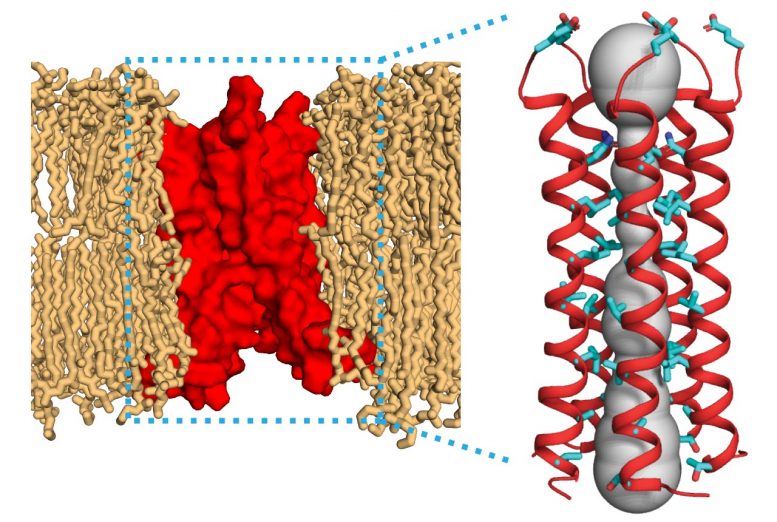
The protein, which acts as an ion channel, could be a target for new drugs against the SARS-CoV-2 virus.
MIT chemists have determined the molecular structure of a protein found in the SARS-CoV-2 virus. This protein, called the envelope protein E, forms a cation-selective channel and plays a key role in the virus’s ability to replicate itself and stimulate the host cell’s inflammation response.
If researchers could devise ways to block this channel, they may be able to reduce the pathogenicity of the virus and interfere with viral replication, says Mei Hong, an MIT professor of chemistry. In this study, the researchers investigated the binding sites of two drugs that block the channel, but these drugs bind only weakly, so they would not be effective inhibitors of the E protein.
“Our findings could be useful for medicinal chemists to design alternative small molecules that target this channel with high affinity,” says Hong, who is the senior author of the new study.
MIT graduate student Venkata Mandala is the lead author of the paper, which appears in Nature Structural and Molecular Biology. Other authors include MIT postdoc Matthew McKay, MIT graduate students Alexander Shcherbakov and Aurelio Dregni, and Antonios Kolocouris, a professor of pharmaceutical chemistry at the University of Athens.
Structural challenges
Hong’s lab specializes in studying the structures of proteins that are embedded in cell membranes, which are often challenging to analyze because of the disorder of the lipid membrane. Using nuclear magnetic resonance (NMR) spectroscopy, she has previously developed several techniques that allow her to obtain accurate atomic-level structural information about these membrane-embedded proteins.
When the SARS-CoV-2 / COVID-19 outbreak began earlier this year, Hong and her students decided to focus their efforts on one of the novel coronavirus proteins. She narrowed in on the E protein partly because it is similar to an influenza protein called the M2 proton channel, which she has previously studied. Both viral proteins are made of bundles of several helical proteins.
“We determined the influenza B M2 structure after about 1.5 years of hard work, which taught us how to clone, express, and purify a virus membrane protein from scratch, and what NMR experimental strategies to take to solve the structure of a homo-oligomeric helical bundle,” Hong says. “That experience turned out to be the perfect training ground for studying SARS-CoV-2 E.”
The researchers were able to clone and purify the E protein in two and half months. To determine its structure, the researchers embedded it into a lipid bilayer, similar to a cell membrane, and then analyzed it with NMR, which uses the magnetic properties of atomic nuclei to reveal the structures of the molecules containing those nuclei. They measured the NMR spectra for two months, nonstop, on the highest-field NMR instrument at MIT, a 900-megahertz spectrometer, as well as on 800- and 600-megahertz spectrometers.
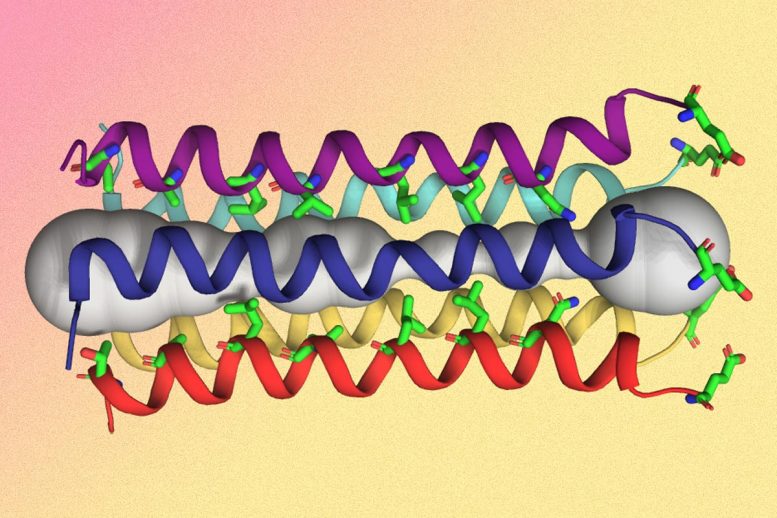
Hong and her colleagues found that the part of the E protein that is embedded in the lipid bilayer, known as the transmembrane domain, assembles into a bundle of five helices. The helices remain largely immobile within this bundle, creating a tight channel that is much more constricted than the influenza M2 channel.
Interestingly, the SARS-CoV-2 E protein looks nothing like the ion channel proteins of influenza and HIV-1 viruses. In flu viruses, the equivalent M2 protein is much more mobile, while in HIV-1, the equivalent Vpu protein has a much shorter transmembrane helix and a wider pore. How these distinct structural features of E affect its functions in the SARS-CoV-2 virus lifecycle is one of the topics that Hong and her colleagues will study in the future.
The researchers also identified several amino acids at one end of the channel that may attract positively charged ions such as calcium into the channel. They believe that the structure they report in this paper is the closed state of the channel, and they now hope to determine the structure of the open state, which should shed light on how the channel opens and closes.
Fundamental research
The researchers also found that two drugs — amantadine, used to treat influenza, and hexamethylene amiloride, used to treat high blood pressure — can block the entrance of the E channel. However, these drugs only bind weakly to the E protein. If stronger inhibitors could be developed, they could be potential drug candidates to treat COVID-19, Hong says.
The study demonstrates that basic scientific research can make important contributions toward solving medical problems, she adds.
“Even when the pandemic is over, it is important that our society recognizes and remembers that fundamental scientific research into virus proteins or bacterial proteins must continue vigorously, so we can preempt pandemics,” Hong says. “The human cost and economic cost of not doing so are just too high.”
Reference: “Structure and drug binding of the SARS-CoV-2 envelope protein transmembrane domain in lipid bilayers” by Venkata S. Mandala, Matthew J. McKay, Alexander A. Shcherbakov, Aurelio J. Dregni, Antonios Kolocouris and Mei Hong, 11 November 2020, Nature Structural & Molecular Biology.
DOI: 10.1038/s41594-020-00536-8
The research was funded by the National Institutes of Health and the MIT School of Science Sloan Fund.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.