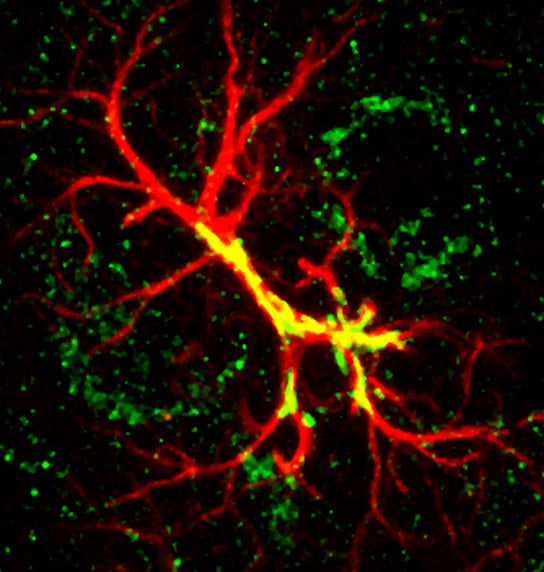
A new study from Duke University provides a close-up of synapse refinement and identifies that the protein hevin is crucial in this process.
Durham, North Carolina – Shortly after birth, human brains expand rapidly with the experience of an entirely new world. During this period, neurons in the newborn brain compete with one another to form lasting connections, called synapses.
A new study by Duke researchers provides a close-up of synapse refinement and identifies a protein that is crucial in this process. Disruptions in the protein, called hevin, have previously been linked to autism, depression, and suicide, but the molecule’s role in the developing brain was mostly unknown until now.
The researchers focused on tiny protrusions of the neuron called spines that harbor synaptic connections. Neuroscience has long assumed that these little nubs serve as sites for single synapses.
But this study, which appeared early online last month in the open access journal eLife, shows that in the brains of newborn mice, some of the spines initially receive two or more inputs. As the brain matures, the spines then receive one. A technique known as three-dimensional electron microscopy made this observation possible.
“I was very excited about this finding,” said first author William Christopher Risher, a postdoctoral researcher in the laboratory of senior author Çagla Eroglu. “I went to check the literature to see if anyone’s really described [multiple-synapse spines] before. And there really hasn’t been much.”
The group also found that mice that are missing the gene that codes for the protein hevin retain more of these multiple synapses compared with normal mice. As the developing brain prunes away synapses to become more efficient, this could present problems.
Hevin was first identified in the miniscule spaces between synapses in 1990. However, gene expression studies showed that it is actually churned out by non-neuronal cells called astrocytes.
Interested in the relationship between astrocytes, synapse formation, and disease, Eroglu’s group showed in 2011 that hevin triggers the formation of new neural connections. “That was the first description of hevin’s function in the nervous system,” said Eroglu, an assistant professor of cell biology and neurobiology, and a member of the Duke Institute for Brain Sciences.
“We continued studying this protein because it is abundant in many brain regions, [both] when synapses are forming and also during adulthood,” Eroglu said.
In the cortex, an area of the brain important for complex thought and awareness, hevin encourages inputs from the thalamus — a part of the brain that acts as a relay center for sensory and motor information — while it discourages inputs from local neurons within the cortex, the group found.
The spines that receive multiple synapses tend to be occupied by both cortical and thalamic connections at the same time, suggesting that these spines are sites for synaptic competition.
The balance of those two types of connections in the cortex could go awry in neurological diseases such as autism and depression, Eroglu said. The group is now studying the molecular mechanisms of hevin and its potential contribution to health and disease.
Other authors include Sagar Patel, Jonnathan Singh Alvarado, Osman Calhan, Il Hwan Kim, Akiyoshi Uezu, and Scott Soderling of Duke’s Cell Biology Department; Srishti Bhagat and Nicole Calakos of Duke Neurology Department; Louis-Jan Pilaz and Debra Silver of Duke’s Molecular Genetics and Microbiology Department; and Daniel Wilton and Beth Stevens of Boston Children’s Hospital, Department of Neurology, Harvard Medical School.
This research was funded by the National Institutes of Health (R01 DA031833, 2T32NS51156-6, NRSA 1F32NS08328 01A1, NS059957, MH103374, NS083897, NS071008), the Holland-Trice Fellowship, the Wakeman Fellowship, the Esther and Joseph Klingenstein Fund, and Alfred P. Sloan Foundation.
Reference: “Astrocytes refine cortical connectivity at dendritic spines” by Sagar Patel, Il Hwan Kim, Akiyoshi Uezu, Srishti Bhagat, Daniel K Wilton, Louis-Jan Pilaz, Jonnathan Singh Alvarado, Osman Y Calhan, Debra L Silver, Beth Stevens, Nicole Calakos, Scott H Soderling and Cagla Eroglu, 17 December 2014, eLife.
DOI: 10.7554/eLife.04047
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.