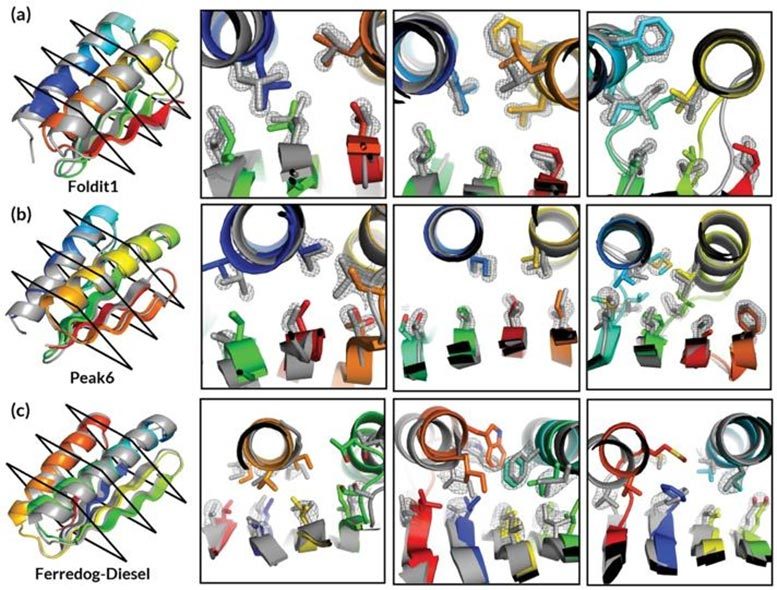
The unique ways in which proteins fold dictate their interplay with diseases and medicines, so understanding their twists and turns is key to designing effective drugs.
While new drug design is serious work, discovering how proteins fold can be fun, too: A crowdsourcing game called Foldit allows players to try different fold configurations for points and rankings. In a recent challenge, Foldit players came up with 56 designs that were reproduced as stable structures in a lab.
And X-ray studies at Berkeley Lab’s Advanced Light Source verified that three of these proteins’ real-life structures match the player-generated models. A fourth protein design was verified using a technique known as nuclear magnetic resonance spectroscopy.
These players had little or no training in this field, though their models proved to be as realistic as those created by experts and sophisticated computer programs. Their successes were highlighted in a scientific letter published in the journal Nature.
Now, the University of Washington researchers behind Foldit hope to introduce new challenges that could lead players to design actual drugs that work better than those now available. These efforts could benefit the growing field of biopharmaceuticals in which copies of existing proteins and modified versions of them are used to combat cancer and other diseases.
“Now that we’ve seen gamers can accurately design folded proteins, we’d like to challenge the gamers with more difficult design problems,” said Brian Koepnick, a research scientist at the University of Washington and Foldit developer.
Reference: “De novo protein design by citizen scientists” by Brian Koepnick, Jeff Flatten, Tamir Husain, Alex Ford, Daniel-Adriano Silva, Matthew J. Bick, Aaron Bauer, Gaohua Liu, Yojiro Ishida, Alexander Boykov, Roger D. Estep, Susan Kleinfelter, Toke Nørgård-Solano, Linda Wei, Foldit Players, Gaetano T. Montelione, Frank DiMaio, Zoran Popović, Firas Khatib, Seth Cooper and David Baker, 5 June 2019, Nature.
DOI: 10.1038/s41586-019-1274-4
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.