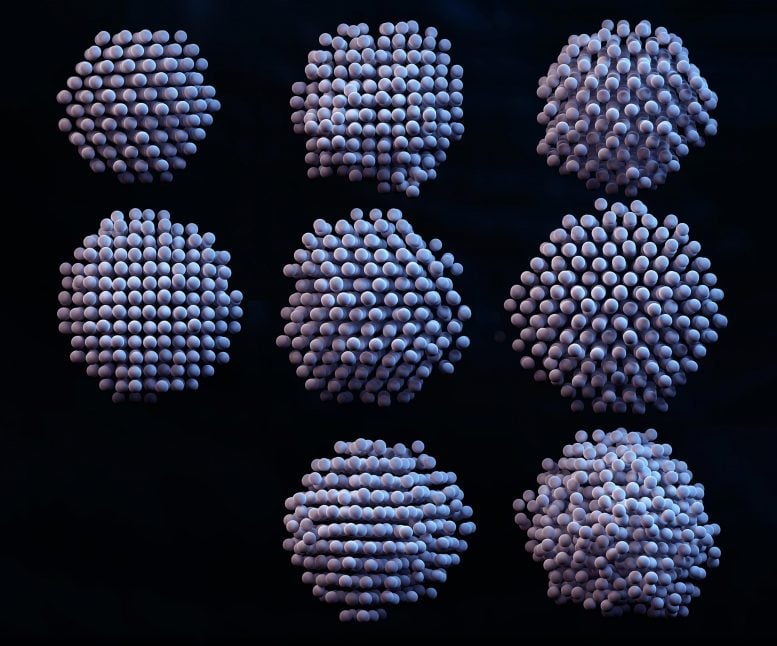

Liquid phase electron microscopy illuminates 3D atomic structures of platinum nanoparticles, advancing full control of nanoengineering.
What do you see in the picture above? Merely a precisely-drawn three-dimensional picture of nanoparticles? Far more than that, nanotechnologists will say, due to a new study published in the journal Science. Whether a material catalyzes chemical reactions or impedes any molecular response is all about how its atoms are arranged. The ultimate goal of nanotechnology is centered around the ability to design and build materials atom by atom, thus allowing scientists to control their properties in any given scenario. However, atomic imaging techniques have not been sufficient to determine the precise three-dimensional atomic arrangements of materials in liquid solution, which would tell scientists how materials behave in everyday life, such as in water or blood plasma.
3D density maps, atomic position maps, and strain maps of 8 reconstructed nanocrystals show critical differences between the individual particles.
Researchers at the Center for Nanoparticle Research within the Institute for Basic Science (IBS, South Korea), in collaboration with Dr. Hans Elmlund at Monash University’s Biomedicine Discovery Institute in Australia and Dr. Peter Ercius at Lawrence Berkeley National Laboratory’s Molecular Foundry in the USA, have reported a new analytic methodology that can resolve the 3D structure of individual nanoparticles with atomic-level resolution. The 3D atomic positions of individual nanoparticles can be extracted with a precision of 0.02 nm — six times smaller than the smallest atom: hydrogen. In other words, this high-resolution method detects individual atoms and how they are arranged within a nanoparticle.
The researchers call their development 3D SINGLE (Structure Identification of Nanoparticles by Graphene Liquid cell Electron microscopy) and utilize mathematical algorithms to derive 3D structures from a set of 2D imaging data acquired by one of the most powerful microscopes on Earth. First, a nanocrystal solution is sandwiched in-between two graphene sheets which are each just a single atom thick (image below). “If a fish bowl were made of a thick material, it would be hard to see through it. Since graphene is the thinnest and strongest material in the world, we created graphene pockets that allow the electron beam of the microscope to shine through the material while simultaneously sealing the liquid sample,” explains PARK Jungwon, one of the corresponding authors of the study (assistant professor at the School of Chemical and Biological Engineering in Seoul National University).
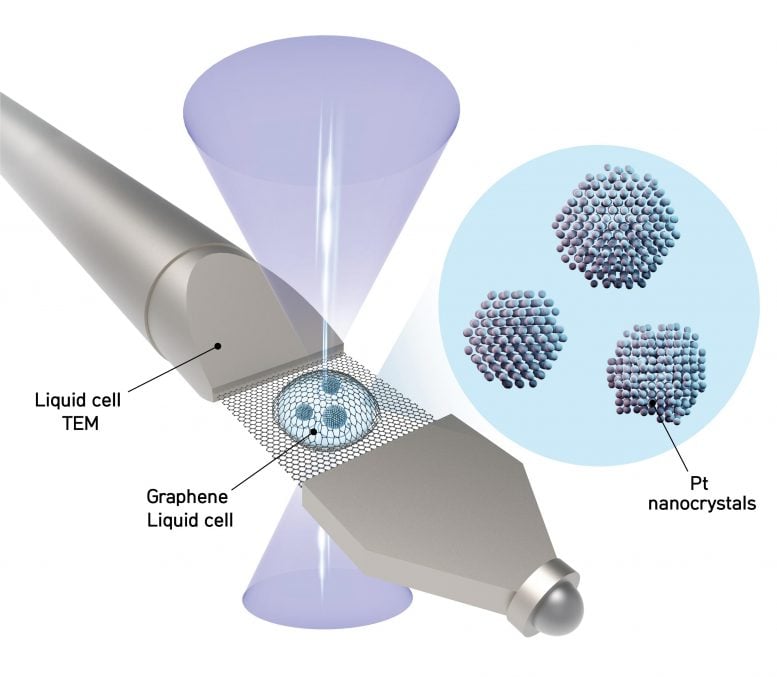
The researchers obtain movies at 400 images per second of each nanoparticle freely rotating in liquid using a high-resolution transmission electron microscope (TEM). The team then applies their reconstruction methodology to combine the 2D images into a 3D map showing the atomic arrangement. Locating the precise position of each atom tells researchers how the nanoparticle was created and how it will interact in chemical reactions.
