
Dr. Eimear Kenny, a renowned professor of Medicine, Genetics, and Genomic Sciences at the Icahn School of Medicine at Mount Sinai, has made substantial contributions to the international Human Pangenome Reference Consortium, leading to the creation of a more inclusive human pangenome reference.
Eimear Kenny, PhD, had just completed undergrad and was working in her first computational genomics job more than 20 years ago when scientists announced the first (nearly) complete sequencing of the human genome—what was considered at the time to be the fundamental blueprint for all humans. The Human Genome Project aimed to map the entire genome in an effort to accelerate the diagnosis and eventual treatment of common and rare diseases.
Now, Dr. Kenny, a Professor of Medicine, and Genetics and Genomic Sciences, at the Icahn School of Medicine at Mount Sinai, can count herself as one of a handful of elite scientists worldwide whose vital contributions have led to the creation of the new human “pangenome” reference, a collection of genome sequences that captures significantly more human diversity.
Details on these novel developments were described in several Nature papers published last month. The work was led by the international Human Pangenome Reference Consortium, a group funded by the National Human Genome Research Institute (NHGRI), part of the National Institutes of Health. Dr. Kenny is a Principal Investigator and lead scientist of the consortium.
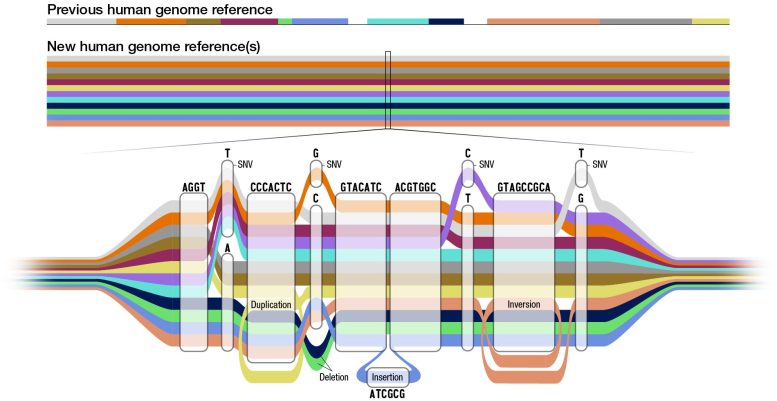
A genome is the set of DNA instructions that helps each living creature develop and function. Genome sequences differ slightly among individuals. In the case of humans, any two peoples’ genomes are, on average, more than 99 percent identical. The small differences contribute to each person’s uniqueness and can provide insights about their health, helping to diagnose disease, predict outcomes and guide medical treatments.
“We have had a single human reference for the past 20 years, and this genome reference has been extraordinarily powerful. It is a resource that has driven the sequencing of the genomes of tens or hundreds of millions of humans on the planet,” says Dr. Kenny, Professor of Medicine, and Genetics and Genomics Sciences at the Icahn School of Medicine at Mount Sinai, who is a co-author of the work. “However, it is limited in that most of the reference sequence only represents one person on the planet, so when you have rarer sequences or only in certain people, they are not represented. Therefore, we needed to really think about how to update the human reference and make it much more representative of diverse humans all over the world, which is what we have now done.”
Goals for the Pangenome Reference
The new pangenome reference includes genome sequences of 47 people, and the researchers aim to increase that number to 350 by mid-2024. Because each person carries a paired set of chromosomes, the current reference includes 94 distinct genome sequences, with a goal of reaching 700 distinct genome sequences by the completion of the project.
“Basic researchers and clinicians who use genomics need access to a reference sequence that reflects the remarkable diversity of the human population. This will help make the reference useful for all people, thereby helping to reduce the chances of propagating health disparities,” says Eric Green, MD, PhD, NHGRI director. “Creating and enhancing a human pangenome reference aligns with NHGRI’s goal of striving for global diversity in all aspects of genomics research, which is crucial to advance genomic knowledge and implement genomic medicine in an equitable way.”
Dr. Kenny, who is also Founding Director of the Institute for Genomic Health at Icahn Mount Sinai, leads research at the interface of genomics, medicine, and computer science to accelerate the use of genomics information in routine clinical care to improve human health.
She uses machine learning approaches and massive-scale databases of genomic information for discovery of novel genetic variants impacting disease risk. She also oversees large clinical trials in on implementing genomic medicine in diverse populations.
That expertise has paid off.
“Across many individuals, my role in this consortium was to contribute to this international scientific effort, and, in particular, help select the genomes that make up the new pangenome reference so that this resource would best benefit many people around the planet,” she says.
Population Genetics and Community Engagement
Dr. Kenny co-led a team using population genetics approaches, community engagement, and outreach to include genomes from diverse populations in the pangenome. This will help address issues of underrepresentation and bias in genomics research, and can improve the accuracy and generalizability of research findings across different populations.
The Human Genome Project completed in 2003 covered about 92 percent of the total human genome sequence. Recent technological advances such as long-read DNA sequencing, which reads longer stretches of the DNA at a time, helped researchers fill in those gaps to create the first complete human genome sequence. The developments were reported in a set of six papers in the April 1, 2022, issue of Science, along with companion papers published in several other journals. These findings were incorporated into the current pangenome reference.
