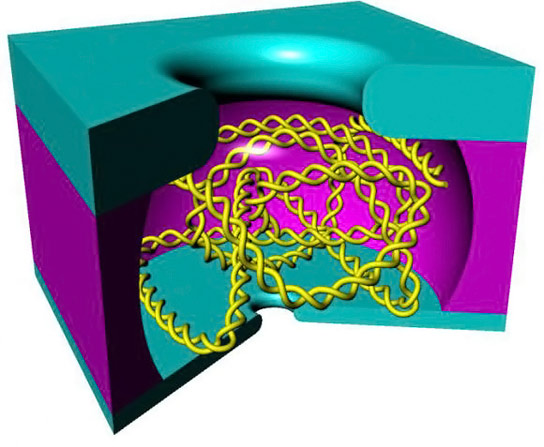
Brown University researchers have designed a nanoscale cage that can trap a single DNA strand and allow before-and-after sequencing of the same DNA strand.
Providence, Rhode Island (Brown University) — Despite having a diameter tens of thousands of times smaller than a human hair, nanopores could be the next big thing in DNA sequencing. By zipping DNA molecules through these tiny holes, scientists hope to one day read off genetic sequences in the blink of an eye.
Now, researchers from Brown University have taken the potential of nanopore technology one step further. They have combined a nanopore with a tiny cage capable of trapping and holding a single DNA strand after it has been pulled through the pore. While caged, biochemical experiments can be performed on the strand, which can then be zipped back through the nanopore to look at how the strand has changed.
“We see this as a very interesting enabling technique,” said Derek Stein, associate professor of physics and engineering at Brown, who helped develop the technology with his graduate students. “It allows you for the first time to look at the same molecule before and after any kind of chemical reaction that may have taken place.”
A paper describing the device is published in Nature Communications.
The device looks a bit like a minuscule hollowed-out hockey puck. On one flat side is a nanopore, and on the other side is a somewhat larger hole. When immersed in a solution containing DNA, an electric current across the nanopore grabs a single strand and pulls it into the hollow chamber. Once there, the strand has a natural tendency to curl into a tangled ball. That ball is too large to fit out of the hole on the other side, but that hole can be used to introduce additional molecules that might react with the trapped DNA. Once a reaction has occurred, the electric current is reversed and the strand is sent back out through the pore, which can look for changes in the strand.
“What we’ve made is basically a very small test tube,” said Xu Liu, who led the work while he was a graduate student at Brown. “We can do biochemistry on the single strand in that very confined space.”
The key to the technology, Liu said, was making that test tube small, but not too small. If it were too small, the DNA wouldn’t have enough room to curl up, which would cause it to squirt out the hole at the top of the device. Using some theoretical calculations and a bit of trial and error, the researchers settled on a cage that’s about 1.5 micrometers square.
Liu then tested the technology using what’s called a restriction enzyme, which cuts DNA molecules at particular sequences. After an intact DNA molecule was pulled through the pore into the cage, the researchers applied the enzyme through the hole in the top of the device. If all went as planned, the enzyme should have cut the strand into four pieces. When they pulled the molecule back through the pore, they detected four distinct signals, indicating that the experiment had worked as expected.
The researchers say the device could be used for all kinds of experiments with DNA. For example, scientists use molecules called hybridization probes to look for specific sequences in a DNA molecule. The probes bind to target sequences, creating a bulge in the DNA strand that a nanopore could easily detect.
“There was always a problem of knowing what the DNA looked like before the probe was applied,” Stein said. “This is a way of making sure you can measure the same molecule before anything is done to it, and then after. That wasn’t possible before with nanopores because the molecule would drift away.”
Liu, who recently received his Ph.D. at Brown, is now working at a nanopore start-up company, where he plans to continue to develop the technology.
The research was supported by the National Science Foundation (CBET0846505) and by Oxford Nanopore Technologies Ltd.
Reference: “Entropic cages for trapping DNA near a nanopore” by Xu Liu, Mirna Mihovilovic Skanata and Derek Stein, 4 February 2015, Nature Communications.
DOI: 10.1038/ncomms7222
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.