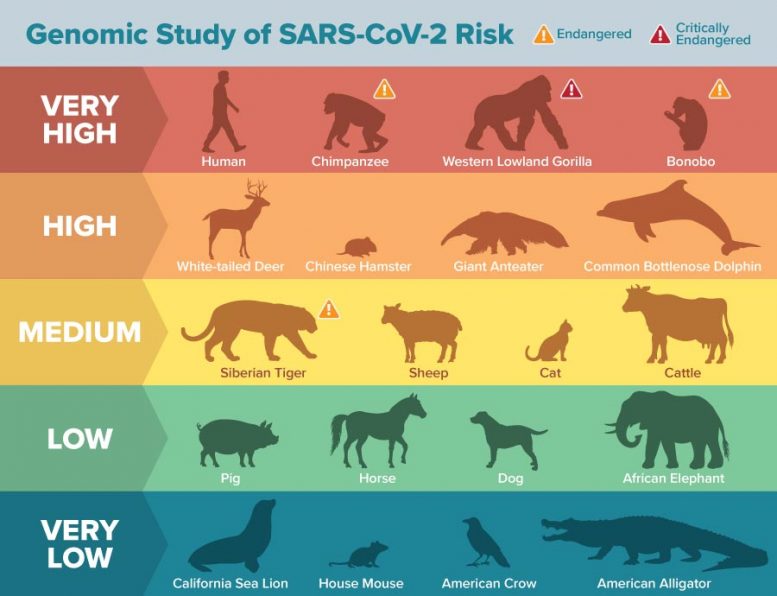
- A large number of mammals can potentially be infected by SARS-CoV-2 through their ACE2 receptors
- Some endangered species are at highest-risk for SARS-CoV-2 infection through ACE2
- These species are potentially vulnerable to spillover of SARS-CoV-2 from humans and should be the focus of surveillance and conservation efforts
- The study can assist in the identification of an intermediate host or hosts for SARS-CoV-2
Humans are not the only species facing a potential threat from SARS-CoV-2, the novel coronavirus that causes COVID-19, according to a new study from the University of California, Davis.
An international team of scientists used genomic analysis to compare the main cellular receptor for the virus in humans — angiotensin converting enzyme-2, or ACE2 — in 410 different species of vertebrates, including birds, fish, amphibians, reptiles, and mammals.
ACE2 is normally found on many different types of cells and tissues, including epithelial cells in the nose, mouth, and lungs. In humans, 25 amino acids of the ACE2 protein are important for the virus to bind and gain entry into cells.
The researchers used these 25 amino acid sequences of the ACE2 protein, and modeling of its predicted protein structure together with the SARS-CoV-2 spike protein, to evaluate how many of these amino acids are found in the ACE2 protein of the different species.
“Animals with all 25 amino acid residues matching the human protein are predicted to be at the highest risk for contracting SARS-CoV-2 via ACE2,” said Joana Damas, first author for the paper and a postdoctoral research associate at UC Davis. “The risk is predicted to decrease the more the species’ ACE2 binding residues differ from humans.”
About 40 percent of the species potentially susceptible to SARS-CoV-2 are classified as “threatened” by the International Union for Conservation of Nature and may be especially vulnerable to human-to-animal transmission. The study was published August 21 in the Proceedings of the National Academy of Sciences.
“The data provide an important starting point for identifying vulnerable and threatened animal populations at risk of SARS-CoV-2 infection,” said Harris Lewin, lead author for the study and a distinguished professor of evolution and ecology at UC Davis. “We hope it inspires practices that protect both animal and human health during the pandemic.”
Endangered species predicted to be at risk
Several critically endangered primate species, such as the Western lowland gorilla, Sumatran orangutan, and Northern white-cheeked gibbon, are predicted to be at very high risk of infection by SARS-CoV-2 via their ACE2 receptor.
Other animals flagged as high risk include marine mammals such as gray whales and bottlenose dolphins, as well as Chinese hamsters.