“I’m delighted about the advancements in genomics technology that we have today. This new era of long-read sequencing, along with other capabilities, allows us to get much higher resolution of genomic sequences and, in particular, more accurately identify larger genomic variants called structural variants, which have been until now very difficult to detect with the short-read technology. This has enabled us to accelerate the rate at which we can find medically relevant variants and dramatically reduce sequencing costs,” says Dr. Kenny.
Importantly, knowing these variants better, Dr. Kenny says, will help elucidate which genes are truly rare or whether they may just be more common in certain parts of the world.
“The other significant aspect is that we are really trying to make a resource that is truly working toward global representativeness. We need to have a path toward recognizing that humans everywhere on the planet need resources available to them that best work for them,” says Dr. Kenny.
For more on this breakthrough, see:
- Human Pangenome Reference: A Deeper Understanding of Worldwide Genomic Diversity
- A Crystal Clear Image of Human Genomic Diversity
- Release of the New Human Pangenome Reference
“A draft human pangenome reference” by Wen-Wei Liao, Mobin Asri, Jana Ebler, Daniel Doerr, Marina Haukness, Glenn Hickey, Shuangjia Lu, Julian K. Lucas, Jean Monlong, Haley J. Abel, Silvia Buonaiuto, Xian H. Chang, Haoyu Cheng, Justin Chu, Vincenza Colonna, Jordan M. Eizenga, Xiaowen Feng, Christian Fischer, Robert S. Fulton, Shilpa Garg, Cristian Groza, Andrea Guarracino, William T. Harvey, Simon Heumos, Kerstin Howe, Miten Jain, Tsung-Yu Lu, Charles Markello, Fergal J. Martin, Matthew W. Mitchell, Katherine M. Munson, Moses Njagi Mwaniki, Adam M. Novak, Hugh E. Olsen, Trevor Pesout, David Porubsky, Pjotr Prins, Jonas A. Sibbesen, Jouni Sirén, Chad Tomlinson, Flavia Villani, Mitchell R. Vollger, Lucinda L. Antonacci-Fulton, Gunjan Baid, Carl A. Baker, Anastasiya Belyaeva, Konstantinos Billis, Andrew Carroll, Pi-Chuan Chang, Sarah Cody, Daniel E. Cook, Robert M. Cook-Deegan, Omar E. Cornejo, Mark Diekhans, Peter Ebert, Susan Fairley, Olivier Fedrigo, Adam L. Felsenfeld, Giulio Formenti, Adam Frankish, Yan Gao, Nanibaa’ A. Garrison, Carlos Garcia Giron, Richard E. Green, Leanne Haggerty, Kendra Hoekzema, Thibaut Hourlier, Hanlee P. Ji, Eimear E. Kenny, Barbara A. Koenig, Alexey Kolesnikov, Jan O. Korbel, Jennifer Kordosky, Sergey Koren, HoJoon Lee, Alexandra P. Lewis, Hugo Magalhães, Santiago Marco-Sola, Pierre Marijon, Ann McCartney, Jennifer McDaniel, Jacquelyn Mountcastle, Maria Nattestad, Sergey Nurk, Nathan D. Olson, Alice B. Popejoy, Daniela Puiu, Mikko Rautiainen, Allison A. Regier, Arang Rhie, Samuel Sacco, Ashley D. Sanders, Valerie A. Schneider, Baergen I. Schultz, Kishwar Shafin, Michael W. Smith, Heidi J. Sofia, Ahmad N. Abou Tayoun, Françoise Thibaud-Nissen, Francesca Floriana Tricomi, Justin Wagner, Brian Walenz, Jonathan M. D. Wood, Aleksey V. Zimin, Guillaume Bourque, Mark J. P. Chaisson, Paul Flicek, Adam M. Phillippy, Justin M. Zook, Evan E. Eichler, David Haussler, Ting Wang, Erich D. Jarvis, Karen H. Miga, Erik Garrison, Tobias Marschall, Ira M. Hall, Heng Li and Benedict Paten, 10 May 2023, Nature.
DOI: 10.1038/s41586-023-05896-x
“Increased mutation rate and gene conversion within human segmental duplications” by Mitchell R. Vollger, Philip C. Dishuck, William T. Harvey, William S. DeWitt, Xavi Guitart, Michael E. Goldberg, Allison N. Rozanski, Julian Lucas, Mobin Asri, Human Pangenome Reference Consortium, Katherine M. Munson, Alexandra P. Lewis, Kendra Hoekzema, Glennis A. Logsdon, David Porubsky, Benedict Paten, Kelley Harris, PingHsun Hsieh and Evan E. Eichler, 10 May 2023. Nature.
DOI: 10.1038/s41586-023-05895-y
“Recombination between heterologous human acrocentric chromosomes” by Andrea Guarracino, Silvia Buonaiuto, Leonardo Gomes de Lima, Tamara Potapova, Arang Rhie, Sergey Koren, Boris Rubinstein, Christian Fischer, Human Pangenome Reference Consortium, Jennifer L. Gerton, Adam M. Phillippy, Vincenza Colonna and Erik Garrison, 10 May 2023, Nature.
DOI: 10.1038/s41586-023-05976-y
“Pangenome graph construction from genome alignment with minigraph-cactus” by Glenn Hickey, Jean Monlong, Jana Ebler, Adam M. Novak, Jordan M. Eizenga, Yan Gao, Human Pangenome Reference Consortium, Tobias Marschall, Heng Li and Benedict Paten, 10 May 2023, Nature Biotechnology.
DOI: 10.1038/s41587-023-01793-w
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.