Scientists at the University of California, Santa Cruz, together with a consortium of researchers, have released a draft of the first human pangenome, which combines the genetic material of 47 individuals from diverse ancestral backgrounds to offer a more accurate representation of global genomic diversity. This new pangenome adds 119 million DNA bases to the existing reference genome, increasing the detection of variants in the human genome. The pangenome was produced by the Human Pangenome Reference Consortium and is available on the UCSC Genome Browser. The project is expected to continue until 2024, when a final pangenome with genomic information from 350 individuals is planned for release.
University of California, Santa Cruz scientists, along with a consortium of researchers, have released a draft of the first human pangenome—a new, usable reference for genomics that combines the genetic material of 47 individuals from different ancestral backgrounds to allow for a deeper, more accurate understanding of worldwide genomic diversity.
By adding 119 million bases—the “letters” in DNA sequences—to the existing genomics reference, the pangenome provides a representation of human genetic diversity that was not possible with a single reference genome. It is highly accurate, more complete and dramatically increases the detection of variants in the human genome, as shown in a collection of groundbreaking papers published today (May 10, 2023) in the journals Nature, Genome Research, Nature Biotechnology, and Nature Methods.
The pangenome was produced by the Human Pangenome Reference Consortium (HPRC), which is co-led by UCSC’s Associate Professor of Biomolecular Engineering Benedict Paten and Assistant Professor of Biomolecular Engineering Karen Miga and is now available for use in an assembly hub on the UCSC Genome Browser. More than a dozen UCSC researchers and students are contributors to this project, which will continue into 2024 when the researchers plan to release a final pangenome with genomic information from 350 individuals.
“We are introducing more diversity and equity into the reference by sampling diverse human beings and including them in this structure that everyone can use,” said Paten, who is the senior author on the main marker paper. “One genome isn’t enough to represent everybody—the pangenome will ultimately be something that is inclusive and representative.”
Understanding Genomic Variation
Each person’s genome varies slightly—by about 0.4 percent compared to the next person, on average—and understanding these differences can provide insight into their health, help to diagnose disease, predict medical outcomes, and guide treatments. Using the pangenome reference will improve scientists’ ability to detect and understand variation in future studies.
Typically when scientists and clinicians study an individual’s genome to look for variation, they compare that individuals’ DNA to that of a standard reference to determine where there are differences of one or more base pairs. Until now, the reference genome has primarily been represented by a single sequence for each human chromosome, mostly sourced from one individual. But, this reference is nearly 20 years old and fundamentally limited in that it can not represent the wealth of genetic variations present in the human population. This introduces an issue called reference bias into genome analysis.
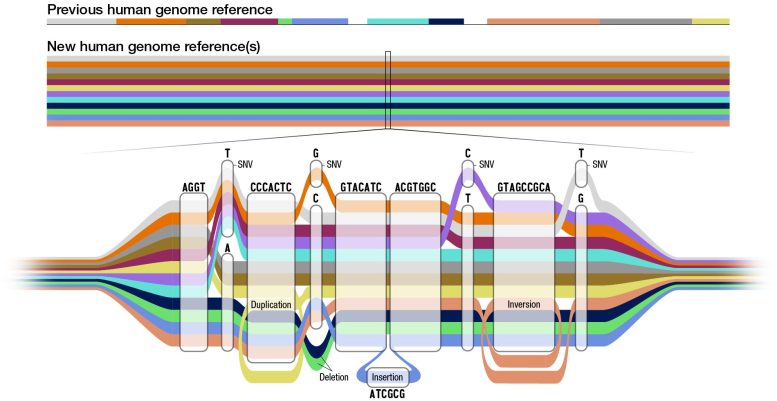
In contrast, the new pangenome is a reference that combines the genomes of 47 individuals from various ancestral backgrounds. The pangenome looks like a linear reference in areas where the sequences have the same bases, and expands to show the areas where there are differences. It represents many different versions of the human genome sequence at the same time, and gives scientists a more accurate point of comparison for variation that is present in some populations but not others.
