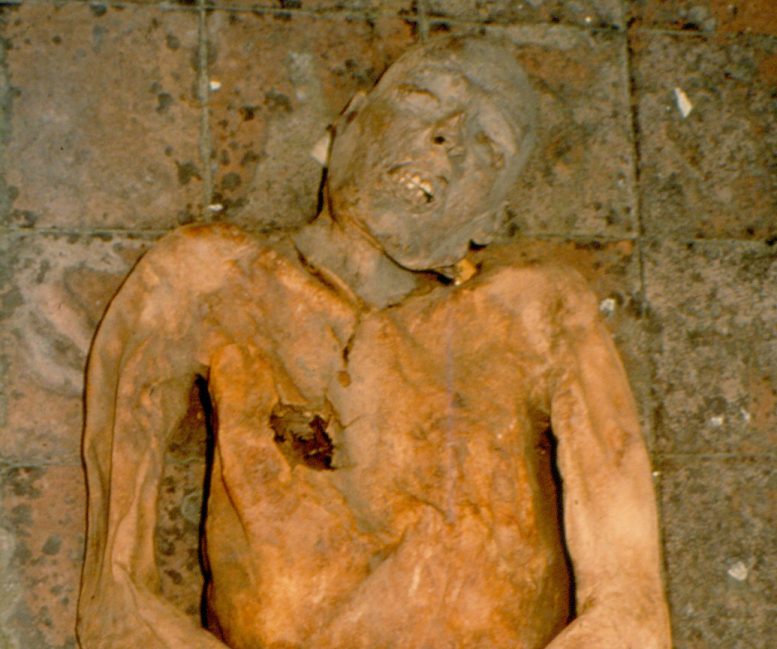
Using pieces of a gallstone from a mummy from the 1500s, researchers have been able to reconstruct the E. coli genome. Credit: Division of Paleopathology of the University of Pisa
Researchers used fragments taken from an Italian mummy to recreate the genome of a centuries-old strain of E. coli
Using fragments taken from a 16th-century mummy’s gallstone, a multinational team headed by scientists from McMaster University and the University of Paris Cité has identified and reconstructed the first ancient E. coli genome.

George Long is a co-lead author of the study and a graduate student of bioinformatics in the Department of Anthropology. Credit: McMaster University
The finding was recently published in the journal Communications Biology.
Despite being a major cause of death and morbidity and significant public health concerns, E. coli does not cause pandemics. It is referred to as a commensal, a kind of bacterium that lives inside of us and may infect its host when conditions are right, such as during times of stress, underlying illness, or immunodeficiency.
According to experts, many details of its evolutionary history, such as when it acquired new genes and antibiotic resistance, are still unknown.
There are no historical records of deaths brought on by commensals like E. coli, in contrast to well-known pandemics like the Black Death, which persisted for centuries and killed as many as 200 million people worldwide. However, its impact on human health and mortality was likely enormous.
“A strict focus on pandemic-causing pathogens as the sole narrative of mass mortality in our past misses the large burden that stems from opportunistic commensals driven by the stress of lives lived,” says evolutionary geneticist Hendrik Poinar, who is director of McMaster’s Ancient DNA Centre and a principal investigator at the university’s Michael G. DeGroote Institute for Infectious Disease Research.
Healthy humans and animals often have modern E. coli in their intestines. While the majority of types are benign, a few of them may cause severe, even deadly food poisoning outbreaks and bloodstream infections. The resilient and adaptable bacterium is known to be particularly resistant to treatment.
Researchers now have a benchmark for comparing how the genome of the present bacteria has changed and adapted over the last 400 years thanks to the discovery of a 400-year-old ancestor.
The well-preserved corpses of a group of Italian nobility whose mummified remains were utilized in the new study were found at the Abbey of Saint Domenico Maggiore near Naples in 1983.
For the study, the researchers conducted a detailed analysis of one of the individuals, Giovani d’Avalos. A Neapolitan noble from the Renaissance period, he was 48 when he died in 1586, and thought to have suffered from chronic inflammation of the gallbladder due to gallstones.
“When we were examining these remains, there was no evidence to say this man had E. coli. Unlike an infection like smallpox, there are no physiological indicators. No one knew what it was,” explains the lead author of the study, George Long, a graduate student of bioinformatics at McMaster who conducted the analysis with co-lead author Jennifer Klunk, a former graduate student in the university’s Department of Anthropology.
The technological feat is particularly remarkable because E. coli is both complex and ubiquitous, living not only in the soil but also in our own microbiomes. Researchers had to meticulously isolate fragments of the target bacterium, which had been degraded by environmental contamination from many sources. They used the recovered material to reconstruct the genome.
“It was so stirring to be able to type this ancient E. coli and find that while unique it fell within a phylogenetic lineage characteristic of human commensals that are today still causing gallstones,” says Erick Denamur, the leader of the French team that was involved in the strain characterization.
“We were able to identify what was an opportunistic pathogen, dig down to the functions of the genome, and to provide guidelines to aid researchers who may be exploring other, hidden pathogens,” says Long.
Reference: “A 16th century Escherichia coli draft genome associated with an opportunistic bile infection” by George S. Long, Jennifer Klunk, Ana T. Duggan, Madeline Tapson, Valentina Giuffra, Lavinia Gazzè, Antonio Fornaciari, Sebastian Duchene, Gino Fornaciari, Olivier Clermont, Erick Denamur, G. Brian Golding, and Hendrik Poinar, 16 June 2022, Communications Biology.
DOI: 10.1038/s42003-022-03527-1
The study was funded by the Canadian Institute of Advanced Research.









Be the first to comment on "Scientists Successfully Reconstruct Ancient Genome Using a 600-Year-Old Mummy"