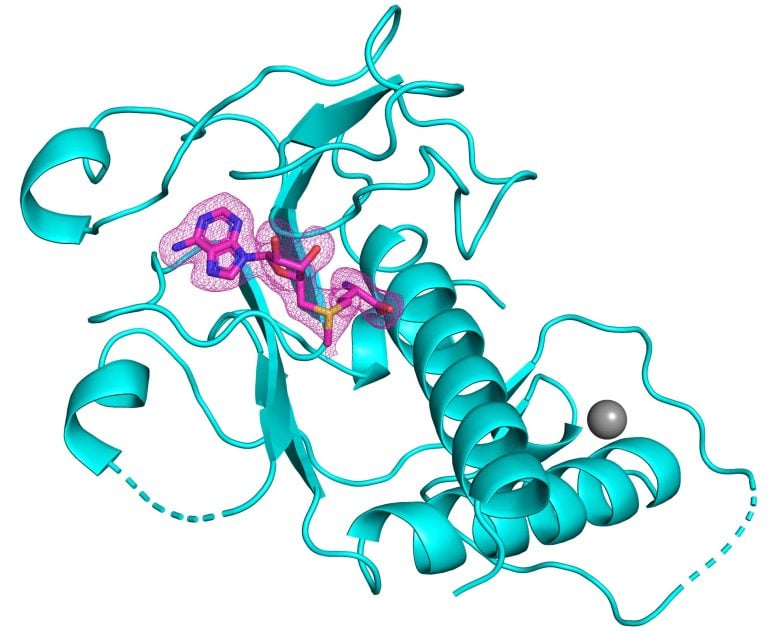
Researchers unravel the crystal structure of a key enzyme of SARS-CoV-2, paving the way for new antivirals.
A high-resolution crystal structure of an enzyme essential to the survival of SARS-CoV-2, the virus that causes COVID-19 has been produced by a team of Mount Sinai researchers. The discovery could help in the development of urgently needed new antivirals to combat current and future coronaviruses.
The enzyme, known as nsp14, contains a critically important region known as the RNA methyltransferase domain. This region has eluded previous attempts by the scientific community to characterize its three-dimensional crystal structure. A paper published in the September 8 online edition of Nature Structural & Molecular Biology describes the innovative process.
“Being able to visualize the shape of the methyltransferase domain of nsp14 at high resolution gives us insights into how to design small molecules that fit into its active site, and thus inhibit its essential chemistry,” says senior author Aneel Aggarwal, PhD. He is Professor of Pharmacological Sciences at the Icahn School of Medicine at Mount Sinai. “With this structural information, and in collaboration with medicinal chemists and virologists, we can now design small molecule inhibitors to add to the family of antivirals that go hand-in-hand with vaccines to combat SARS-CoV-2.”
Antiviral Research and the Importance of Multiple Inhibitors
Prescription antivirals that target key enzymes of SARS-CoV-2 include nirmatrelvir for the main protease (MPro) enzyme, and molnupiravir and remdesivir for the RNA polymerase (nsp12) enzyme. Research to develop new antivirals targeting different enzymatic activities has been accelerating in laboratories around the world, and Mount Sinai’s discovery has added significantly to that effort.
“Part of what drives our work,” says Dr. Aggarwal, “is the knowledge gained from treating HIV—that you typically need a cocktail of inhibitors for maximum impact against the virus.”
The Mount Sinai research team actually developed three crystal structures of nsp14, each with different cofactors. From these, they identified the best scaffold for the design of antivirals for inhibiting the RNA methyltransferase activity that the enzyme enables and the virus needs to survive. According to their scheme, the antiviral would take the place of the natural cofactor S-adenosylmethionine, therefore preventing the methyltransferase chemistry from occurring. The crystal structures that the researchers have elucidated have been made available to the public. They can now serve as guides for biochemists and virologists globally to engineer these compounds.
Fusion-Assisted Crystallization: A Breakthrough in nsp14 Research
Making the discovery possible was the ability of scientists to clear a hurdle that had prevented others in the past from creating three-dimensional crystals of the nsp14 methytransferase domain. “We employed an approach known as fusion-assisted crystallization,” explains lead author Jithesh Kottur, PhD. He is a postdoctoral fellow at Icahn Mount Sinai, and a crystallographer and biochemist. “It involves fusing the enzyme with another small protein that helps it to crystalize.”
Dr. Aggarwal is an internationally recognized structural biologist. He underscores the importance of ongoing investigative work by researchers in his field against a virus that has led to millions of deaths globally. “The virus evolves so quickly that it can develop resistance to the antivirals now available, which is why we need to continue developing new ones,” he observes. “Because of the high sequence conservation of nsp14 across coronaviruses and their variants (meaning it does not mutate much), our study will aid in the design of broad-spectrum antivirals for both present and future coronavirus outbreaks.”
Reference: “High resolution structures of the SARS-CoV-2 N7-methyltransferase inform therapeutic development” by Jithesh Kottur, Olga Rechkoblit, Richard Quintana-Feliciano, Daniela Sciaky and Aneel K. Aggarwal, 8 September 2022, Nature Structural & Molecular Biology.
DOI: 10.1038/s41594-022-00828-1
Funding: NIH/National Institutes of Health, DOE/US Department of Energy, NIH/National Institute of General Medical Sciences, DOE/US Department of Energy
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
1 Comment
Statistically, here in the US the reported number of Covid-19 deaths in 30 months is roughly equal to the projected number of deaths due to medical errors (e.g. Johns Hopkins researchers in May of 2016) during the same time period. Now age 78, as I’ve written many times before in many places in various ways, the three leading causes of premature mortality in the US are 1) medically unrecognized/undiagnosed long-term chronic subclinical non-IgE-mediated food (minimally) allergy reactions (e.g. preexisting conditions and comorbidities; premature aging and preventable diseases), 2) FDA approved food poisoning (namely added MSG and soy; others) and 3) ignorance and incompetence related medical errors. Any additional funding/research should be in areas of immediately cost effective and life saving prevention, not in areas of costly perpetual illness, premature demise and unlikely biochemical and/or genetic cures.