“One genome can’t possibly represent all of the rich variation we know can be observed and studied around the world,” said Miga, Director of the HPRC Production Center at UCSC. “The No. 1 goal of the human pangenome reference is to try to broaden the representation of a reference resource to be more inclusive and more equitable for studying the human species, as a collection of references and not just one.”
Genomic variation can be small, consisting of differences of just one or a few DNA bases, or it can be large structural variants, classified as variants that are 50 base pairs or larger. These larger, structural variants can have important health implications. Until now, researchers have been unable to identify more than 70 percent of the structural variants that exist in human genomes due to limited technologies and the bias of using a single reference sequence.
Of the 119 million new bases added to the reference with the pangenome, roughly 90 million of these derive from structural variation. Structural variants are complex and may be inversions of sequences, insertions, deletions, or tandem repeats—a segment of two or more bases repeated numerous times. These new bases will help researchers to study regions in the genome for which there was previously no reference, and potentially be able to associate structural variants with disease in future studies.
“Now, we can map to more structural variants, so we’re finding features and areas in the genome that just weren’t there before,” Miga said. “That’s exciting because it’s allowing us to look at gene regulation in a unique way that we couldn’t study before, because those areas probably would have been inappropriately mapped or just ignored altogether.”
Using the pangenome reference for genomic analysis increases the detection of structural variants by 104 percent as compared to detection using the standard reference. The pangenome reference also increases the accuracy of calling small variants, those just a few bases long, by about 34 percent because of the increased amount of data present in the pangenome.
Each human carries a paired set of chromosomes—one set inherited from the mother and one from the father. The individual genomes present in the pangenome reference contains haplotype-resolved information, meaning it can confidently distinguish the two parental sets of chromosomes—a major scientific feat. Having this information will help scientists better understand how various genes and diseases are inherited.
This also means the current reference actually includes 94 distinct genome sequences, with the goal of getting to 700 by 2024.
Creating the Pangenome
The pangenome was made possible through the development of advanced computational techniques to align the multiple genome sequences into one, usable reference in a structure called a pangenome graph. Paten and researchers in the UCSC Computational Genomics lab helped lead the HPRC efforts to develop the algorithmic methods needed to create this pangenome graph structure.
Because of the methods used in this project, all of the genomes within the pangenome reference are of extremely high quality and accuracy, covering more than 99 percent of each human genome with more than 99 percent accuracy.
“In the linear reference, we had only one sequence, one representation of each gene,” said Mobin Asri, a bioinformatics Ph.D. candidate at UCSC and co-first author on the main paper. “But we know that our genes have different variations in the human population. Using the pangenome graph, we want to have all of those variations in a single structure—and a graph is a natural way to do this.”
The HPRC project relies heavily on long- and ultra long-read sequencing technology to read DNA from biological samples. With recent advances, these techniques can now decode thousands to millions of base pairs of the genome at once. The long stretches of DNA reads are then assembled via specialized algorithms into more complete genomic sequences. Ideally each assembled sequence should represent the sequence of one chromosome.
Long reads contain errors about one percent of the time and current assembly algorithms are
not perfect, which can cause the assembled sequences to be erroneous in some locations. To check for and correct these errors, the individual genomes that have been sequenced and assembled move through multiple tools, including a reliability pipeline developed by Asri. Once having been processed by these tools, the researchers can ensure the assemblies are accurate and complete.
After moving through Asri’s pipeline, the various genomes are compiled via complex algorithmic methods into the pangenome graph structure. Visually, the graph genome allows researchers to view differences in the various reference sequences as diverging areas in otherwise shared paths.
