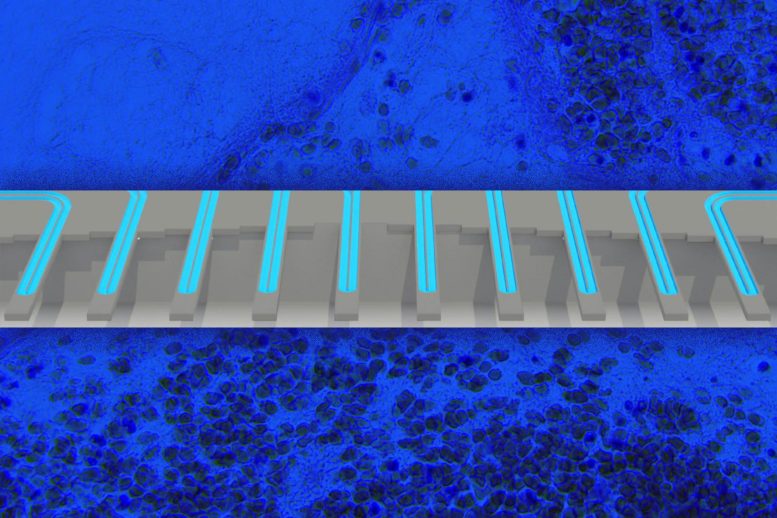

Researchers at MIT and Dana-Farber Cancer Institute have developed a new way to determine whether an individual cancer patient will respond to a particular drug. Their method involves exposing cells to the drug and then measuring changes in their mass using a device similar to the one shown. Credit: Figure courtesy of the researchers. Background: iStockPhoto
A new study shows a link between patient survival and changes in tumor cell mass after glioblastoma treatment.
Researchers at MIT and Dana-Farber Cancer Institute have developed a new way to determine whether individual patients will respond to a specific cancer drug or not. This kind of test could help doctors to choose alternative therapies for patients who don’t respond to the therapies normally used to treat their cancer.
The new technique, which involves removing tumor cells from patients, treating the cells with a drug, and then measuring changes in the cells’ mass, could be applied to a wide variety of cancers and drug treatments, says Scott Manalis, the David H. Koch Professor of Engineering in the departments of Biological Engineering and Mechanical Engineering, and a member of the Koch Institute for Integrative Cancer Research.
“Essentially all of the clinically used cancer drugs either directly or indirectly stop the growth of cancer cells,” Manalis says. “That’s why we think measuring mass could offer a universal readout of the effects of a lot of different types of drug mechanisms.”
The new study, which focused on glioblastoma, an aggressive form of brain cancer, is part of a collaboration between the Koch Institute and Dana-Farber Precision Medicine programs to find new biomarkers and diagnostic tests for cancer.
Manalis and Keith Ligon, director of the Center for Patient Derived Models at Dana-Farber and an associate professor at Harvard Medical School, are the senior authors of the study, which was published on October 5, 2021, in Cell Reports. The lead authors of the paper are Max Stockslager SM ’17, PhD ’20 and Dana-Farber research technician Seth Malinowski.
Measuring cancer cells
Glioblastoma, which is diagnosed in about 13,000 Americans per year, is incurable, but radiation and drug treatment can help to prolong patients’ expected lifespan. Most do not survive longer than one to two years.
“With this disease, you don’t have much time to make adjustments. So, if you take an ineffective drug for six months, that’s pretty significant,” Ligon says. “This kind of assay could help to speed up the learning process for each individual patient and help with decision-making.”
Patients diagnosed with glioblastoma are usually given a chemotherapy drug called temozolomide (TMZ). However, this drug only helps about 50 percent of patients.
Currently, doctors can use a genetic marker — methylation of a gene called MGMT — to predict whether patients will respond to TMZ treatment. Patients who have this marker usually respond better to the drug. However, the marker doesn’t offer reliable predictions for all patients because of other genetic factors. For patients who don’t respond to TMZ, there are a handful of alternative drugs available, Ligon says, or patients can choose to participate in a clinical trial.
In recent years, Manalis and Ligon have been working on a new approach to predicting patient responses, which is based on measuring how tumor cells respond to treatment, rather than genomic signatures. This approach is known as functional precision medicine.
