
Sequencing the last 8% of the human genome has taken 20 years and the invention of new techniques for reading long sequences of the genetic code, which consists of the nucleotides C, T, G and A. The entire genome consists of more than 3 billion nucleotides. Credit: Ernesto del Aguila III, NHGRI
Repetitive DNA sequences around centromere show history of human genetic variation.
Scientists lied a little when they revealed the entire sequencing of the human genome in 2003. In actuality, almost 20 years later, approximately 8% of the genome has never been completely sequenced, due to highly repetitive DNA segments that are difficult to match with the rest of the genome. However, a three-year-old team has finally filled in the gaps in the remaining DNA, giving scientists and physicians the first complete, gap-free genome sequencing.
The recently completed genome, termed T2T-CHM13, is a significant improvement over the existing reference genome, GRCh38, which is used by physicians and scientists to check for disease-linked mutations as well as to study the evolution of human genetic diversity.
The new DNA sequences, among other things, provide previously unknown details about the area around the centromere, which is where chromosomes are seized and tugged apart as cells split, ensuring that each “daughter” cell acquires the right amount of chromosomes. Variability within this area might potentially provide fresh information about how our ancestors developed in Africa.
“Uncovering the complete sequence of these formerly missing regions of the genome told us so much about how they’re organized, which was totally unknown for many chromosomes,” said Nicolas Altemose, a postdoctoral researcher at the University of California, Berkeley, and co-author of four new articles describing the completed genome. “Before, we just had the blurriest picture of what was there, and now it’s crystal clear down to single base pair resolution.”
Altemose is first author of one paper that describes the base pair sequences around the centromere. A paper explaining how the sequencing was done will appear in the April 1 print edition of the journal Science, while Altemose’s centromere paper and four others describing what the new sequences tell us are summarized in the journal with the full papers posted online. Four companion papers, including one for which Altemose is co-first author, also will appear online April 1 in the journal Nature Methods.
The sequencing and analysis were performed by a team of more than 100 people, the so-called Telemere-to-Telomere Consortium, or T2T, named for the telomeres that cap the ends of all chromosomes. The consortium’s gapless version of all 22 autosomes and the X sex chromosome is composed of 3.055 billion base pairs, the units from which chromosomes and our genes are built, and 19,969 protein-coding genes. Of the protein-coding genes, the T2T team found about 2,000 new ones, most of them disabled, but 115 of which may still be expressed. They also found about 2 million additional variants in the human genome, 622 of which occur in medically relevant genes.
“In the future, when someone has their genome sequenced, we will be able to identify all of the variants in their DNA and use that information to better guide their health care,” said Adam Phillippy, one of the leaders of T2T and a senior investigator at the National Human Genome Research Institute (NHGRI) of the National Institutes of Health. “Truly finishing the human genome sequence was like putting on a new pair of glasses. Now that we can clearly see everything, we are one step closer to understanding what it all means.”
The evolving centromere
The new DNA sequences in and around the centromere total about 6.2% of the entire genome, or nearly 190 million base pairs, or nucleotides. Of the remaining newly added sequences, most are found around the telomeres at the end of each chromosome and in the regions surrounding ribosomal genes. The entire genome is made of just four types of nucleotides, which, in groups of three, code for the amino acids used to build proteins. Altemose’s main research involves finding and exploring areas of the chromosomes where proteins interact with DNA.
