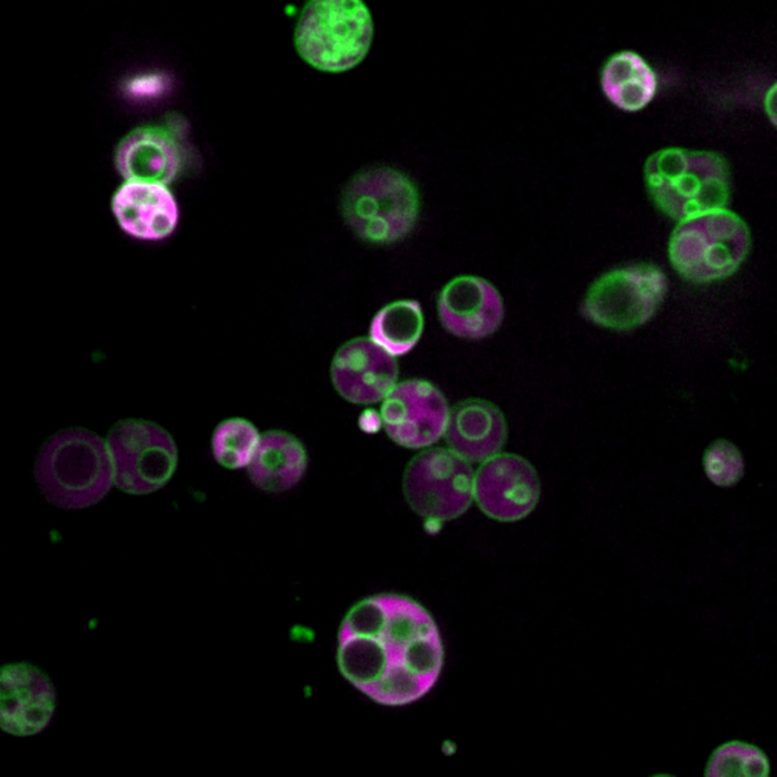
Newly discovered peroxisome subcompartments may enhance fat processing and reshape our understanding of cell metabolism and related diseases.
Discovery “requires us to rethink everything we thought we knew about peroxisomes.”
In his first year of graduate school, Rice University biochemist Zachary Wright discovered something hidden inside a common piece of cellular machinery that’s essential for all higher-order life from yeast to humans.
What Wright saw in 2015 — subcompartments inside organelles called peroxisomes — is described in a study published today in Nature Communications.
“This is, without a doubt, the most unexpected thing our lab has ever discovered,” said study co-author Bonnie Bartel, Wright’s Ph.D. advisor and a member of the National Academy of Sciences. “This requires us to rethink everything we thought we knew about peroxisomes.”
Peroxisomes are compartments where cells turn fatty molecules into energy and useful materials, like the myelin sheaths that protect nerve cells. In humans, peroxisome dysfunction has been linked to severe metabolic disorders, and peroxisomes may have wider significance for neurodegeneration, obesity, cancer, and age-related disorders.
Much is still unknown about peroxisomes, but their basic structure — a granular matrix surrounded by a sacklike membrane — wasn’t in question in 2015. Bartel said that’s one reason Wright’s discovery was surprising.

Fluorescent Imaging Unveils Hidden Structures
“We’re geneticists, so we’re used to unexpected things. But usually they don’t come in Technicolor,” she said, referring to another surprising thing about Wright’s find: beautiful color images that show both the walls of the peroxisome subcompartments and their interiors. The images were possible because of bright fluorescent reporters, glowing protein tags that Wright employed for the experiments. Biochemists modify the genes of model organisms — Bartel’s lab uses Arabidopsis plants — to tag them with fluorescent proteins in a controlled way that can reveal clues about the function and dysfunction of specific genes, including some that cause diseases in people, animals, and plants.
Wright, now a postdoctoral research associate in Bartel’s lab, was testing a new reporter in 2015 when he spotted the peroxisome subcompartments.
“I never thought Zach did anything wrong, but I didn’t think it was real,” Bartel said. She thought the images must be the result of some sort of artifact, a feature that didn’t really exist inside the cell but was instead created by the experiment.