Building an Accessible Resource
All of the first 47 diploid genomes in the draft pangenome were sourced from individuals who participated in the 1000 Genomes Project (1000G), an influential effort which created a catalog of common human genetic variation from openly consented samples and was completed in 2015. The open consent status of these samples allow any researcher to access the resource without the privacy barriers that typically accompany genome research, with the aim of making the pangenome accessible to as many people as possible.
“Becoming a common resource is something that’s fundamental to the success of a human pangenome reference,” Miga said. “It has to have the ability to be accessible and open around the world to all researchers so we can use it as the foundation.”
The HPRC team is focused on outreach to ensure that the pangenome is a useful resource that will be utilized in clinics around the world. This means facilitating annotations, feedback, and input from the researchers carrying out studies using the pangenome reference.
“The draft pangenome is an important proof of principle that we hope is going to influence a lot of people and get them thinking about the pangenome and how it might affect their work,” Paten said. “Looking ahead, we see a lot of engagement with other groups—it takes a lot of different people to build something that is going to become a big community resource.”
Along with a focus on accessibility, the HPRC project has a dedicated ethics team focused on the social and legal implications of this project. They are working to anticipate challenging issues and help guide informed consent, prioritize the study of different samples, explore possible regulatory issues pertaining to clinical adoption, and work with international and Indigenous communities to incorporate their genome sequences in these broader efforts.
Continuing the Legacy and Future Work
The human pangenome is a continuation of decades-long efforts from scientists at UC Santa Cruz to understand the biological code that underlies human life.
In 2000, Jim Kent, then a UCSC graduate student and now a research scientist at the Genomics Institute and director of the UCSC Genome Browser, wrote the code that assembled the first working draft of the human genome. UCSC scientists published it with open access to anyone who wanted to use it. Since then, UCSC has been at the forefront of genomics research.
In April 2022, UCSC’s Karen Miga co-led the Telomere-to-Telomere consortium to assemble the first complete sequencing of a human genome, filling in missing, complex regions of reference that had long eluded scientists.
“Since 2000, we’ve had a series of increasingly more accurate representations of one genome,” said David Haussler, Scientific Director of the UCSC Genomics Institute who led the UCSC team on the original Human Genome Project and advises on the pangenome project. “But no matter how accurately you represent one genome, that’s not going to represent all of humanity. Now is a turning point: no longer genomics of the one standard human genome, but genomics for everybody.”
The researchers are making progress toward the goal of completing the full pangenome by 2024. The team is in the process of recruiting new individuals to represent some populations not included in the 1000 Genomes Project, particularly people of Middle Eastern and African ancestry. Miga, as the director of the Data Production Center at UCSC, will spearhead these efforts going forward.
In addition to completing the final pangenome reference, the researchers are working toward forming an international human pangenome project that would establish partnerships with researchers across the world. These partnerships would include a two-way skills and knowledge exchange, aimed to bring the skills and technology needed to create high-quality reference genomes into the hands of researchers worldwide so they can carry out their own research.
