Reference: “Antibody-Free Rapid Detection of SARS-CoV-2 Proteins Using Corona Phase Molecular Recognition to Accelerate Development Time” by Soo-Yeon Cho, Xiaojia Jin, Xun Gong, Sungyun Yang, Jianqiao Cui and Michael S. Strano, 26 October 2021, Analytical Chemistry.
DOI: 10.1021/acs.analchem.1c02889
The research was funded by a National Institutes of Health Rapid Acceleration of Diagnostics (RADx) grant.








While the MSM condemns the use of ivermectin, the most populated state in India just declared they are officially COVID free after promoting widespread use of the safe, proven medicine. In addition to this, Ivermectin attaches to covid spikes and prevents them from binding to ACE2. Get your Ivermectin today while you still can!
Benki 🔥🔥🔥🔥🔥🔥