UIC researchers prove that drugs designed for bacteria have potential to act on human cells.
According to researchers at the University of Illinois Chicago, the antibiotics used to treat common bacterial infections, like pneumonia and sinusitis, may also be used to treat human diseases, like cancer. Theoretically, at least.
As outlined in a new Nature Communications study, the UIC College of Pharmacy team has shown in laboratory experiments that eukaryotic ribosomes can be modified to respond to antibiotics in the same way that prokaryotic ribosomes do.

Fungi, plants, and animals — like humans — are eukaryotes; they are made up of cells that have a clearly defined nucleus. Bacteria, on the other hand, are prokaryotes. They are made up of cells, which do not have a nucleus and have different structures, sizes, and properties. The ribosomes of eukaryotic and procaryotic cells, which are responsible for the protein synthesis needed for cell growth and reproduction, are also different.
“Some antibiotics, used for treating bacterial infections, work in an interesting way. They bind to the ribosome of bacterial cells and very selectively inhibit protein synthesis. Some proteins are allowed to be made, but others are not,” said Alexander Mankin, the Alexander Neyfakh Professor of Medicinal Chemistry and Pharmacognosy at the UIC College of Pharmacy and senior author of the study. “Without these proteins being made, bacteria die.”
When people use antibiotics to treat an infection, the cells of the patient are not affected because the drugs are not designed to bind to the differently shaped ribosomes of eukaryotic cells.
“Because there are many human diseases caused by the expression of unwanted proteins — this is common in many types of cancer or neurodegenerative diseases, for example — we wanted to know if it would be possible to use an antibiotic to stop a human cell from making the unwanted proteins, and only the unwanted proteins,” Mankin said.
To answer this question, Mankin and study first author Maxim Svetlov, research assistant professor with the department of pharmaceutical sciences, looked to yeast, a eukaryote with cells similar to human cells.

The research team, which included partners from Germany and Switzerland, performed a “cool trick,” Mankin said. “We engineered the yeast ribosome to be more bacteria-like.”
Unlocking Ribosomal Control Through Genetics
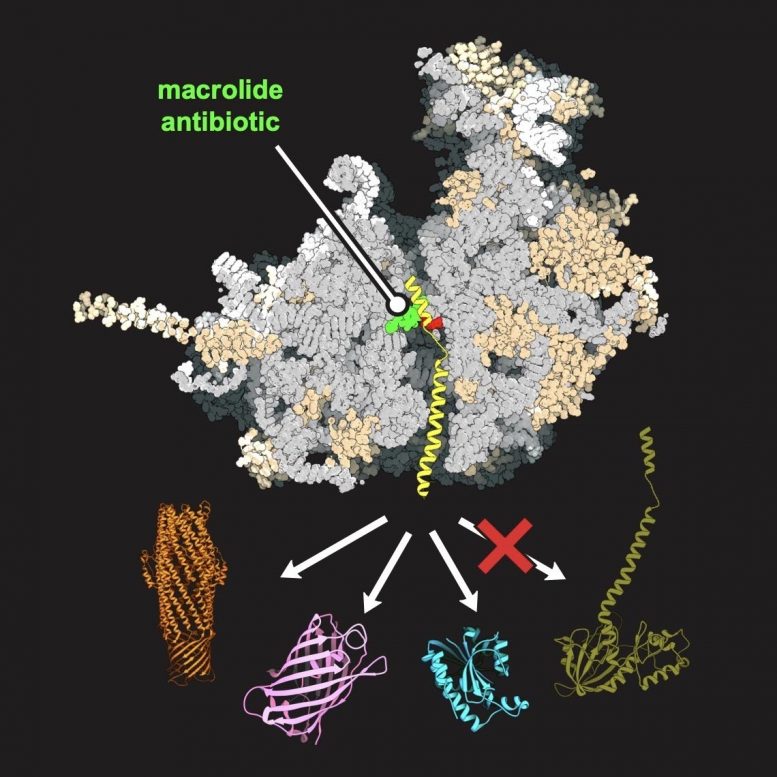
Mankin and Svetlov’s team used biochemistry and fine genetics to change one nucleotide of more than 7,000 in yeast ribosomal RNA, which was enough to make a macrolide antibiotic — a common class of antibiotics that works by binding to bacterial ribosomes — act on the yeast ribosome. Using this yeast model, the researchers applied genomic profiling and high-resolution structural analysis to understand how every protein in the cell is synthesized and how the macrolide interacts with the yeast ribosome.
“Through this analysis, we understood that depending on a protein’s specific genetic signature — the presence of a ‘good’ or ‘bad’ sequence — the macrolide can stop its production on the eukaryotic ribosome or not,” Mankin said. “This showed us, conceptually, that antibiotics can be used to selectively inhibit protein synthesis in human cells and used to treat human disorders caused by ‘bad’ proteins.”
The experiments of the UIC researchers provide a staging ground for further studies. “Now that we know the concepts work, we can look for antibiotics that are capable of binding in the unmodified eukaryotic ribosomes and optimize them to inhibit only those proteins that are bad for a human,” Mankin said.
Reference: “Context-specific action of macrolide antibiotics on the eukaryotic ribosome” by Maxim S. Svetlov, Timm O. Koller, Sezen Meydan, Vaishnavi Shankar, Dorota Klepacki, Norbert Polacek, Nicholas R. Guydosh, Nora Vázquez-Laslop, Daniel N. Wilson and Alexander S. Mankin, 14 May 2021, Nature Communications.
DOI: 10.1038/s41467-021-23068-1
Additional co-authors of the study are Dorota Klepacki and Nora Vázquez-Laslop of UIC; Timm Koller and Daniel Wilson of the University of Hamburg; Sezen Meydan and Nicholas Guydosh of the National Institutes of Health; and Norbert Polacek and Vaishnavi Shankar of the University of Bern.
This work was supported by grants from the National Institutes of Health (R35 GM127134, DK075132, 1FI2GM137845), the German Research Foundation (WI3285/6-1), and the Swiss National Science Foundation (31003A_166527).
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.