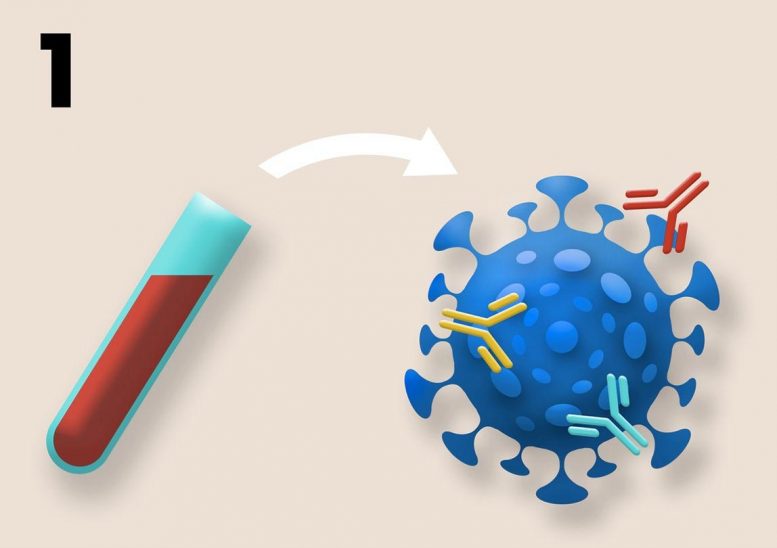
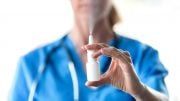
The series of illustrations on this page are a schematic illustrating three ways that standard samples from COVID-19 clinical trials can be repurposed to assess the risk that vaccine resistance will evolve. 1. The complexity of B-cell and T-cell responses can be measured using blood samples. Different neutralizing antibodies are depicted above in different colors. More complex responses indicate more evolutionarily robust immunity. Credit: Kennedy et al, 2020 (PLOS Biology, CC BY 4.0)
Similar to bacteria evolving resistance to antibiotics, viruses can evolve resistance to vaccines, and the evolution of SARS-CoV-2 could undermine the effectiveness of vaccines that are currently under development, according to a paper published today (November 9, 2020) in the open-access journal PLOS Biology by David Kennedy and Andrew Read from Pennsylvania State University, USA. The authors also offer recommendations to vaccine developers for minimizing the likelihood of this outcome.
“A COVID-19 vaccine is urgently needed to save lives and help society return to its pre-pandemic normal,” said David Kennedy, assistant professor of biology. “As we have seen with other diseases, such as pneumonia, the evolution of resistance can quickly render vaccines ineffective. By learning from these previous challenges and by implementing this knowledge during vaccine design, we may be able to maximize the long-term impact of COVID-19 vaccines.”
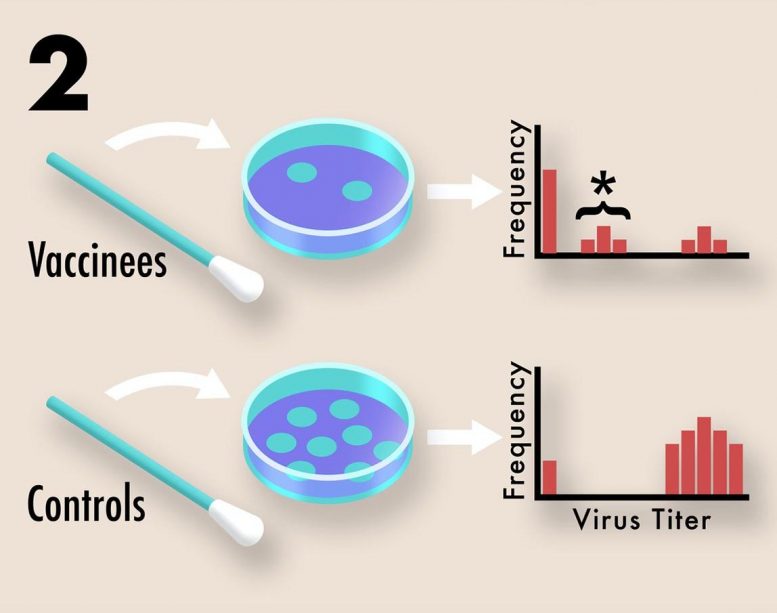
2. The effect of vaccination on transmission potential can be assessed by collecting viral titer data using routine nasal swabs. Plaque assays from multiple vaccinated and control individuals are compiled into a histogram. Undetectable viral titers suggest little or no transmission potential, due to either complete immune protection or the absence of exposure. High viral titers suggest high transmission potential due to the absence of a protective immune response. Intermediate viral titers, marked above with an asterisk, suggest moderate transmission potential due to partial vaccine protection. Intermediate titers indicate an increased risk for resistance evolution since pathogen diversity can be generated within hosts and selection can act during transmission between hosts. Credit: Kennedy et al, 2020 (PLOS Biology, CC BY 4.0)
The researchers specifically suggest that the standard blood and nasal-swab samples taken during clinical trials to quantify individuals’ responses to vaccination may also be used to assess the likelihood that the vaccines being tested will drive resistance evolution. For example, the team proposes that blood samples can be used to assess the redundancy of immune protection generated by candidate vaccines by measuring the types and amounts of antibodies and T-cells that are present.
