
Dragons and Brain Evolution
These days, dragons are keeping Game of Thrones fans on their toes. But according to recent research conducted by Max Planck scientists on the brain of the Australian bearded dragon Pogona vitticeps, they are also offering significant insights into the evolution of vertebrate brains.
Early tetrapods (animals with four limbs) made the move from aquatic to terrestrial environments 320 million years ago, which resulted in the three main clades of vertebrates today: reptiles, birds (an offshoot of the reptilian tree), and mammals. All tetrapod brains possess a similar basal architecture established during early development because of common ancestry.
It is unclear, however, how variations in this common “Bauplan” contributed to clade-specific characteristics. To answer this issue, researchers at the Max Planck Institute for Brain Research in Frankfurt created a molecular atlas of the dragon brain and compared it to one from mice. Contrary to conventional opinion, which holds that a mammalian brain is a combination of an ancient “reptilian” brain and modern mammalian traits, their results imply that both reptilian and mammalian brains have developed distinct clade-specific neuron types and circuits from a common ancestral set.
“Neurons are the most diverse cell types in the body. Their evolutionary diversification reflects alterations in the developmental processes that produce them and may drive changes in the neural circuits they belong to”, says Professor Gilles Laurent, Director at the Max Planck Institute for Brain Research who led the new study published in Science.
“For example, distinct brain areas do not work in isolation, suggesting that the evolution of interconnected regions, such as the thalamus and cerebral cortex, might in some way be correlated. Also, a brain area in reptiles and mammals that derived from a common ancestral structure might have evolved in such a way that it remains ancestral in one clade today, while it is “modern” in the other. Conversely, it could be that both clades now contain a mix of common (ancient) and specific (novel) neuron types. These are the sorts of questions that our experiments tried to address”, Laurent adds.
While traditional approaches to comparing developmental regions and projections in the brain do not have the necessary resolution to reveal these similarities and differences, Laurent and his team took a cellular transcriptomic approach. Using a technique called single-cell RNA sequencing that detects a large fraction of the RNA molecules (transcriptomes) present in single cells, the scientists generated a cell-type atlas of the brain of the Australian bearded dragon Pogona vitticeps and compared it to existing mouse brain datasets.
Transcriptomic Comparisons Reveal Shared Classes of Neuron Types
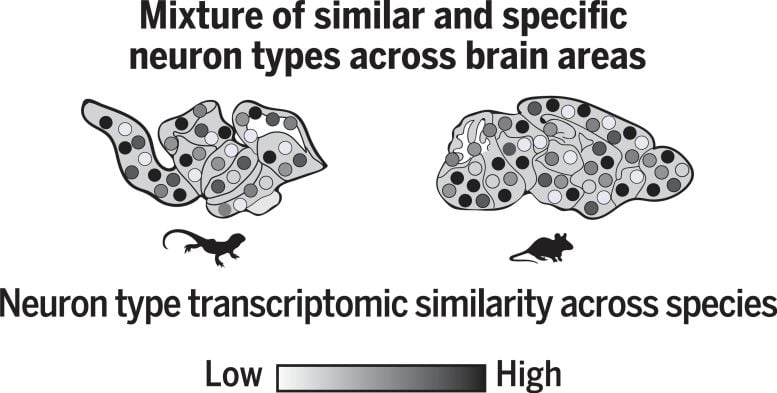
“We profiled over 280,000 cells from the brain of Pogona and identified 233 distinct types of neurons”, explains David Hain, a graduate student in the Laurent Lab and co-first author of the study. “Computational integration of our data with mouse data revealed that these neurons can be grouped transcriptomically in common families, that probably represent ancestral neuron types”, says Hain. In addition, he found that most areas of the brain contain a mix of common (ancient) and specific (novel) neuron types, as shown in the figure below.
Graduate student Tatiana Gallego-Flores used histological techniques to map these cell types throughout the dragon brain and observed (among others) that neurons in the thalamus could be grouped in two transcriptomic and anatomical domains, defined by their connectivity to other regions of the brain. Because these connected regions have had different fates in mammals and in reptiles, one of these regions being highly divergent, comparing the thalamic transcriptomes of these two domains proved to be very interesting. Indeed, it revealed that transcriptomic divergence matched that of the target regions.
“This suggests that neuronal transcriptomic identity somewhat reflects, at least in part, the long-range connectivity of a region to its targets. Since we do not have the brains of ancient vertebrates, reconstructing the evolution of the brain over the past half billion years will require connecting together very complex molecular, developmental, anatomical, and functional data in a way that is self-consistent. We live in very exciting times because this is becoming possible”, concludes Laurent.
Reference: “Molecular diversity and evolution of neuron types in the amniote brain” by David Hain, Tatiana Gallego-Flores, Michaela Klinkmann, Angeles Macias, Elena Ciirdaeva, Anja Arends, Christina Thum, Georgi Tushev, Friedrich Kretschmer, Maria Antonietta Tosches and Gilles Laurent, 2 September 2022, Science.
DOI: 10.1126/science.abp8202
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.