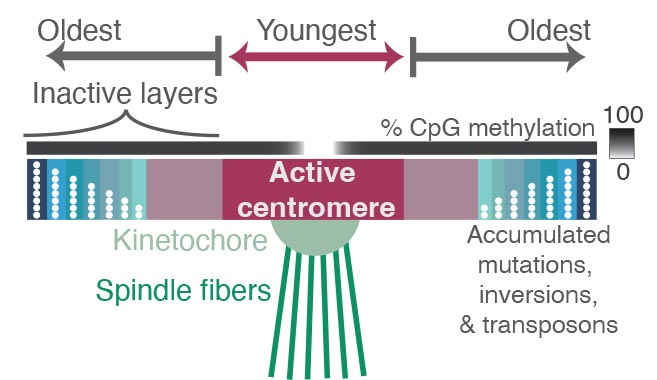
The spindles (green) that pull chromosomes apart during cell division are attached to a protein complex called the kinetochore, which latches onto the chromosome at a place called the centromere — a region containing highly repetitive DNA sequences. Comparing the sequences of these repeats revealed where mutations have accumulated over millions of years, reflecting the relative age of each repeat. Repeats in the active centromere tend to be the youngest and most recently duplicated sequences in the region, and they have strikingly low DNA methylation. Surrounding the active centromere on both sides are older repeats, probably the relics of former centromeres, with the oldest ones farthest from the active centromere. The researchers hope that new experimental methods will help reveal why centromeres evolve from the middle, as well as why this pattern is so closely associated with binding by the kinetochore and with low DNA methylation. Credit: Nicolas Altemose, UC Berkeley
“Without proteins, DNA is nothing,” said Altemose, who earned a Ph.D. in bioengineering jointly from UC Berkeley and UC San Francisco in 2021 after having received a D.Phil. in statistics from Oxford University. “DNA is a set of instructions with no one to read it if it doesn’t have proteins around to organize it, regulate it, repair it when it’s damaged and replicate it. Protein-DNA interactions are really where all the action is happening for genome regulation, and being able to map where certain proteins bind to the genome is really important for understanding their function.”
After the T2T consortium sequenced the missing DNA, Altemose and his team used new techniques to find the place within the centromere where a big protein complex called the kinetochore solidly grips the chromosome so that other machines inside the nucleus can pull chromosome pairs apart.
“When this goes wrong, you end up with missegregated chromosomes, and that leads to all kinds of problems,” he said. “If that happens in meiosis, that means you can have chromosomal anomalies leading to spontaneous miscarriage or congenital diseases. If it happens in somatic cells, you can end up with cancer — basically, cells that have massive misregulation.”
What they found in and around the centromeres were layers of new sequences overlaying layers of older sequences, as if through evolution new centromere regions have been laid down repeatedly to bind to the kinetochore. The older regions are characterized by more random mutations and deletions, indicating they’re no longer used by the cell. The newer sequences where the kinetochore binds are much less variable, and also less methylated. The addition of a methyl group is an epigenetic tag that tends to silence genes.
All of the layers in and around the centromere are composed of repetitive lengths of DNA, based on a unit about 171 base pairs long, which is roughly the length of DNA that wraps around a group of proteins to form a nucleosome, keeping the DNA packaged and compact. These 171 base pair units form even larger repeat structures that are duplicated many times in tandem, building up a large region of repetitive sequences around the centromere.
The T2T team focused on only one human genome, obtained from a non-cancerous tumor called a hydatidiform mole, which is essentially a human embryo that rejected the maternal DNA and duplicated its paternal DNA instead. Such embryos die and transform into tumors. But the fact that this mole had two identical copies of the paternal DNA — both with the father’s X chromosome, instead of different DNA from both mother and father — made it easier to sequence.
The researchers also released this week the complete sequence of a Y chromosome from a different source, which took nearly as long to assemble as the rest of the genome combined, Altemose said. The analysis of this new Y chromosome sequence will appear in a future publication.
