A bold project to read the complete genetic sequences of every known vertebrate species reaches its first milestone by publishing new methods and the first 25 high-quality genomes.
It’s one of the most audacious projects in biology today – reading the entire genome of every bird, mammal, lizard, fish, and all other creatures with backbones.
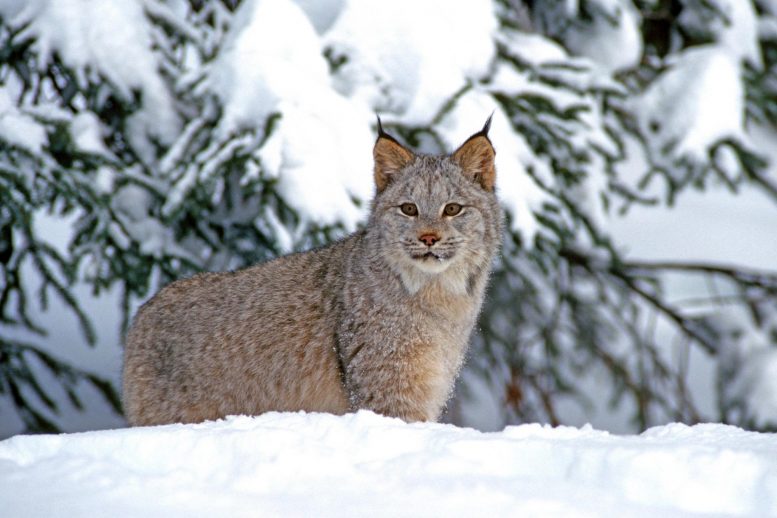
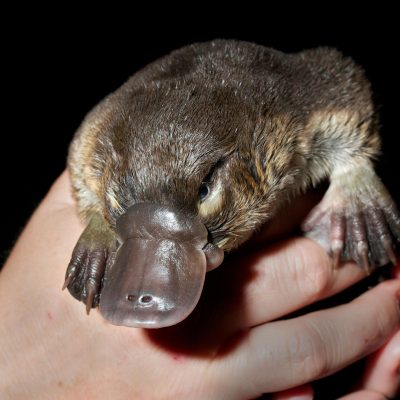
And now comes the first major payoff from the Vertebrate Genomes Project (VGP): near complete, high-quality genomes of 25 species, Howard Hughes Medical Institute (HHMI) Investigator Erich Jarvis with scores of coauthors report April 28, 2021, in the journal Nature. These species include the greater horseshoe bat, the Canada lynx, the platypus, and the kākāpō parrot – one of the first high-quality genomes of an endangered vertebrate species.
The paper also lays out the technical advances that let scientists achieve a new level of accuracy and completeness and paves the way for decoding the genomes of the roughly 70,000 vertebrate species living today, says HHMI Investigator and study coauthor David Haussler, a computational geneticist at the University of California, Santa Cruz (UCSC). “We will get a spectacular picture of how nature actually filled out all the ecosystems with this unbelievably diverse array of animals.”
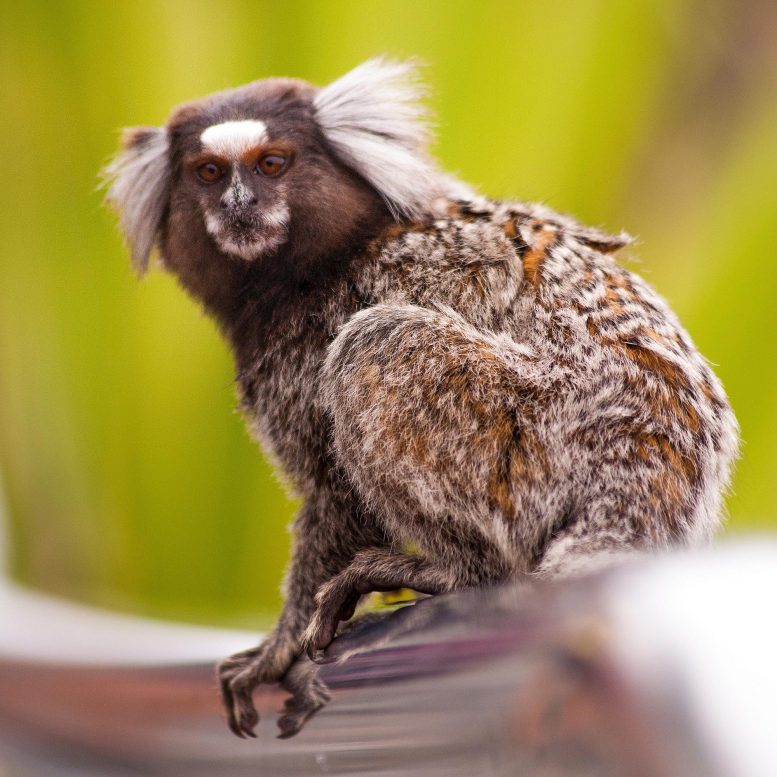
Together with a slew of accompanying papers, the work is beginning to deliver on that promise. The project team has discovered previously unknown chromosomes in the zebra finch genome, for example, and a surprise finding about genetic differences between marmoset and human brains. The new research also offers hope for saving the kākāpō and the endangered vaquita dolphin from extinction.
