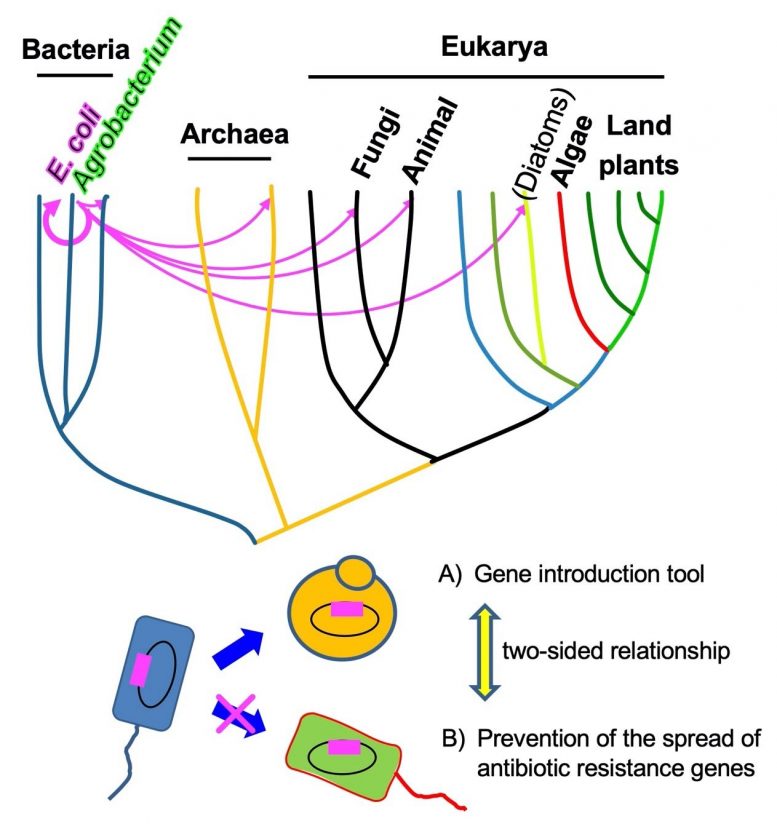
Mutant genes can promote genetic transfer across taxonomic kingdoms.
Bacteria do not sexually reproduce, but that does not stop them from exchanging genetic information as it evolves and adapts. During conjugal transfer, a bacterium can connect to another bacterium to pass along DNA and proteins. Escherichia coli bacteria, commonly called E. coli, can transfer at least one of these gene-containing plasmids to organisms across taxonomic kingdoms, including to fungi and protists. Now, researchers from Hiroshima University have a better understanding of this genetic hat trick, which has potential applications as a tool to promote desired characteristics or suppress harmful ones across genetic hosts.
They recently published their results in Frontiers in Microbiology.
Plasmids transfer from one bacterium — the donor — to another — the recipient. A particular kind of plasmid, called IncP1, can be hosted by a variety of bacteria and, seemingly as a result of its broad hosts, can transfer DNA to recipients beyond bacteria. The hypothesis is that the plasmid contains genes cultivated from different hosts and donors, resulting in this unique ability.
Donor Chromosome’s Role in Conjugation
“Although conjugation factors encoded on plasmids have been extensively analyzed, those on the donor chromosome have not,” said paper author Kazuki Moriguchi, associate professor, Program of Basic Biology, Graduate School of Integrated Sciences for Life, Hiroshima University.
There have been some studies on the various genes, according to Moriguchi, but the function of the genes was not examined, so it is not clear how they were related to the conjugation mechanism.
In this study, the researchers conducted a genome-wide survey on an extensive collection of bacteria mutants as donors to yeast. The mutants were engineered to have specific genes “knocked out” in order to study how the overall system performs without the presence of that specific gene, allowing researchers to infer information about the gene’s function.
“We focused on ‘up’ mutants that have the ability to accelerate conjugative transfer to both prokaryotes and eukaryotes as they could be potent donor strains applicable to gene introduction tools,” Moriguchi said, noting how IncP1’s ability to transmit genetic material across kingdoms could be used to develop precise tools to introduce genes capable of changing how the bacteria perform certain functions or react to changes in their environments.
Three Key Mutants and Their Mysterious Targets
Out of 3,884 mutants surveyed, three were identified that could conjugate across E. coli or from E. coli to yeast without accumulating genetic material, indicating that the genes worked together. The researchers analyzed the genes but were unable to elucidate the exact target or targets of conjugation mechanism that allows for cross-kingdom transfer. However, their analysis did reveal how the genes appear to work.
Two of the genes work to repress the unknown target in the E. coli donor. Simultaneously, the third gene is inactivated, allowing another unknown target to resume activity.
“The results suggest that the unknown target factors of these three genes form a complex in order to activate or repress the conjugation, either directly or indirectly at an identical step or steps of the IncP1 conjugation machinery, although the exact mechanism beyond this phenomenon remains unknown,” Moriguchi said.
According to Moriguchi, the data collected in this study can help facilitate the breeding of donor strains from various bacteria, each of which carries a high affinity with target organisms in addition to having a high conjugation ability.
Reference: “Isolation and Analysis of Donor Chromosomal Genes Whose Deficiency Is Responsible for Accelerating Bacterial and Trans-Kingdom Conjugations by IncP1 T4SS Machinery” by Fatin Iffah Rasyiqah Mohamad Zoolkefli, Kazuki Moriguchi, Yunjae Cho, Kazuya Kiyokawa, Shinji Yamamoto and Katsunori Suzuki, 20 May 2021, Frontiers in Microbiology.
DOI: 10.3389/fmicb.2021.620535
Co-authors include Fatin Iffah Rasyiqah Mohamad Zoolkefli, Shinji Yamamoto and Katsunori Suzuki, Department of Biological Science, Graduate School of Science; Suzuki and Kazuya Kiyokawa, Program of Basic Biology, Graduate School of Integrated Sciences for Life; and Yunjae Cho, Department of Biological Science, Faculty of Science.
The Japan Society for the Promotion of Science funded this work in part.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
1 Comment
… how would one see the mimicry explained in the world, after all…