Domestic animals such as cats, cattle, and sheep were found to have a medium risk, and dogs, horses, and pigs were found to have low risk for ACE2 binding. How this relates to infection and disease risk needs to be determined by future studies, but for those species that have known infectivity data, the correlation is high.
In documented cases of SARS-COV-2 infection in mink, cats, dogs, hamsters, lions, and tigers, the virus may be using ACE2 receptors, or they may use receptors other than ACE2 to gain access to host cells. Lower propensity for binding could translate to lower propensity for infection, or lower ability for the infection to spread in an animal or between animals once established.
Because of the potential for animals to contract the novel coronavirus from humans, and vice versa, institutions including the National Zoo and the San Diego Zoo, which both contributed genomic material to the study, have strengthened programs to protect both animals and humans.
“Zoonotic diseases and how to prevent human-to-animal transmission is not a new challenge to zoos and animal care professionals,” said co-author Klaus-Peter Koepfli, senior research scientist at Smithsonian-Mason School of Conservation and former conservation biologist with the Smithsonian Conservation Biology Institute’s Center for Species Survival and Center for Conservation Genomics. “This new information allows us to focus our efforts and plan accordingly to keep animals and humans safe.”
The authors urge caution against overinterpreting the predicted animal risks based on the computational results, noting the actual risks can only be confirmed with additional experimental data. The list of animals can be found here.
Research has shown that the immediate ancestor of SARS-CoV-2 likely originated in a species of bat. Bats were found to be at very low risk of contracting the novel coronavirus via their ACE2 receptor, which is consistent with actual experimental data.
Whether bats directly transmitted the novel coronavirus directly to humans, or whether it went through an intermediate host, is not yet known, but the study supports the idea that one or more intermediate hosts were involved. The data allow researchers to zero in on which species might have served as an intermediate host in the wild, assisting efforts to control a future outbreak of SARS-CoV-2 infection in human and animal populations.
Reference: “Broad host range of SARS-CoV-2 predicted by comparative and structural analysis of ACE2 in vertebrates” by Joana Damas, Graham M. Hughes, Kathleen C. Keough, Corrie A. Painter, Nicole S. Persky, Marco Corbo, Michael Hiller, Klaus-Peter Koepfli, Andreas R. Pfenning, Huabin Zhao, Diane P. Genereux, Ross Swofford, Katherine S. Pollard, Oliver A. Ryder, Martin T. Nweeia, Kerstin Lindblad-Toh, Emma C. Teeling, Elinor K. Karlsson and Harris A. Lewin, 21 August 2020, Proceedings of the National Academy of Sciences.
DOI: 10.1073/pnas.2010146117
Additional authors on the study include: Marco Corbo, UC Davis Genome Center; Graham M. Hughes and Emma C. Teeling, University College Dublin, Ireland; Kathleen C. Keough and Katherine S. Pollard, Gladstone Institutes and UC San Francisco; Corrie A. Painter, Nicole S. Persky, Diane P. Genereux, Ross Swofford, Kerstin Lindblad-Toh and Elinor K. Karlsson, Broad Institute of MIT and Harvard, Cambridge, Mass.; Michael Hiller, Max Planck Institute of Molecular Cell Biology and Genetics, Dresden, Germany; Andreas R. Pfenning, Carnegie Mellon University, Pittsburgh; Huabin Zhao, Wuhan University, Wuhan, China; Oliver A. Ryder, San Diego Zoo Institute for Conservation Research, Escondido, and UC San Diego; Martin T. Nweeia, Harvard School of Dental Medicine, Boston, and Smithsonian Institution, Washington, D.C.
The research in this study was coordinated as part of the Genome 10K Organization, which includes the Bat1K, Zoonomia, the Vertebrate Genomes Project and the Earth BioGenome Project. Genomic information for the study was also provided the National Center for Biotechnology Information’s GenBank, the San Diego Zoo’s Frozen Zoo and the Smithsonian’s Global Genome Initiative. This work was supported by the Robert and Rosabel Osborne Endowment.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.