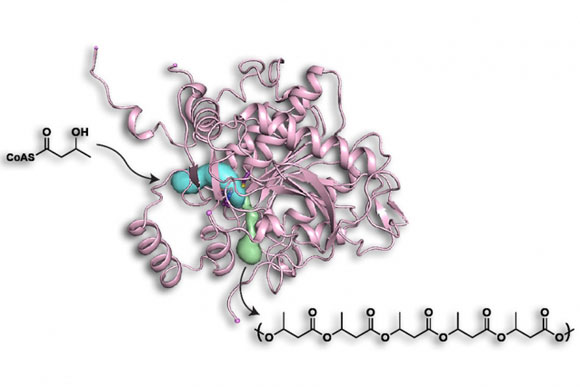
Researchers at MIT have determined the structure of a bacterial enzyme that can produce biodegradable plastics, an advance that could help chemical engineers tweak the enzyme to make it even more industrially useful.
The enzyme generates long polymer chains that can form either hard or soft plastics, depending on the starting materials that go into them. Learning more about the enzyme’s structure could help engineers control the polymers’ composition and size, a possible step toward commercial production of these plastics, which, unlike conventional plastic formed from petroleum products, should be biodegradable.
“I’m hoping that this structure will help people in thinking about a way that we can use this knowledge from nature to do something better for our planet,” says Catherine Drennan, an MIT professor of chemistry and biology and Howard Hughes Medical Institute Investigator. “I believe you want to have a good fundamental understanding of enzymes like this before you start engineering them.”
Drennan and JoAnne Stubbe, the Novartis Professor of Chemistry Emeritus and a professor emeritus of biology, are the senior authors of the study, which appears in the Journal of Biological Chemistry. The paper’s lead author is graduate student Elizabeth Wittenborn.
An elusive structure
The enzyme polyhydroxyalkanoate (PHA) synthase is found in nearly all bacteria, which use it to produce large polymers that store carbon when food is scarce. The bacterium Cupriavidus necator can store up to 85 percent of its dry weight as these polymers.
The enzyme produces different types of polymers depending on the starting material, usually one or more of the numerous variants of a molecule called hydroxyalkyl-coenzyme A, where the term alkyl refers to a variable chemical group that helps determine the polymers’ properties. Some of these materials form hard plastics, while others are softer and more flexible or have elastic properties that are more similar to rubber.
PHA synthase holds great interest for chemists and chemical engineers because it can string together up to 30,000 subunits, or monomers, in a precisely controlled way.
“What nature can do in this case and many others is make huge polymers, bigger than what humans can make,” Stubbe says. “And they have uniform molecular weight, which makes the properties of these polymers distinct.”
Drennan, Stubbe, and other chemists have been pursuing this enzyme’s structure for many years, but until now it has proven stubbornly elusive because of the difficulty in crystallizing the protein. Crystallization is a necessary step to performing X-ray crystallography, which reveals the atomic and molecular structure of the protein.
Two former graduate students, Marco Jost, and Yifeng Wei, who are also co-authors on the paper, worked on the crystallization as a side project and succeeded just before leaving MIT.
Once the researchers had the crystals, Wittenborn collected and analyzed the resulting crystallographic data to come up with the structure. The analysis revealed that PHA synthase consists of two identical subunits that form what is known as a dimer. Each subunit has an active site in which the polymerization occurs, thus eliminating an earlier proposal that the active site would be located at the dimer interface.
The analysis also revealed that the enzyme has two openings — one where the starting materials enter and another that allows the growing polymer chain to exit.
“The coenzyme A part of the substrate has to come back out because you have to put in another monomer,” Stubbe says. “There’s a lot of gymnastics that are going on, which I think makes this fascinating.”
The location of the entry channel was obvious as a gaping hole bordered by highly conserved amino acids, that is, amino acids that have remained constant as the enzyme has evolved. The exit channel was more difficult to identify because it is a much smaller opening, but the researchers were able to find it in part because it is also surrounded by conserved amino acids.
“The conserved residues form an arc-like network around the exit channel,” Wittenborn says. “They’re almost completely surrounding a very narrow portion of the channel, and we think they’re there to help secure the protein as the polymer starts to push its way through this tube.”
New framework
Drennan’s lab now plans to try to solve structures of the enzyme while it is bound to substrates and products, which should yield even more information critical to understanding how it works.
“This is the beginning of a new era of studying these systems where we now have this framework, and with every experiment we do, we’re going to be learning more,” Drennan says.
Some biotechnology companies have pursued making PHAs using PHA synthase and other enzymes needed to make the polymer, and one company is now using it to make polymers for medical use. Although for the most part the process is not cost-efficient enough to be economically competitive with low-cost conventional plastics derived from oil, the technology has enabled the production of unique PHA polymer compositions that can be used for specialty polymer additives, latex, and medical applications.
The new structural information yielded by this study will have little impact on cost but may open up the possibility of other new materials and applications, says Kristi Snell, the chief scientific officer and vice president of research at Yield10 Bioscience/Metabolix, which recently sold its PHA biopolymer technology to another company.
“The structure and mechanism of this enzyme has been a big question for over 20 years, and finding the structure could provide insight to help researchers make better polymers with unique properties,” Snell says.
Reference: “Structure of the Catalytic Domain of the Class I Polyhydroxybutyrate Synthase from Cupriavidus necator” by Elizabeth C. Wittenborn, Marco Jost, Yifeng Wei, JoAnne Stubbe and Catherine L. Drennan, November 2016, Journal of Biological Chemistry.
DOI: 10.1074/jbc.M116.756833
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.