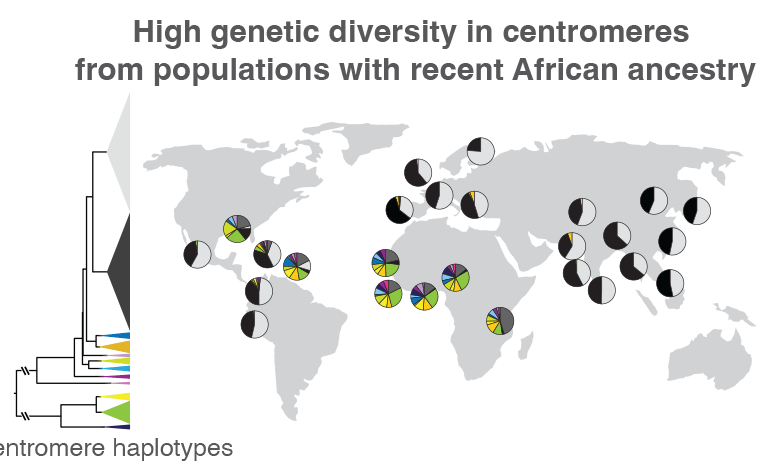
When the researchers compared centromeric regions of 1,600 people from around the world, they found that those without recent African ancestry mostly had two types of sequence variations. The proportions of these two variations are represented by the black and light gray wedges within the circles, which are placed on the map near the location where each group of individuals was sampled. Those from Africa or other areas with a large proportion of people with recent African ancestry, like the Caribbean, had much more centromeric sequence variation, represented by the multi-colored wedges. Such variations could help track how centromeric regions evolve, as well as how these genetic variants are related to health and disease. Credit: Nicolas Altemose, UC Berkeley
Altemose and his team, which included UC Berkeley project scientist Sasha Langley, also used the new reference genome as a scaffold to compare the centromeric DNA of 1,600 individuals from around the world, revealing major differences in both the sequence and copy number of repetitive DNA around the centromere. Previous studies have shown that when groups of ancient humans migrated out of Africa to the rest of the world, they took only a small sample of genetic variants with them. Altemose and his team confirmed that this pattern extends into centromeres.
“What we found is that in individuals with recent ancestry outside the African continent, their centromeres, at least on chromosome X, tend to fall into two big clusters, while most of the interesting variation is in individuals who have recent African ancestry,” Altemose said. “This isn’t entirely a surprise, given what we know about the rest of the genome. But what it suggests is that if we want to look at the interesting variation in these centromeric regions, we really need to have a focused effort to sequence more African genomes and do complete telomere-to-telomere sequence assembly.”
DNA sequences around the centromere could also be used to trace human lineages back to our common ape ancestors, he noted.
“As you move away from the site of the active centromere, you get more and more degraded sequence, to the point where if you go out to the furthest shores of this sea of repetitive sequences, you start to see the ancient centromere that, perhaps, our distant primate ancestors used to bind to the kinetochore,” Altemose said. “It’s almost like layers of fossils.”
Long-read sequencing a game changer
The T2T’s success is due to improved techniques for sequencing long stretches of DNA at once, which helps when determining the order of highly repetitive stretches of DNA. Among these are PacBio’s HiFi sequencing, which can read lengths of more than 20,000 base pairs with high accuracy. Technology developed by Oxford Nanopore Technologies Ltd., on the other hand, can read up to several million base pairs in sequence, though with less fidelity. For comparison, so-called next-generation sequencing by Illumina Inc. is limited to hundreds of base pairs.
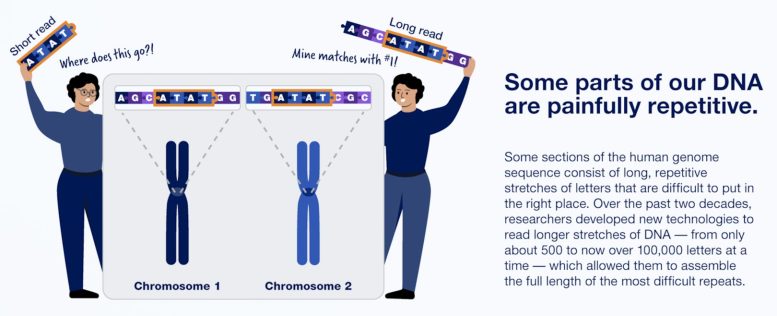
One reason it took 20 years to complete the human genome sequence: much of our DNA is extremely repetitive. Credit: Infographic courtesy of NHGRI, NIH
“These new long-read DNA sequencing technologies are just incredible; they’re such game changers, not only for this repetitive DNA world, but because they allow you to sequence single long molecules of DNA,” Altemose said. “You can begin to ask questions at a level of resolution that just wasn’t possible before, not even with short-read sequencing methods.”
Altemose plans to explore the centromeric regions further, using an improved technique he and colleagues at Stanford developed to pinpoint the sites on the chromosome that are bound by proteins, similar to how the kinetochore binds to the centromere. This technique, too, uses long-read sequencing technology. He and his group described the technique, called Directed Methylation with Long-read sequencing (DiMeLo-seq), in a paper that appeared this week in the journal Nature Methods.