References:
“A draft human pangenome reference” by Wen-Wei Liao, Mobin Asri, Jana Ebler, Daniel Doerr, Marina Haukness, Glenn Hickey, Shuangjia Lu, Julian K. Lucas, Jean Monlong, Haley J. Abel, Silvia Buonaiuto, Xian H. Chang, Haoyu Cheng, Justin Chu, Vincenza Colonna, Jordan M. Eizenga, Xiaowen Feng, Christian Fischer, Robert S. Fulton, Shilpa Garg, Cristian Groza, Andrea Guarracino, William T. Harvey, Simon Heumos, Kerstin Howe, Miten Jain, Tsung-Yu Lu, Charles Markello, Fergal J. Martin, Matthew W. Mitchell, Katherine M. Munson, Moses Njagi Mwaniki, Adam M. Novak, Hugh E. Olsen, Trevor Pesout, David Porubsky, Pjotr Prins, Jonas A. Sibbesen, Jouni Sirén, Chad Tomlinson, Flavia Villani, Mitchell R. Vollger, Lucinda L. Antonacci-Fulton, Gunjan Baid, Carl A. Baker, Anastasiya Belyaeva, Konstantinos Billis, Andrew Carroll, Pi-Chuan Chang, Sarah Cody, Daniel E. Cook, Robert M. Cook-Deegan, Omar E. Cornejo, Mark Diekhans, Peter Ebert, Susan Fairley, Olivier Fedrigo, Adam L. Felsenfeld, Giulio Formenti, Adam Frankish, Yan Gao, Nanibaa’ A. Garrison, Carlos Garcia Giron, Richard E. Green, Leanne Haggerty, Kendra Hoekzema, Thibaut Hourlier, Hanlee P. Ji, Eimear E. Kenny, Barbara A. Koenig, Alexey Kolesnikov, Jan O. Korbel, Jennifer Kordosky, Sergey Koren, HoJoon Lee, Alexandra P. Lewis, Hugo Magalhães, Santiago Marco-Sola, Pierre Marijon, Ann McCartney, Jennifer McDaniel, Jacquelyn Mountcastle, Maria Nattestad, Sergey Nurk, Nathan D. Olson, Alice B. Popejoy, Daniela Puiu, Mikko Rautiainen, Allison A. Regier, Arang Rhie, Samuel Sacco, Ashley D. Sanders, Valerie A. Schneider, Baergen I. Schultz, Kishwar Shafin, Michael W. Smith, Heidi J. Sofia, Ahmad N. Abou Tayoun, Françoise Thibaud-Nissen, Francesca Floriana Tricomi, Justin Wagner, Brian Walenz, Jonathan M. D. Wood, Aleksey V. Zimin, Guillaume Bourque, Mark J. P. Chaisson, Paul Flicek, Adam M. Phillippy, Justin M. Zook, Evan E. Eichler, David Haussler, Ting Wang, Erich D. Jarvis, Karen H. Miga, Erik Garrison, Tobias Marschall, Ira M. Hall, Heng Li and Benedict Paten, 10 May 2023, Nature.
DOI: 10.1038/s41586-023-05896-x
“Increased mutation rate and gene conversion within human segmental duplications” by Mitchell R. Vollger, Philip C. Dishuck, William T. Harvey, William S. DeWitt, Xavi Guitart, Michael E. Goldberg, Allison N. Rozanski, Julian Lucas, Mobin Asri, Human Pangenome Reference Consortium, Katherine M. Munson, Alexandra P. Lewis, Kendra Hoekzema, Glennis A. Logsdon, David Porubsky, Benedict Paten, Kelley Harris, PingHsun Hsieh and Evan E. Eichler, 10 May 2023. Nature.
DOI: 10.1038/s41586-023-05895-y
“Recombination between heterologous human acrocentric chromosomes” by Andrea Guarracino, Silvia Buonaiuto, Leonardo Gomes de Lima, Tamara Potapova, Arang Rhie, Sergey Koren, Boris Rubinstein, Christian Fischer, Human Pangenome Reference Consortium, Jennifer L. Gerton, Adam M. Phillippy, Vincenza Colonna and Erik Garrison, 10 May 2023, Nature.
DOI: 10.1038/s41586-023-05976-y
“Pangenome graph construction from genome alignment with minigraph-cactus” by Glenn Hickey, Jean Monlong, Jana Ebler, Adam M. Novak, Jordan M. Eizenga, Yan Gao, Human Pangenome Reference Consortium, Tobias Marschall, Heng Li and Benedict Paten, 10 May 2023, Nature Biotechnology.
DOI: 10.1038/s41587-023-01793-w
Other UCSC researchers on the main paper include Marina Haukness, Glenn Hickey, Julian Lucas, Jean Monlong, Xian Chang, Jordan Eizenga, Charles Markello, Adam Novak, Hugh Olsen, and Trevor Pesout.
Funding for the HPRC was primarily provided by the National Human Genome Research Institute.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.