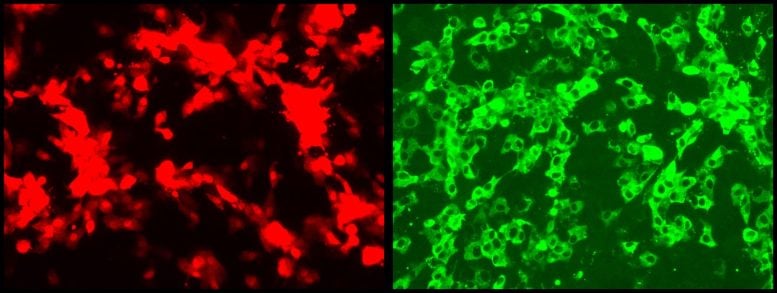
New “reporter viruses” developed by Texas Biomed researchers make it much easier to observe SARS-CoV-2 and its variants in cells and live animals in the lab, and enables faster screening of potential anti-viral drugs, vaccines, and neutralizing antibodies.
A version of SARS-CoV-2, the virus that causes COVID-19 disease, has been successfully modified to glow brightly in cells and animal tissues, providing a real-time way to track the spread and intensity of viral infection as it happens in animal models, researchers at Texas Biomedical Research Institute (Texas Biomed) report in the journal The Proceedings of the National Academy of Sciences (PNAS).
“Now we can track where the virus goes in animal models for COVID-19,” said virologist Luis Martinez-Sobrido, Ph.D., a Professor at Texas Biomed, and senior paper author. “Being able to see how the virus progresses, and which organs and cell types it specifically targets, will be a big help for understanding the virus and optimizing anti-viral drugs and vaccines.”

In addition to tracking the virus, Martinez-Sobrido and his collaborators have already begun using the reporter viruses to screen how well neutralizing antibodies work against different variants of concern, as recently reported in the Journal of Virology.
Turning Up the Lights
To make the reporter virus, Martinez-Sobrido, and his team combined several advanced molecular biology tools to add the genetic sequence for the fluorescent or bioluminescent “reporter” proteins to the virus genetic code. As the virus’s code is replicated and transcribed, so too is the code for the glowing proteins.
In an earlier study, the team replaced one of the virus’s genes with the gene for the glowing proteins, but this resulted in a very dim signal – the gene was not expressed enough to be easily detected in animals. To turn up the brightness, the researchers had to figure out how to get the virus to produce larger quantities of the reporter proteins.
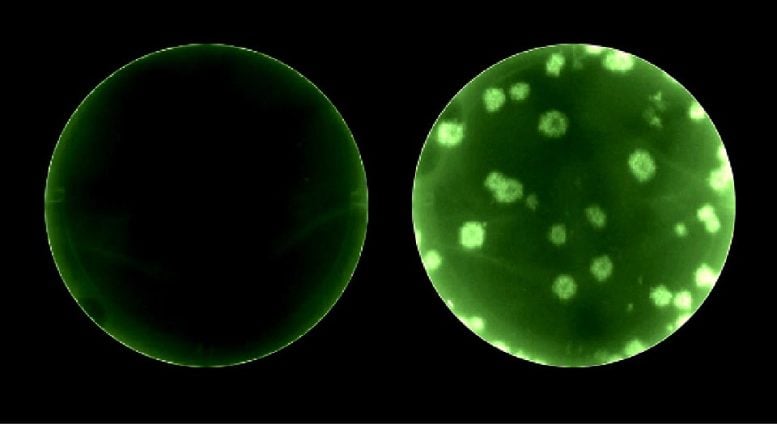
Their solution: they inserted the reporter gene next to a different gene in SARS-CoV-2, specifically, the gene coding for the nucleocapsid protein. “It’s the most expressed protein in SARS-CoV-2,” said molecular biologist Chengjin Ye, Ph.D., a member of Martinez-Sobrido’s lab. This time, the signal was so bright, “it almost blinded me when I looked through the fluorescent microscope,” he said.
Faster Screens
The reporter proteins work in cells and live animal models, in combination with imaging systems that detect the wavelengths of light emitted by the proteins. Being able to observe viral load and location visually offers many advantages over other methods. It is much simpler and faster, saving time and materials.
“Instead of needing a large team to screen 2,000 compounds to see if they work against the virus, one person could do that with a reporter virus in a few hours,” Ye said.
It also enables tracking the virus in the same animal throughout the course of infection and treatment, reducing the number of animals needed to gain similar insights.
