
Study finds that the pace of evolution in organisms differs depending on the type of cells involved.
A new study finds that two species of worms have maintained strikingly similar patterns of gene activation and deactivation, even though they diverged from a common ancestor 20 million years ago.
The findings were recently published in the journal Science.
“It was just remarkable, with this evolutionary distance, that we should see such coherence in gene expression patterns,” said Dr. Robert Waterston, professor of genome sciences at the University of Washington School of Medicine in Seattle and a co-senior author of the paper. “I was surprised how well everything lined up.”
Broadly expressed genes resist change
Gene expression patterns tended to stay the same, or what evolutionary biologists call “conserved,” when changes would affect many types of cells, Waterston said.
“If the gene is broadly expressed in many cell types across the organism, it may be difficult to change expression,” he said. “But if the gene expression is limited to a single cell type or a few cell types, maybe it can succeed.”
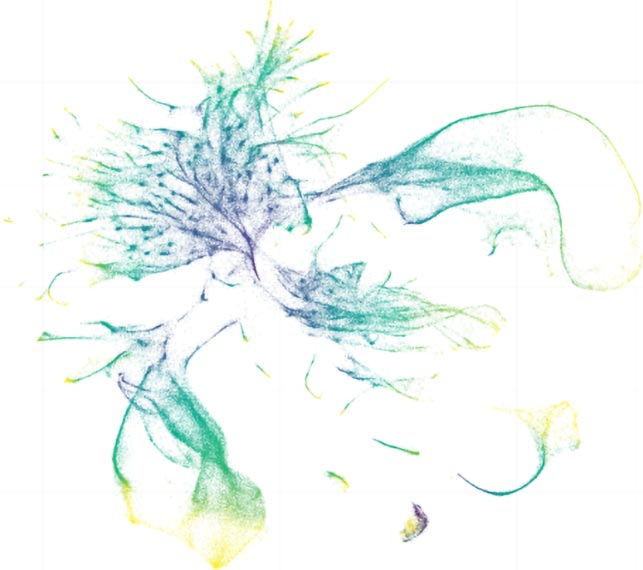
Divergence more common in specialized cells
When gene expression differed between the two worms, the changes were more likely to appear in specialized cell types. For instance, expression patterns in cells responsible for basic functions like muscle or digestion tended to remain stable, while those in cells involved in sensing and responding to the environment were more likely to change.
“Genes related to neuronal function, for example, seem to diverge more rapidly—perhaps because changes were needed to adapt to new environments—but for now, that’s speculation,” said Christopher R. L. Large, a postdoctoral fellow in the Department of Genetics at the Perelman School of Medicine at the University of Pennsylvania and the paper’s lead author. Large earned his Ph.D. in genome sciences from the UW School of Medicine.
Transparent worms as ideal model organisms
In the study, researchers compared gene expression patterns in two soil-dwelling roundworms, Caenorhabditis elegans and Caenorhabditis briggsae. Both species are well suited for studying development because they are small, about a millimeter long, made up of roughly 550 cells when fully developed, and transparent. These traits allow scientists to watch their cells divide and develop in real time. Notably, these worms share many of their approximately 20,000 genes with more complex organisms, including humans.
All the cells in both worms have been identified and mapped. Despite 20 million years of evolution, the two worms retain nearly identical body plans and cell types, with an almost one-to-one correspondence that makes them ideal subjects for comparison.
The goal of the study was to compare gene expression in every cell type of the two worms to determine what changes had occurred since they split from their common ancestor.
Single-cell RNA sequencing reveals developmental changes
To do this, the researchers measured levels of messenger RNA in every cell at various stages of embryonic development by using a technique called single-cell RNA sequencing.
Messenger RNA, or mRNA, carries the instructions for making proteins from active genes to the cell’s protein-making machinery. High levels of mRNA from a gene indicate that it is active. Low levels mean it’s inactive.
With the single-cell RNA sequencing technique, the researchers tracked the changes in the individual cells during the worms’ embryonic development from when the embryo was a ball of 28 mostly undifferentiated cells to when most cell types had developed into their near-final form, a process that takes about 12 hours
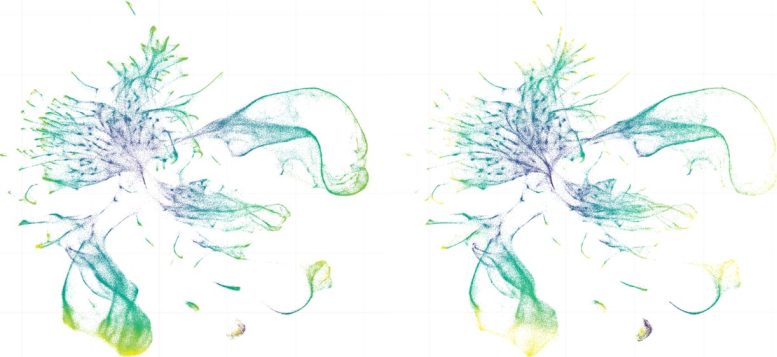
“We’ve been studying the evolution of development since the 1970s,” said Dr. Junhyong Kim, professor of biology and director of the Penn Genome Frontiers Institute, a co-senior author of the study. “But this is the first time that we’ve been able to compare development in every single cell of two different organisms.”
Kim said the finding that some gene expression was conserved wasn’t surprising, because of how similar the worms’ bodies are. But it was surprising that when there were changes, those changes appeared to have no effect on the body plan.
The study describes where and when gene expression patterns differ between the species but doesn’t yet explain why, said Dr. John Isaac Murray, associate professor of genetics at the Perelman School of Medicine and the study’s third senior author.
“It’s hard to say whether any of the differences we observed were due to evolutionary adaptation or simply the result of genetic drift, where changes happen randomly,” he said. “But this approach will allow us to explore many unanswered questions about evolution.”
Reference: “Lineage-resolved analysis of embryonic gene expression evolution in C. elegans and C. briggsae” by Christopher R. L. Large, Rupa Khanal, LaDeana Hillier, Chau Huynh, Connor Kubo, Junhyong Kim, Robert H. Waterston and John I. Murray, 19 June 2025, Science.
DOI: 10.1126/science.adu8249
This study was supported by the National Institutes of Health (HD105819, HG010478, HG007355) and the National Sciences Foundation (PRFB2305513).
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.