Meanwhile, the T2T consortium is partnering with the Human PanGenome Reference Consortium to work toward a reference genome that represents all of humanity.
“Instead of just having one reference from one human individual or one hydatidiform mole, which isn’t even a real human individual, we should have a reference that represents everybody,” Altemose said. “There are various ideas about how to accomplish that. But what we need first is a grasp of what that variation looks like, and we need lots of high-quality individual genome sequences to accomplish that.”
His work on the centromeric regions, which he called “a passion project,” was funded by postdoctoral fellowships. The leaders of the T2T project were Karen Miga of UC Santa Cruz, Evan Eichler of the University of Washington, and Adam Phillippy of NHGRI, which provided much of the funding. Other UC Berkeley co-authors of the centromere paper are Aaron Streets, assistant professor of bioengineering; Abby Dernburg and Gary Karpen, professors of molecular and cell biology; project scientist Sasha Langley; and former postdoctoral fellow Gina Caldas.
For related research, see Hidden Regions Revealed in First Complete Sequence of a Human Genome.
Reference: “Complete genomic and epigenetic maps of human centromeres” by Nicolas Altemose, Glennis A. Logsdon, Andrey V. Bzikadze, Pragya Sidhwani, Sasha A. Langley, Gina V. Caldas, Savannah J. Hoyt, Lev Uralsky, Fedor D. Ryabov, Colin J. Shew, Michael E. G. Sauria, Matthew Borchers, Ariel Gershman, Alla Mikheenko, Valery A. Shepelev, Tatiana Dvorkina, Olga Kunyavskaya, Mitchell R. Vollger, Arang Rhie, Ann M. McCartney, Mobin Asri, Ryan Lorig-Roach, Kishwar Shafin, Julian K. Lucas, Sergey Aganezov, Daniel Olson, Leonardo Gomes de Lima, Tamara Potapova, Gabrielle A. Hartley, Marina Haukness, Peter Kerpedjiev, Fedor Gusev, Kristof Tigyi, Shelise Brooks, Alice Young, Sergey Nurk, Sergey Koren, Sofie R. Salama, Benedict Paten, Evgeny I. Rogaev, Aaron Streets, Gary H. Karpen, Abby F. Dernburg, Beth A. Sullivan, Aaron F. Straight, Travis J. Wheeler, Jennifer L. Gerton, Evan E. Eichler, Adam M. Phillippy, Winston Timp, Megan Y. Dennis, Rachel J. O’Neill, Justin M. Zook, Michael C. Schatz, Pavel A. Pevzner, Mark Diekhans, Charles H. Langley, Ivan A. Alexandrov and Karen H. Miga, 1 April 2022, Science.
DOI: 10.1126/science.abl4178








So when can it be used as a weapon?
it already is being used as a weapon to convince mankind they evolved from an ape
This is probably the most common misunderstanding of evolution I’ve seen. Man didn’t evolve from apes. Man and apes share a common ancestor.
anyone who keeps saying mankind evolved from an ape doesn’t understand what evolution is. We have a common ancestor with apes, NOT that we evolved from apes. Why is this so hard to understand? Among all of the animals of the animal kingdom, we LOOK like them physically yet people find it hard to fathom that we may be related. it’s more surprising that people don’t see that yet can say with certainty that there is an invisible man somewhere who magically makes things appear.
so you’re saying something gave birth to both monkeys and humans. makes 100% since.
100% since, what? Evolution? Yes, I agree.
Kenny ken ken,
Evolution clearly did not create human beings. You obviously are locked into the Khazarian Deep State Matrix. Good luck coming back to the land of reality.
Hilarious to see Atheists whose concept of God is Santa Claus. Our entire universe is a fractal projection from a single point. Thus it all exists in the mind of God. We are here with all our defects to struggle and fight. And through this we may develop a gestalt intelligent society probably connected through AgI.
However it will be interesting to see if Advanced Crisper type techniques will be able to improve our DNA possibly removing repetitive junk or editing out genetic diseases. Or possibly worst of all increase the length of our telemares.
It seems that we had this ability in Antideluvian eras. But it ended in disaster.
This is the most significant project in human history. It will lead to many incites into the human body. It will lead to shorter trials and better testing for new medication.