Tracking Variants
The team adapted the reporter viruses to express different colored proteins attached to SARS-CoV-2 variants of concern, which they described in a separate paper in the Journal of Virology. Critically, this approach has enabled them to test how well a neutralizing antibody works against two variants in one test well, at the same time.

“This is a significant advantage for saving time and resources, especially with so many basic materials like plastics and reagents in such high demand and limited supply due to the pandemic,” says Kevin Chiem, Ph.D. candidate and member of Martinez-Sobrido’s lab. “As new variants emerge, we can easily adapt the system and quickly screen for how well antibodies work against them.”
Powerful and Accurate
Importantly, the group demonstrated the reporter viruses behave the same as a wild-type version of the virus. This is thanks to the fact they did not remove any viral genes, and because they designed the reporter protein to immediately separate from the virus’s nucleocapsid protein so it functions normally. Their research shows reporter protein brightness correlates well with viral load, although protein accumulation can occur over several days leading to a slightly stronger signal in some cases.
The advancement relies on several powerful techniques, including reverse genetics techniques to generate recombinant SARS-CoV-2, which link together pieces of genetic code to produce the full virus.
Martinez-Sobrido and his team have shared their recombinant SARS-CoV-2 and the noninfectious precursor materials, called plasmids, with more than 100 labs around the world. They can now share the reporter viruses with qualified labs with biocontainment safety level (BSL)-3 access, which is necessary to work with SARS-CoV-2, to help combat the still ongoing COVID-19 pandemic.
“We feel it is our responsibility to share these new tools and technologies with other researchers around the world to help bring the pandemic to an end as quickly as possible,” Martinez-Sobrido said.
Reference: “Analysis of SARS-CoV-2 infection dynamic in vivo using reporter-expressing viruses” by Chengjin Ye, Kevin Chiem, Jun-Gyu Park, Jesus A. Silvas, Desarey Morales Vasquez, Julien Sourimant, Michelle J. Lin, Alexander L. Greninger, Richard K. Plemper, Jordi B. Torrelles, James J. Kobie, Mark R. Walter, Juan Carlos de la Torre and Luis Martinez-Sobrido, 24 September 2021, Proceedings of the National Academy of Sciences.
DOI: 10.1073/pnas.2111593118
Collaborators on these projects include Jun-Gyu Park, Jesus A. Silvas, Desarey Morales Vasquez, and Jordi B. Torrelles at Texas Biomed; Julien Sourimant and Richard K. Plemper at The Center for Translational Antiviral Research at Georgia State University; Michelle J. Lin and Alexander L. Greninger at University of Washington; James J. Kobie, Mark R. Walter and Michael S. Piepenbrink at University of Alabama at Birmingham; and Juan Carlos de la Torre at The Scripps Research Institute.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
1 Comment
Sᴛᴀʀᴛ ᴡᴏʀᴋɪɴɢ ғʀᴏᴍ ʜᴏᴍᴇ! Gʀᴇᴀᴛ ᴊᴏʙ ғᴏʀ Ever, sᴛᴀʏ-ᴀᴛ-ʜᴏᴍᴇ ᴍᴏᴍs ᴏʀ ᴀɴʏᴏɴᴇ ɴᴇᴇᴅɪɴɢ ᴀɴ ᴇxᴛʀᴀ ɪɴᴄᴏᴍᴇ… Yᴏᴜ ᴏɴʟʏ ɴᴇᴇᴅ ᴀ ᴄᴏᴍᴘᴜᴛᴇʀ ᴀɴᴅ ᴀ ʀᴇʟɪᴀʙʟᴇ ɪɴᴛᴇʀɴᴇᴛ ᴄᴏɴɴᴇᴄᴛɪᴏɴ… Mᴀᴋᴇ $100 ʜᴏᴜʀʟʏ ᴀɴᴅ ᴜᴘ ᴛᴏ $13000 ᴀ ᴍᴏɴᴛʜ ʙʏ ғᴏʟʟᴏᴡɪɴɢ connection ᴀᴛ ᴛʜᴇ ʙᴏᴛᴛᴏᴍ .Yᴏᴜ ᴄᴀɴ ʜᴀᴠᴇ ʏᴏᴜʀ ғɪʀsᴛ ᴄʜᴇᴄᴋ ʙʏ ᴛʜᴇ ᴇɴᴅ ᴏғ ᴛʜɪs ᴡᴇᴇᴋ………► 𝐖𝐰𝐰.𝐄𝐱𝐭𝐫𝐚𝐑𝐢𝐜𝐡𝟏.𝐜𝐨𝐦