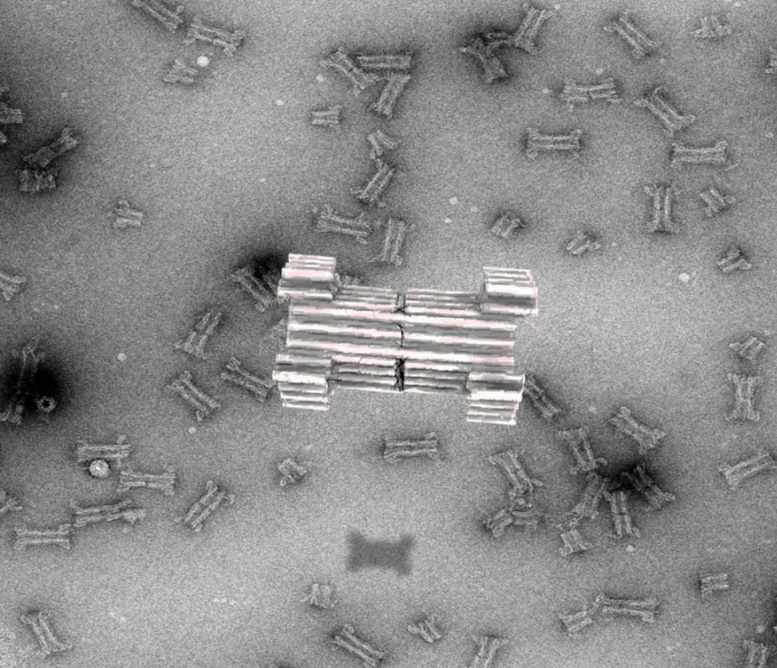

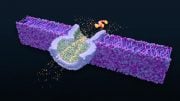
The construction of a synthetic DNA nanopore capable of selectively translocating protein-size macromolecules across lipid bilayers. Credit: Rasmus Peter Thomsen, Aarhus University
A scientific collaboration led by researchers at iNANO/Department of Molecular Biology and Genetics at Aarhus University and the Department of Chemistry at the University of Copenhagen has resulted in the construction of a synthetic DNA nanopore capable of selectively translocating protein-size macromolecules across lipid bilayers.
In 2015, the first commercial nanopore DNA sequencing device was introduced by Oxford Nanopore Technologies. Based on a synthetically engineered transmembrane protein, nanopore sequencing allows long DNA strands to be channeled through the central lumen of the pore where changes in the ionic current work as a sensor of the individual bases in the DNA. This technique was a key milestone for DNA sequencing and the achievement was only made possible after decades of research.
Since then, researchers have tried to extend this principle and build larger pores to accommodate proteins for sensing purposes, but a major challenge has been the limited understanding of artificial protein design. As an alternative, a new technique based on the artificial folding of DNA into complex structures, the so-called 3D-origami technique, first reported by the AU group in 2009, has emerged. Compared to proteins, DNA origami has been shown to have an unprecedented design space for constructing nanostructures that mimic and extend naturally occurring complexes.
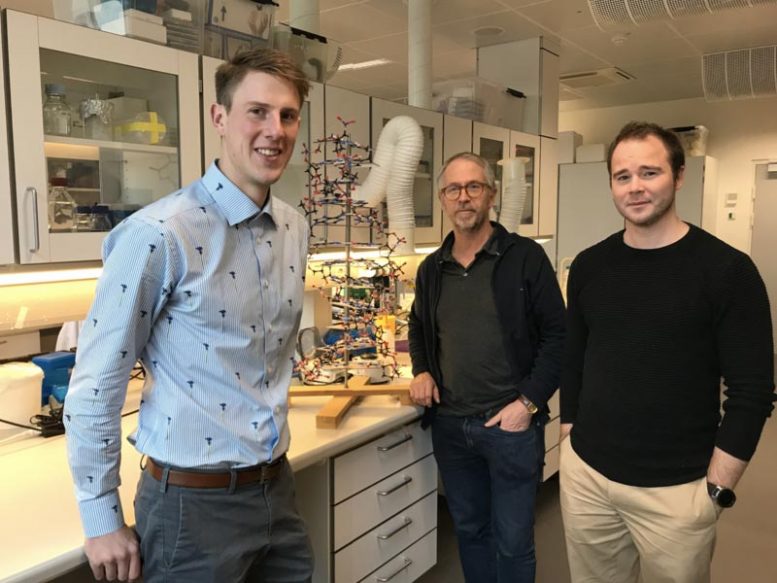

The researchers from Aarhus University behind the scientific article (from left): Rasmus P. Thomsen, Jørgen Kjems and Rasmus Schøler Sørensen. Credit: Anne Færch Nielsen, Aarhus University
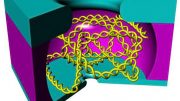
In a new article, published in Nature Communications, the researchers now report the creation of a large synthetic nanopore made from DNA. This nanopore structure is capable of translocating large protein-sized macromolecules between compartments separated by a lipid bilayer. In addition, a functional gating system was introduced inside the pore to enable the biosensing of very few molecules in the solution.
With the use of powerful optical microscopes, the researchers could follow the flow of molecules through individual nanopores. By introducing a controllable plug in the pore, it was furthermore possible to size-selectively control the flow of protein-size molecules and demonstrate label-free, real-time, bio-sensing of a trigger molecule.
Lastly, the pore was equipped with a set of controllable flaps, allowing targeted insertion into membranes displaying particular signal molecules. In the future, this mechanism will potentially enable the insertion of the sensor specifically into diseased cells and may allow diagnosis at the single-cell level.
Reference: “A large size-selective DNA nanopore with sensing applications” by Rasmus P. Thomsen, Mette Galsgaard Malle, Anders Hauge Okholm, Swati Krishnan, Søren S.-R. Bohr, Rasmus Schøler Sørensen, Oliver Ries, Stefan Vogel, Friedrich C. Simmel, Nikos S. Hatzakis and Jørgen Kjems, 11 December 2019, Nature Communications.
DOI: 10.1038/s41467-019-13284-1







Be the first to comment on "Scientists Create Synthetic Nanopores Made From DNA"