
Many of the fundamental principles in biology and essentially all pathways regulating development were identified in so-called genetics screens. Originally pioneered in the fruit fly Drosophila and the nematode C. elegans, genetic screens involve inactivation of many genes one by one. By analyzing the consequences of gene loss, scientists can draw conclusions about its function. This way, for example, all genes required for formation of a brain can be identified.
Genetic screens can routinely be carried out in flies and worms. In humans, a wealth of knowledge exists about genetic disorders and the consequences of disease-relevant mutations, but their systematic analysis was impossible. Now, the Knoblich lab at IMBA has developed a groundbreaking technique allowing hundreds of genes to be analyzed in parallel in human tissue. They named the new technology CRISPR-LICHT and published their findings in the journal Science.
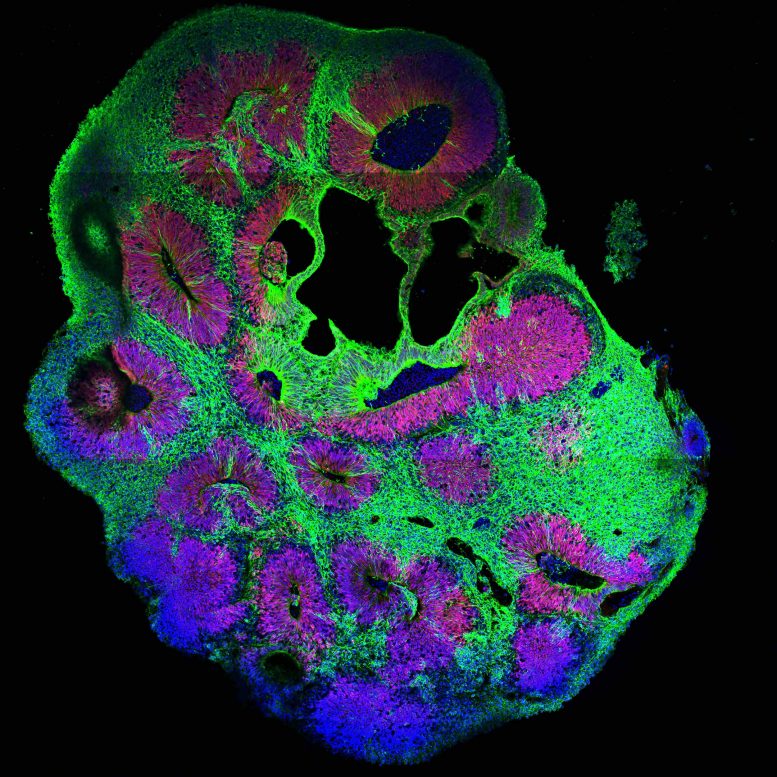
By using cerebral organoids, a 3D cell culture model for the human brain developed in Jürgen Knoblich’s group at IMBA, hundreds of mutations can now be analyzed for their role in the human brain using CRISPR-LICHT.
“The basis of the technique is a combination of the well-known CRISPR-Cas9 technology, which was awarded the Nobel Prize in October 2020, and a dual-barcoding method. The key trick was to apply a guide RNA, but also a genetic barcode, a piece of DNA that we add to the genome of the cells we use to grow organoids. This allows us to see the entire cell lineage of each organoid, while a second barcode lets us to count the number of cells generated by each starting cell. This reduces noise and so we can determine the effect of each guide RNA on the number of cells produced during organoid growth. To describe our approach, we termed the method CRIPSR-Lineage Tracing at Cellular resolution in Heterogenous Tissue (CRISPR-LICHT),” explains co-first author Dominik Lindenhofer, PhD student at IMBA.
The researchers applied CRISPR-LICHT to microcephaly, a genetic disorder caused by a reduction in brain size and severe mental impairment in patients. By means of this revolutionary new technology, the scientists screened through all genes suspected to play a role in the disease.
“Not only were we able to identify microcephaly genes with CRISPR-LICHT, but we also pinpointed a specific mechanism involved in controlling the size of the brain,” says IMBA postdoc and co-first author Christopher Esk. The endoplasmic reticulum (ER) was identified as main hub in controlling extracellular matrix protein secretion. This mechanism affects the integrity of tissue, and thus the brain size and was identified as one cause of microcephaly.
Genetic screens in Drosophila have long been an established tool in genome-wide screenings and have a long tradition in Vienna. The “Vienna Drosophila Research Center” (VDRC), co-developed by scientists from IMBA, is the only Drosophila stock center in Europe and boasts one of the worldwide largest fly collections for functional gene studies. Jürgen Knoblich, IMBA Scientific Director and group leader, also has his roots in fly genetics and, thanks to the fruit fly, gained important insights into the role of stem cells for brain development.
“We are very excited to report that we now can routinely do genetic screens in complex organoid systems. The method can be applied to other organoid models and to any disease affecting organ formation. It is a completely new approach to analyzing brain disorders, and bears incredible future potential, as it can be applied to any brain disease, including autism. Our work was only possible due to the collaborative spirit at the Vienna BioCenter, as groups from our neighboring institutes of the Max Perutz Labs and the Institute of Molecular Pathology also substantially contributed to the ideas that helped develop the novel technology,” says Jürgen Knoblich, last author of the study.
Reference: “A human tissue screen identifies a regulator of ER secretion as a brain size determinant” by Christopher Esk, Dominik Lindenhofer, Simon Haendeler, Roelof A. Wester, Florian Pflug, Benoit Schroeder, Joshua A. Bagley, Ulrich Elling, Johannes Zuber, Arndt von Haeseler and Jürgen A. Knoblich, 29 October 2020, Science.
DOI: 10.1126/science.abb5390
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.