Also the earth is flat
Bad. Call me when you find cures for Ms, lupus, diabetes etc.
Sorry..typo! That should have read BFD
It is a BFD for those very reasons..and you just outed yourself.
“When scientists announced the complete sequence of the human genome in 2003, they were fudging a bit.”
So then why would anyone be inclined to believe that this new reporting is any more trustworthy?
Where’s the transgender sequence? If no such sequence exists, can we stop the Emperor’s New Clothes fantasy and acknowledge that transgenderism is predicated on wished and feelings, NOT facts/science. Sorry, feelings are NOT facts.
Sorry…I now understand that gender and sex are two different things. I was conflating them. I apologize for my bigoted narrative.
Let us hope that a gene or two related to fluency and spelling ability in one’s so-called native language is found.
I agre 100 purcent. That wud be profownd!
A friend of mine commented to me that he believes gays have a genetic defect that explain why they are attracted to others (in a sense that they can’t reproduce). This makes total sense, and it would be an amazing scientific discovery to eventually eliminate all types of genetic defects, to include those that believe that you can “be” a different gender than your sex.
You are terribly misinformed if you believe that homosexuality and transgenderism are “defects” of an individual’s genetic code. You’re bordering on Eugenic Theory with your disgusting diatribe.
Do you think there’s a genetic purpose to being homosexual? It appears as though maybe you’re the misinformed one. One of our primary genetic goals is to reproduce. It’s possible that the genetic code of a homosexual person is “aware” of a potential defect “down the line”, and the genetic code has changed to cease it’s own replication. It’s not “disgusting diatribe” if it makes sense scientifically. I personally find it disgusting when a person is so intolerant to the opinions of others, and refuses to accept their ideas. I’m pretty sure there’s a word for that…
The issue that is widely debated is whether or not homosexuality is a genetic status, or a social one; i.e. is it a choice or not? As far as I can tell, the best debates on this subject are solid in concluding that it’s one or the other. I’ve never heard of another point of view, but I’m open to suggestion.
I’ll disagree with your initial premise that it makes total sense that gays have a genetic defect.
There are multiple logic problems with these statements that are attempting to support a presumption that homosexuality is a “defect”. Genetic mutations/adaptations occur and they either end up increasing the odds of propagation or decreasing the odds. From that single perspective, the mutation or adaptation can be seen as advantageous or disadvantageous. Logic problem 1; you are implying that homosexuals do not reproduce. This is false as both gay men and women have children and pass their genes on. One can have sex for pleasure and reproduction. Since homosexuality only seems to be more manifest in the world today, one would have to assume if there was a genetic mutation it was a beneficial one. Logic problem 2; you state homosexuality as a binary problem of either being a choice or deterministic. The highest influence I have ever seen attributed to psychological behavior stemming from genetics is in the range of 30-50% based on twin studies. That leaves at least as much influence to non-genetic factors, and thus, could never be eliminated. Logic problem 3; The only purpose of sexual attraction is reproduction. Sexuality is a spectrum of behaviors, many of which do not involve intercourse and reproduction. Sexual orientation is just one aspect of sexuality. The complexity of the behavior makes it virtually impossible for there to be a targetable set of genes to turn off. Logic problem 4; Labelling something a defect is a subjective choice. I have already established that homosexuality has in no way been something that nature sees as beneficial in terms of natural selection. Humans have exhibited homosexuality since the start of written history and have only multiplied. So what is a defect? Height, hair color, baldness, freckles, left handedness, stuttering, muscle mass, skin color, albinism, eye color, etc. Attempting to eliminate any defect, no matter how subjective weakens the natural selection process where by both random and environmentally induced mutations play out to see what change leads to a more robust species. Over manipulation of genes without a complete understanding of the consequences more than likely would result in a decline in the overall species, much like the practice of royal inbreeding.
So, after the big bang, elements and compounds were formed….when and where exactly did the first DNA appear? The first bit of organic material.
No scientist can explain it. Some scientists babel nonsense about amino acids but they just don’t know.
DEVOLUTION. The movie ‘Idiocracy’ might as well have been prophecy.
Any science denier who takes advantage of modern technology as a result of science are the biggest hypocrites of all.