The study defined the atomic structures of eight platinum nanoparticles – platinum is the most valuable of the precious metals, used in a number of applications such as catalytic materials for energy storage in fuel cells and petroleum refinement. Even though all of the particles were synthesized in the same batch, they displayed important differences in their atomic structures which affect their performance.
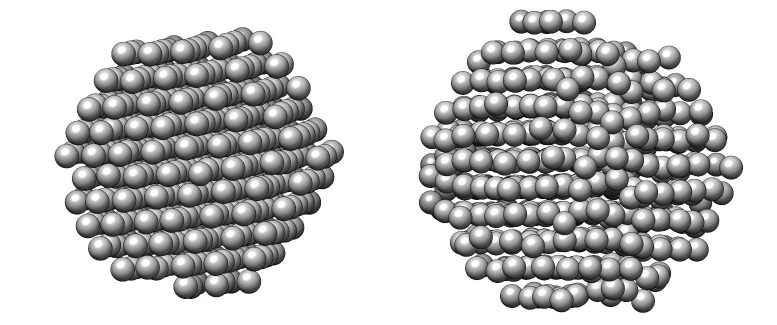
“Now it is possible to experimentally determine the precise 3D structures of nanomaterials that had only been theoretically speculated. The methodology we developed will contribute to fields where nanomaterials are used, such as fuel cells, hydrogen vehicles, and petrochemical synthesis,” says Dr. KIM Byung Hyo, the first author of the study. Notably, this methodology can measure the atomic displacement and strain on the surface atoms of individual nanoparticles. The strain analysis from the 3D reconstruction facilitates characterization of the active sites of nanocatalysts at the atomic scale, which will enable structure-based design to improve the catalytic activities. The methodology can also contribute more generally to the enhancement of nanomaterials’ performance.
“We have developed a groundbreaking methodology for determining the structures that govern the physical and chemical properties of nanoparticles at the atomic level in their native environment. The methodology will provide important clues in the synthesis of nanomaterials. The algorithm we introduced is related to new drug development through structure analysis of proteins and big data analysis, so we are expecting further application to new convergence research,” notes Director HYEON Taeghwan of the IBS Center for Nanoparticle Research.
Reference: “Critical differences in 3D atomic structure of individual ligand-protected nanocrystals in solution” by Byung Hyo Kim, Junyoung Heo, Sungin Kim, Cyril F. Reboul, Hoje Chun, Dohun Kang, Hyeonhu Bae, Hyejeong Hyun, Jongwoo Lim, Hoonkyung Lee, Byungchan Han, Taeghwan Hyeon, A. Paul Alivisatos, Peter Ercius, Hans Elmlund and Jungwon Park, 3 April 2020, Science.
DOI: 10.1126/science.aax3233
This study is jointly supported by the Institute for Basic Science (IBS), Samsung Science and Technology Foundation (SSTF), and the Molecular Foundry (U.S. Department of Energy (DOE) Office of Science User Facility).
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
1 Comment
Could the loose packing shown in some particles be an artifact of them being bound differently against the graphene boundary? It must support vibrational states? Even though the loose structure looks somewhat random, its range of displacements from the uniform positions seems uniform. Do you have any way to monitor the time variations of the graphene carbon positions at a distance from and around the droplet? Or, during fabrication (not mentioned here) could there be microstates responsible? You just say “defects” with no real guess at what might have happened.
This is really stretching, but can you use this to sort the nanoparticles? “Good” ones go that way, “bad ones” this way, “not clear” go there?? I found in industrial chemistry that the byproduct streams often have high value, once you can get enough to characterize and let people try things with it. That would require a focussed nanobeam. These are really different structures, so they should be easy to identify. The size falls into a range that is the frontier of many superresolution techniques. The computers can probably handle sorting without stretching current off-the-shelf methods. I think the low latency methods from AI optical might help.
Thank you so much for this elegant and brilliant way of looking at nanoparticles. It would be too much to hope it could be applied to proteins and viruses? I am getting tired of being stuck at home, so I keep dreaming about getting real time precise atomic coordinates for complete viruses. That is probably impossible, but I can’t stop thinking about it in my tireder moments. LOL!!!