“Much like how combination antibiotic therapy delays the evolution of antibiotic resistance, vaccines that are designed to induce a redundant immune response — or one in which the immune system is encouraged to target multiple sites, called epitopes — on the virus’s surface, can delay the evolution of vaccine resistance,” said Andrew Read, Evan Pugh Professor of Biology and Entomology and director of the Huck Institutes of the Life Sciences. “That’s because the virus would have to acquire several mutations, as opposed to just one, in order to survive the host immune system’s attack.”
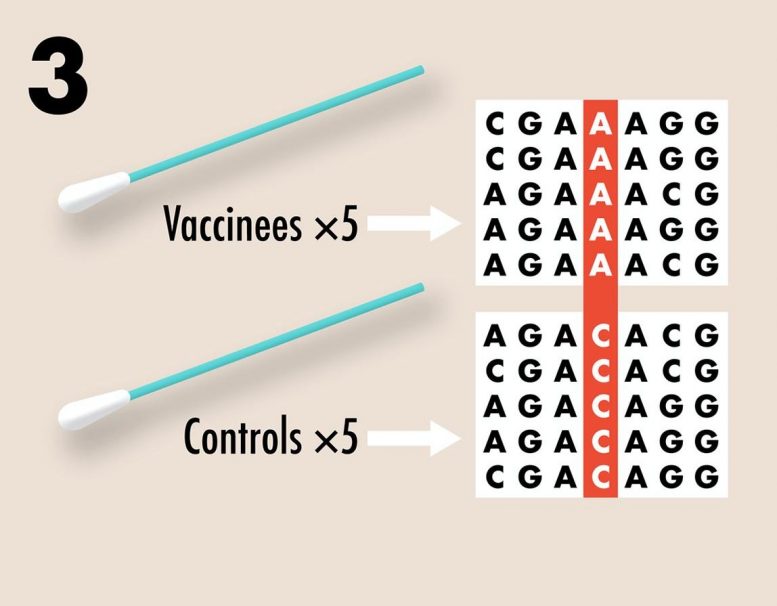
3. Pre-existing variation for vaccine resistance can be assessed by recovering genome sequences from nasopharyngeal swabs of symptomatic COVID-19 cases included in the study. In a placebo controlled, double blind study, any significant differences in the genome sequences of samples from vaccinated and control individuals would suggest at least partial vaccine resistance. Credit: Kennedy et al, 2020 (PLOS Biology, CC BY 4.0)
The researchers also recommend that nasal swabs typically collected during clinical trials may be used to determine the viral titer, or amount of virus present, which can be considered a proxy for transmission potential. They noted that strongly suppressing virus transmission through vaccinated hosts is key to slowing the evolution of resistance, since it minimizes opportunities for mutations to arise and reduces opportunities for natural selection to act on those mutations that do arise.
In addition, the team suggests that the genetic data acquired through nasal swabs can be used to examine whether vaccine-driven selection has occurred. For example, differences in alleles, or forms of genes that arise from mutations, between the viral genomes collected from vaccinated versus unvaccinated individuals would indicate that selection has taken place.
“According to the World Health Organization, at least 198 COVID-19 vaccines are in the development pipeline, with 44 currently undergoing clinical evaluation,” said Kennedy. “We suggest that the risk of resistance be used to prioritize investment among otherwise similarly promising vaccine candidates.”
Reference: “Monitor for COVID-19 vaccine resistance evolution during clinical trials” by David A. Kennedy and Andrew F. Read, 9 November 2020, PLOS Biology.
DOI: 10.1371/journal.pbio.3001000




HELP – Hydrocortisone, eucalyptus, lavender or peppermint help to cure coronavirus and mold allergies. Vaccine propaganda is drug resistance. After years of failing to prescribe prescription flu drugs, the fake news media has evolved an entirely new classification of fake medicine and science to resist cheap cures. A tube of hydrocortisone creme costs only one dollar and only a dab to the nose is needed to cure coronavirus. The FDA has prescribed Lysol, containing eucalyptol, for environmental cleaning. Essential oil of lavender works great in a spray. Peppermint candy does not work fast enough. A half gallon of peppermint ice cream will cure a moderate case of COVID-19 before finishing the bowl. Peppermint tea is a great remedy.
You are a danger to society. Go away.
If you get COVID-19, I sure hope your bowl of ice cream cures you. Moron.
This fake-hope author should be given ample supply of their miracle cure, then taken to a hospital ICU, holding a covid19 patient and be sure they’re exposed. This would be a REAL Clinical trial. Same for ALL of these numbskulls!
Only Donald Trump could save us. Now all is lost…no scientist will actually do anything with vaccines because they are part of the deep state.