“If this was really happening, somebody would have already noticed it,” she recalled thinking.

“Basically, from that point on, I was trying to understand them,” Wright said. He checked his instruments, replicated his experiments and found no evidence of an artifact. He gathered more evidence of the mysterious subcompartments, and eventually wound up at Fondren Library, combing through old studies.
Clues in Forgotten Studies from the 1960s
“I revisited the really old literature about peroxisomes from the ’60s, and saw that they had observed similar things and just didn’t understand them,” he said. “And that idea was just lost.”
There were a number of references to these inner compartments in studies from the ’60s and early ’70s. In each case, the investigators were focused on something else and mentioned the observation in passing. And all the observations were made with transmission electron microscopes, which fell out of favor when confocal microscopy became widely available in the 1980s.
“It’s just much easier than electron microscopy,” Bartel said. “The whole field started doing confocal microscopy. And in the early days of confocal microscopy, the proteins just weren’t that bright.”
Wright was also using confocal microscopy in 2015, but with brighter reporters that made it easier to resolve small features. Another key: He was looking at peroxisomes from Arabidopsis seedlings.
“One reason this was forgotten is because peroxisomes in yeast and mammalian cells are smaller than the resolution of light,” Wright said. “With fluorescence microscopy, you could only ever see a dot. That’s just the limit that light can do.”
Arabidopsis Seedlings Provide a Unique Window
The peroxisomes he was viewing were up to 100 times larger. Scientists aren’t certain why peroxisomes get so large in Arabidopsis seedlings, but they do know that germinating Arabidopsis seeds get all of their energy from stored fat, until the seedling leaves can start producing energy from photosynthesis. During germination, they are sustained by countless tiny droplets of oil, and their peroxisomes must work overtime to process the oil. When they do, they grow several times larger than normal.
“Bright fluorescent proteins, in combination with much bigger peroxisomes in Arabidopsis, made it extremely apparent, and much easier, to see this,” Wright said.
But peroxisomes are also highly conserved, from plants to yeast to humans, and Bartel said there are hints that these structures may be general features of peroxisomes.
“Peroxisomes are a basic organelle that has been with eukaryotes for a very long time, and there have been observations across eukaryotes, often in particular mutants, where the peroxisomes are either bigger or less packed with proteins, and thus easier to visualize,” she said. But people didn’t necessarily pay attention to those observations because the enlarged peroxisomes resulted from known mutations.
Subcompartments Aid Fat Metabolism
The researchers aren’t sure what purpose is served by the subcompartments, but Wright has a hypothesis.
“When you’re talking about things like beta-oxidation, or metabolism of fats, you get to the point that the molecules don’t want to be in water anymore,” Wright said. “When you think of a traditional kind of biochemical reaction, we just have a substrate floating around in the water environment of a cell — the lumen — and interacting with enzymes; that doesn’t work so well if you’ve got something that doesn’t want to hang around in the water.”
“So, if you’re using these membranes to solubilize the water-insoluble metabolites, and allow better access to lumenal enzymes, it may represent a general strategy to more efficiently deal with that kind of metabolism,” he said.
Bartel said the discovery also provides a new context for understanding peroxisomal disorders.
“This work could give us a way to understand some of the symptoms, and potentially to investigate the biochemistry that’s causing them,” she said.
Reference: “Peroxisomes form intralumenal vesicles with roles in fatty acid catabolism and protein compartmentalization in Arabidopsis” by Zachary J. Wright and Bonnie Bartel, 4 December 2020, Nature Communications.
DOI: 10.1038/s41467-020-20099-y
Bartel is the Ralph and Dorothy Looney Professor of BioSciences at Rice. The research was supported by the National Institutes of Health (R01GM079177, R35GM130338, S10RR026399) and the Welch Foundation (C-1309).
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
4 Comments
More lies
Open Access work for my TEM lens, TENS massage for my SERS extents.
It would be nice if they get to the point of the article, like why I should care?
Reading this article, it made me think… Hey! Can we separate/ isolate the peroxisomes from these seedlings and transfer them into a different system (yeast)? It sounds like the e.r. mechanisms just a bit. (I wouldn’t mind a few million metabolizing the ring around my waist lol.)