
DNA sequencing is getting faster and cheaper, and clinicians are now finding uses for it in their practices. A test for malfunctioning genes might show how to treat a tumor or help diagnose the underlying cause of disease. However, raw sequencing data is too complex for most clinicians to directly analyze, and medical institutions are wary of transferring patient data to specialists located elsewhere.
A genome-interpretation company is now offering a hardware solution. Their meter-tall, 275-kilogram (606-pound) black box carries enough storage and processing power to analyze one genome every day, picking out mutations with potential links to diseases, and would be fast enough to inform clinicians for treatment. Some of the most important features of the $125,000-dollar unit is that it is a self-contained object. No patient data is transferred via networks to other places.
Knome, the company in Cambridge, Massachusetts which produces the device, has been approached by a number of companies who want to keep a system within their walls, not outsourcing data in any way. It was unveiled on the 27th of September, and the knoSYS 100 belongs to a class of emerging tools and services to help clinical researchers catch up with the latest advances in genome sequencing and apply it directly to their patients.
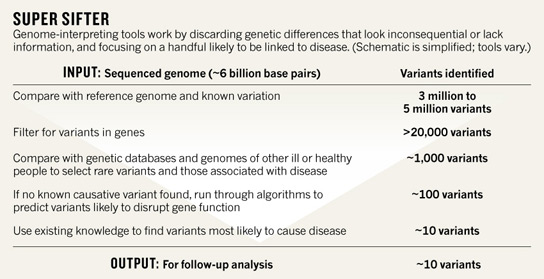
The core of the new Knome system is software, not hardware. It combs through a newly sequenced genome to try and find the places where the human genetic code varies. It annotates these places with existing knowledge. This process is known as genome interpretation and can quickly winnow down the millions of variants found in any individual to a handful that could explain the disease.
For now, the software isn’t meant for clinical diagnosis or medical advice. The clients include drug companies and medical centers researching how to use genome sequencing properly for clinical decisions.
Genome interpretation is far from mature, and its overall reliability depends largely on the quality of the sequences it analyzes. Not all variants can be detected and errors occur at every step of the process.
The protocols and the various annotation tools are still not perfect. There is the challenge of figuring out which detected variants are relevant to the disease in question. Any annotated variant must be tested before it can lead to a clear diagnosis. However, just organizing this information into a useful form is a big step forward, states George Church, genomicist at Harvard Medical School in Boston, Massachusetts, co-founder of Knome. The process is about delivering a high-quality interpretation that’s based on current knowledge, he continues.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.