“The idea behind functional precision medicine is that, for cancer, you could take a patient’s tumor cells, give them the drugs that the patient might get, and predict what would happen, before giving them to the patient,” Ligon says.
Scientists are working on many different approaches to functional precision medicine, and one technique that Manalis and Ligon have been pursuing is measuring changes in cell mass that occur following drug treatment. The approach is based on a technology developed by Manalis’ lab for weighing single cells with extremely high accuracy by flowing them through vibrating microchannels.
Several years ago, Manalis, Ligon, and their colleagues demonstrated that they could use this technology to analyze how two types of cancer, glioblastoma and acute lymphoblastic leukemia, respond to treatment. This result was based on measuring individual cells multiple times after drug treatment, allowing the researchers to calculate how their growth rate changed over time following treatment. They showed that this statistic, which they called mass accumulation rate (MAR), was strongly predictive of whether the cells were susceptible to a given drug.
Using a high-throughput version of this system, which they developed in 2016, they could calculate an accurate MAR using just 100 cells per patient. However, a drawback to the MAR technique is that the cells must remain in the system for several hours, so they can be weighed over and over, in order to calculate the growth rate over time.
In their new study, the researchers decided to see if a simpler and significantly faster approach — measuring subtle changes in single-cell mass distributions between drug-treated and untreated cancer cells — would be able to predict patient survival. They performed a retrospective study with a set of live glioblastoma tumor cells from 69 patients, donated to the Ligon lab and the Dana-Farber Center for Patient Derived Models, and used them to grow spheroid tissue cultures. After separating the cells, the researchers treated them with TMZ and then measured their mass a few days later.
They found that by simply measuring the mass difference between cells before and after treatment, using as few as 2,000 cells per patient sample, they could accurately predict whether the patient had responded to TMZ or not.
Better predictions
The researchers showed that their mass measurement was just as accurate as the MGMT methylation marker, but mass measurement has an added advantage in that it can work in patients for whom the genetic marker doesn’t reveal TMZ susceptibility. For many other types of cancer, there are no biomarkers that can be used to predict drug response.
“Most cancers do not have a genomic marker that can be used at all. What we argue is that this functional approach could work in other situations where you don’t have any option of a genomic marker,” Manalis says.
Because the test works by measuring changes in mass, it can be used to observe the effects of many different types of cancer drugs, regardless of their mechanism of action. TMZ works by arresting the cell cycle, which causes cells to become larger because they can no longer divide but they still increase their mass. Other cancer drugs work by interfering with cell metabolism or damaging their structure, which also affect cell mass.
The researchers’ long-term hope is that this approach could be used to test several different drugs on an individual patient’s cells, to predict which treatment would work best for that patient.
“Ideally we would test the drug the patient was most likely to get, but we would also test for things that would be the backup plan: first-, second-, and third-line therapies, or different combinations of drugs,” says Ligon, who also serves as chief of neuropathology at the Brigham and Women’s Hospital and a consultant in pathology at Boston Children’s Hospital.
Manalis and Ligon have co-founded a company called Travera, which has licensed this technology and is now gathering data from patient samples from several different types of cancer, in hopes of developing clinically validated lab tests that can be used to help patients.
Reference: “Functional drug susceptibility testing using single-cell mass predicts treatment outcome in patient-derived cancer neurosphere models” by Max A. Stockslager, Seth Malinowski, Mehdi Touat, Jennifer C. Yoon, Jack Geduldig, Mahnoor Mirza, Annette S. Kim, Patrick Y. Wen, Kin-Hoe Chow, Keith L. Ligon and Scott R. Manalis, 5 October 2021, Cell Reports.
DOI: 10.1016/j.celrep.2021.109788
The research was funded by the MIT Center for Precision Cancer Medicine, the DFCI Center for Patient Derived Models, the Cancer Systems Biology Consortium of the National Cancer Institute, and the Koch Institute Support (core) Grant from the National Cancer Institute.





Be the first to comment on "Weighing Cancer Cells To Personalize Drug Choices and Help With Treatment Decision-Making"