Study investigates potential for the intestinal microbial community to alter drug safety and efficacy.
Researchers at Princeton University have developed a systematic approach for evaluating how the microbial community in our intestines can chemically transform, or metabolize, oral medications in ways that impact their safety and efficacy.
The new methodology provides a more complete picture of how gut bacteria metabolize drugs, and could aid the development of medications that are more effective, have fewer side effects, and are personalized to an individual’s microbiome.
The study was published today (June 10, 2020) in the journal Cell.
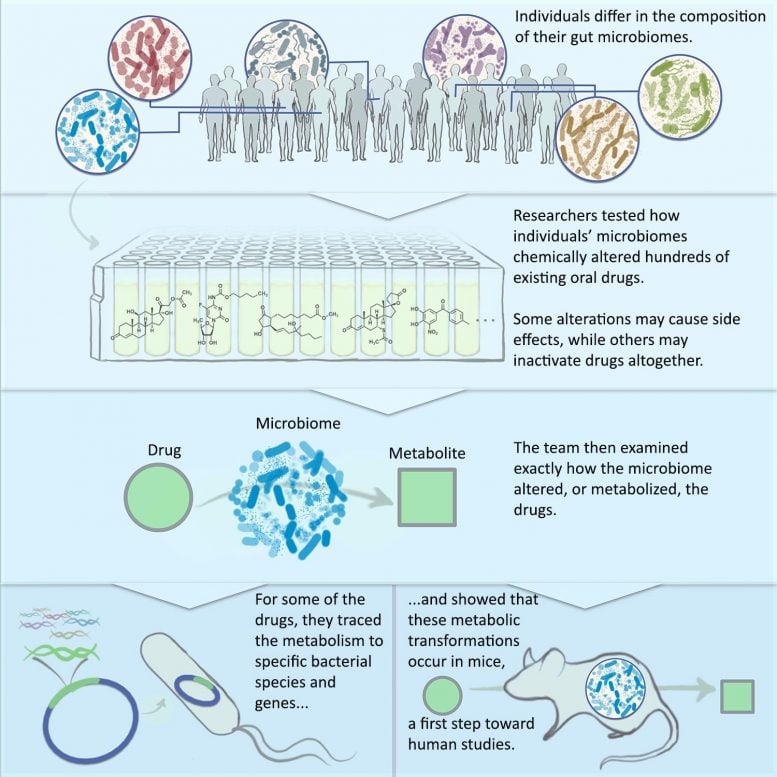
Previous studies have examined how single species of gut bacteria can metabolize oral medications. The new framework enables evaluation of a person’s entire intestinal microbial community at once.
“Basically, we do not run and hide from the complexity of the microbiome, but instead, we embrace it,” said Mohamed S. Donia, assistant professor of molecular biology. “This approach allows us to gain a holistic and more realistic view of the microbiome’s contribution to drug metabolism.”
The team used the approach to evaluate the gut microbiome’s effect on hundreds of common medications already on the market. The intestines are the primary region where pills and liquid medications are absorbed into the body.
The researchers identified 57 cases in which gut bacteria can alter existing oral medications. Eighty percent of those had not been previously reported, emphasizing the potential of the method for revealing unknown drug-microbiome interactions.
These alterations range from converting the medicine into an inactive state — which can reduce its efficacy — to converting the drug into a form that is toxic, potentially causing side effects.
The framework could aid drug discovery by identifying potential drug-microbiome interactions early in development, informing formulation changes. The approach can also help during clinical trials to better analyze the toxicity and efficacy of the drugs being tested.
The intestines are home to hundreds of species of bacteria. The makeup of these communities — what kinds of bacteria and how many of each species — can vary considerably from person to person.
“This inter-person variability underscores why studying a single bacterial species makes it impossible to compare the microbiome’s metabolism of drugs between individuals,” Donia said. “We need to study the entire intestinal microbial community.”
The researchers found that some people’s microbiomes had little effect on a given drug, while other microbiomes had a significant effect, demonstrating how important the community of bacteria — rather than just a single species — is on drug metabolism.
“Everyone’s microbiome is unique, and we were able to see this in our study,” said Bahar Javdan, an M.D.-Ph.D. student in molecular biology and a co-first author on the study. “We observed three main categories — drugs that were consistently metabolized by all the microbiomes in our study, drugs that were metabolized by some and not by others, and drugs that were not subject to any microbiome-derived metabolism.”
The methodological approach could be valuable for personalizing treatment to the microbiome of each patient. For example, the framework could help predict how a certain drug will behave, and suggest changes to the therapeutic strategy if undesired effects are predicted.
“This is a case where medicine and ecology collide,” said Jaime Lopez, a graduate student in the Lewis-Sigler Institute for Integrative Genomics and a co-first author on the study, who contributed the computational and quantitative analysis of the data. “The bacteria in these microbial communities help each other survive, and they influence each other’s enzymatic profiles. This is something you would never capture if you didn’t study it in a community.”
The framework involves four steps for systematically evaluating the intestinal microbiome’s effect on drugs.
First, the researchers collected 21 fecal samples collected from anonymous donors and cataloged the bacterial species living in each individual. They found that the donors each had a unique microbial community living in their guts, and that the majority of these personalized communities can be grown in a lab culturing system that they developed.
Next, they tested 575 FDA-approved drugs to see if they are chemically modified by one of the 21 cultured microbiomes, and then tested a subset of the drugs with all the cultured microbiomes. Here, they found microbiome-derived metabolites that had never been previously reported, as well as ones that have been reported in humans and associated with side effects but their origins were unknown. They found cases where all the donor microbiomes performed the same reactions on the drug, and others where only a subset did.
Then they examined the mechanisms by which some of the modified drugs are altered by the cultured microbiomes. To understand exactly how the transformations occurred, they traced the source of the chemical transformations to particular bacterial species and to particular genes within those bacteria. They also showed that the microbiome-derived metabolism reactions that are discovered in this manner can be recapitulated in a mouse model, the first step in adapting the approach for human drug development.
Reference: ” Personalized mapping of drug metabolism by the human gut microbiome” by Bahar Javdan, Jaime G. Lopez, Pranatchareeya Chankhamjon, Ying-Chiang J.Lee, Raphaella Hull, Qihao Wu, Xiaojuan Wang, Seema Chatterjee and Mohamed S. Donia, 10 June 10, 2020, Cell.
DOI: 10.1016/j.cell.2020.05.001
Additional contributors to the study were postdoctoral research associates Pranatchareeya Chankhamjon, Qihao Wu and Xiaojuan Wang; graduate student in molecular biology Ying-Chiang J. Lee, graduate student at Cambridge University Raphaella Hull; and Seema Chatterjee, research lab manager.
Funding for this study was provided by the Princeton Department of Molecular Biology, the National Institutes of Health, the New Jersey Commission on Cancer and the National Science Foundation.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
1 Comment
THE APPENDIX.
Your ‘vestigial’ dismissal
Is simply inferior
To His ‘nursery and dispensary’
For friendly bacteria.