“These 25 genomes represent a key milestone,” explains Jarvis, VGP chair and a neurogeneticist at The Rockefeller University. “We are learning a lot more than we expected,” he says. “The work is a proof of principle for what’s to come.”

From 10K to 70K
The VGP milestone has been years in the making. The project’s origins date back to the late-2000s, when Haussler, geneticist Stephen O’Brien, and Oliver Ryder, director of conservation genetics at the San Diego Zoo, figured it was time to think big.
Instead of sequencing just a few species, such as humans, and model organisms like fruit flies, why not read the complete genomes of ten thousand animals in a bold “Genome 10K” effort? At the time, though, the price tag was hundreds of millions of dollars, and the plan never really got off the ground. “Everyone knew it was a great idea, but nobody wanted to pay for it,” recalls HHMI Investigator and HHMI Professor Beth Shapiro, an evolutionary biologist at UCSC and a coauthor of the Nature paper.
Plus, scientists’ early efforts at spelling out, or “sequencing,” all the DNA letters in an animal’s genome were riddled with errors. In the original approach used to complete the first rough human genome in 2003, scientists chopped up DNA into short pieces a few hundred letters long and read those letters. Then came the fiendishly difficult job of assembling the fragments in the right order. The methods weren’t up to task, resulting in misassemblies, major gaps, and other mistakes. Often it wasn’t even possible to map genes to individual chromosomes.

The introduction of new sequencing technologies with shorter reads helped make the idea of reading thousands of genomes possible. These rapidly developing technologies slashed costs but also reduced the quality in genome assembly structure. Then in 2015, Haussler and colleagues brought in Jarvis, a pioneer in deciphering the intricate neural circuits that let birds trill new tunes after listening to others’ songs. Jarvis had already shown a knack for managing big, complex efforts. In 2014, he and more than a hundred colleagues sequenced the genomes of 48 bird species, which turned up new genes involved in vocal learning. “David and others asked me to take on leadership of the Genome 10K project,” Jarvis recalls. “They felt I had the personality for it.” Or, as Shapiro puts it: “Erich is a very pushy leader, in a nice way. What he wants to happen, he will make happen.”
Jarvis expanded and rebranded the Genome 10K idea to include all vertebrate genomes. He also helped launch a new sequencing center at Rockefeller that, together with one at the Max Planck Institute in Germany led by former HHMI Janelia Research Campus Group Leader Gene Myers, and another at the Sanger Institute in the UK led by Richard Durbin and Mark Blaxter, is currently producing most of the VGP genome data. He asked Adam Phillippy, a leading genome expert at the National Human Genome Research Institute (NHGRI), to chair the VGP assembly team. Then, he found about 60 top scientists willing to use their own grant money to pay for the sequencing costs at the centers to tackle the genomes they were most interested in. The team also negotiated with the Māori in New Zealand and officials in Mexico to get kākāpō and vaquita samples in “a beautiful example of international collaboration,” says Sadye Paez, program director of the VGP at Rockefeller.
Opening Doors
The massive team of researchers pulled off a series of technological advances. The new sequencing machines let them read DNA chunks 10,000 or more letters long, instead of just a few hundred. The researchers also devised clever methods for assembling those segments into individual chromosomes. They have been able to tease out which genes were inherited from the mother and the father. This solves a particularly thorny problem known as “false duplication,” where scientists mistakenly label maternal and paternal copies of the same gene as two separate sister genes.
“I think this work opens a set of really important doors, since the technical aspects of assembly have been the bottleneck for sequencing genomes in the past,” says Jenny Tung, a geneticist at Duke University, who was not directly involved with the research. Having high-quality sequencing data “will transform the types of questions that people can ask,” she says.
The team’s improved accuracy shows that previous genome sequences are seriously incomplete. In the zebra finch, for example, the team found eight new chromosomes and about 900 genes that had been thought to be missing. Previously unknown chromosomes popped up in the platypus as well, as members of the team reported online in Nature earlier this year. The researchers also plowed through, and correctly assembled, long stretches of repetitive DNA, much of which contain just two of the four genetic letters. Some scientists considered these stretches to be non-functional “junk” or “dark matter.” Wrong. Many of the repeats occur in regions of the genome that code for proteins, says Jarvis, suggesting that the DNA plays a surprisingly crucial role in turning genes on or off.
That’s just the start of what the Nature paper envisions as “a new era of discovery across the life sciences.” With every new genome sequence, Jarvis and his collaborators uncover new – and often unexpected – findings. Jarvis’s lab, for example, has finally nabbed the regulatory region of a key gene parrots and songbirds need to learn tunes; next, his team will try to figure out how it works. The marmoset genome yielded several surprises. While marmoset and human brain genes are largely conserved, the marmoset has several genes for human pathogenic amino acids. That highlights the need to consider genomic context when developing animal models, the team reports in a companion paper in Nature. And in findings published last year in Nature, a group led by Professor Emma Teeling at University College Dublin in Ireland discovered that some bats have lost immunity-related genes, which could help explain their ability to tolerate viruses like SARS-CoV-2, which causes COVID-19.
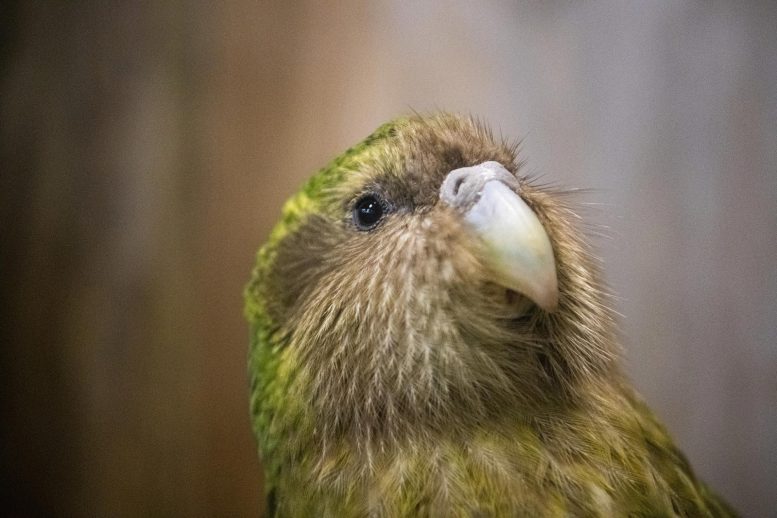

The new information also may boost efforts to save rare species. “It is a critically important moral duty to help species that are going extinct,” Jarvis says. That’s why the team collected samples from a kākāpō named Jane, part of a captive breeding program that has brought the parrot back from the brink of extinction. In a paper published in the new journal Cell Genomics, of the Cell family of journals, Nicolas Dussex at the University of Otago and colleagues described their studies of Jane’s genes along with other individuals. The work revealed that the last surviving kākāpō population, isolated on an island off New Zealand for the last 10,000 years, has somehow purged deleterious mutations, despite the species’ low genetic diversity. A similar finding was seen for the vaquita, with an estimated 10-20 individuals left on the planet, in a study published in Molecular Ecology Resources, led by Phil Morin at the National Oceanic and Atmospheric Administration Fisheries in La Jolla, California. “That means there is hope for conserving the species,” Jarvis concludes.

A Clear Path
The VGP is now focused on sequencing even more species. The project team’s next goal is finishing 260 genomes, representing all vertebrate orders, and then snaring enough funding to tackle thousands more, representing all families. That work won’t be easy, and it will inevitably bring new technical and logistical challenges, Tung says. Once hundreds or even thousands of animals readily found in zoos or labs have been sequenced, scientists may face ethical hurdles obtaining samples from other species, especially when the animals are rare or endangered.
But with the new paper, the path ahead looks clearer than it has in years. The VGP model is even inspiring other large sequencing efforts, including the Earth Biogenome Project, which aims to decode the genomes of all eukaryotic species within 10 years. Perhaps for the first time, it seems possible to realize the dream that Haussler and many others share of reading every letter of every organism’s genome. Darwin saw the enormous diversity of life on Earth as “endless forms most beautiful,” Haussler observes. “Now, we have an incredible opportunity to see how those forms came about.”
Reference: “Towards complete and error-free genome assemblies of all vertebrate species” by Arang Rhie, Shane A. McCarthy, Olivier Fedrigo, Joana Damas, Giulio Formenti, Sergey Koren, Marcela Uliano-Silva, William Chow, Arkarachai Fungtammasan, Juwan Kim, Chul Lee, Byung June Ko, Mark Chaisson, Gregory L. Gedman, Lindsey J. Cantin, Francoise Thibaud-Nissen, Leanne Haggerty, Iliana Bista, Michelle Smith, Bettina Haase, Jacquelyn Mountcastle, Sylke Winkler, Sadye Paez, Jason Howard, Sonja C. Vernes, Tanya M. Lama, Frank Grutzner, Wesley C. Warren, Christopher N. Balakrishnan, Dave Burt, Julia M. George, Matthew T. Biegler, David Iorns, Andrew Digby, Daryl Eason, Bruce Robertson, Taylor Edwards, Mark Wilkinson, George Turner, Axel Meyer, Andreas F. Kautt, Paolo Franchini, H. William Detrich III, Hannes Svardal, Maximilian Wagner, Gavin J. P. Naylor, Martin Pippel, Milan Malinsky, Mark Mooney, Maria Simbirsky, Brett T. Hannigan, Trevor Pesout, Marlys Houck, Ann Misuraca, Sarah B. Kingan, Richard Hall, Zev Kronenberg, Ivan Sović, Christopher Dunn, Zemin Ning, Alex Hastie, Joyce Lee, Siddarth Selvaraj, Richard E. Green, Nicholas H. Putnam, Ivo Gut, Jay Ghurye, Erik Garrison, Ying Sims, Joanna Collins, Sarah Pelan, James Torrance, Alan Tracey, Jonathan Wood, Robel E. Dagnew, Dengfeng Guan, Sarah E. London, David F. Clayton, Claudio V. Mello, Samantha R. Friedrich, Peter V. Lovell, Ekaterina Osipova, Farooq O. Al-Ajli, Simona Secomandi, Heebal Kim, Constantina Theofanopoulou, Michael Hiller, Yang Zhou, Robert S. Harris, Kateryna D. Makova, Paul Medvedev, Jinna Hoffman, Patrick Masterson, Karen Clark, Fergal Martin, Kevin Howe, Paul Flicek, Brian P. Walenz, Woori Kwak, Hiram Clawson, Mark Diekhans, Luis Nassar, Benedict Paten, Robert H. S. Kraus, Andrew J. Crawford, M. Thomas P. Gilbert, Guojie Zhang, Byrappa Venkatesh, Robert W. Murphy, Klaus-Peter Koepfli, Beth Shapiro, Warren E. Johnson, Federica Di Palma, Tomas Marques-Bonet, Emma C. Teeling, Tandy Warnow, Jennifer Marshall Graves, Oliver A. Ryder, David Haussler, Stephen J. O’Brien, Jonas Korlach, Harris A. Lewin, Kerstin Howe, Eugene W. Myers, Richard Durbin, Adam M. Phillippy and Erich D. Jarvis, 28 April 2021, Nature.
DOI: 10.1038/s41586-021-03451-0
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
4 Comments
What? Not the naked mole rat? 🙁
The phrase “In the beginning was the WORD”, has distracted me for some time. I believe truly grasping its significance leads to the effort of the old thinkers – through too much poetry – to unlock the history of our existence and the suggestion that we alone have gathered these formulas in order to reproduce them selectively on our next planet. “In my father’s house are many mansions”. All we’ll need is these spectacular 3-D printers, a little clay; and about a week. So who says this is the first time?
Boring!!!!
To study compĺete genetic sequènces òf a huge numbers of othèr creatuŕes implies equivalent to study a large number of stars in fine sèquence order.