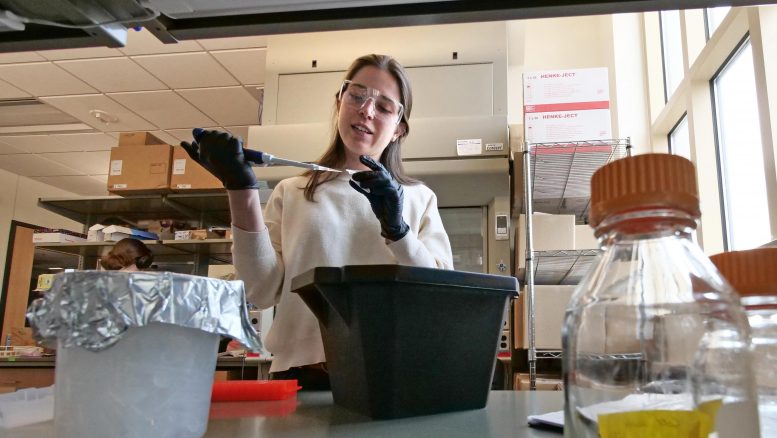

CU Boulder scientists use yeast to forecast virus mutations, paving the way for smarter vaccines and antibody therapies against COVID-19 and beyond.
Researchers at CU Boulder have developed a platform that can quickly identify common mutations on the SARS-CoV-2 virus that allow it to escape antibodies and infect cells.
Published recently in Cell Reports, the research marks a major step toward successfully developing a universal vaccine for not only COVID-19, but also potentially for influenza, HIV and other deadly global viruses.
“We’ve developed a predictive tool that can tell you ahead of time which antibodies are going to be effective against circulating strains of virus,” said lead author Timothy Whitehead, associate professor of chemical and biological engineering. “But the implications for this technology are more profound: If you can predict what the variants will be in a given season, you could get vaccinated to match the sequence that will occur and short-circuit this seasonal variation.”
The research team’s secret ingredient? Baker’s yeast.
They developed a genetically modified version of this innocuous material to express some of SARS-CoV-2’s viral spike proteins along the yeast’s surface, with which they can map resulting mutations that form and escape neutralizing antibodies. The resulting roadmap could inform the development of more effective booster vaccines and tailored antibody treatments for patients with severe cases of COVID-19, said Whitehead.
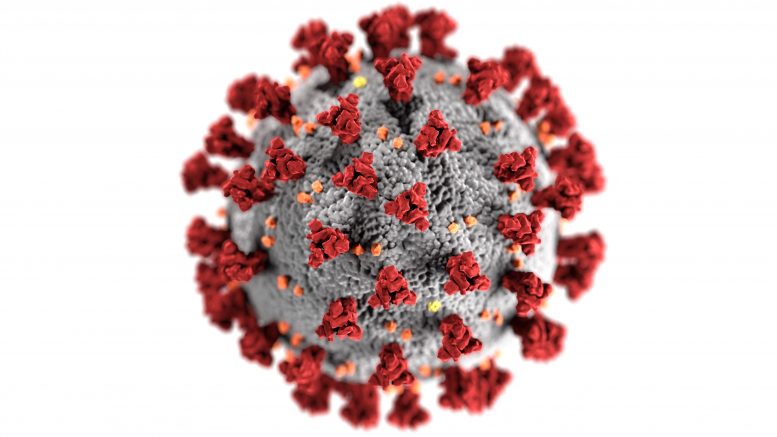
The Key to the Crown
Spike proteins are sharp bumps that stick out from the surface of viruses in the coronavirus family. Under a microscope, they can appear like a crown, which is where coronaviruses—corona being Latin for “crown”—gain their name, and how they bind to cells like a key in a lock. When antibodies recognize them, latch on, and prevent them from binding to cells, they prevent infection.
But when spike proteins mutate, antibodies do a double-take.
“There are mutations on the spike protein that prevent an antibody from going in and recognizing it. Just like getting a new haircut, you look like a different person; this looks like a different virus to that antibody,” said Whitehead.

In the case of the more contagious Delta variant that emerged in 2021, mutations on the spike proteins have made it more contagious and reduced the efficacy of some antibody therapies.
What if there was a way to predict which mutations could emerge next—and therefore prepare for them? Earlier this year, Whitehead’s team set out to find a way.
What Comes After Delta?
Some antibodies which can bind to different locations have been used in treatment cocktails given to COVID-19 patients. But the strains of the virus now circulating within the U.S. are different enough that some of these antibody therapies no longer seem effective, said Whitehead.
So first, the researchers needed to identify mutations on the spike protein that could prevent these antibodies from working. Then they wanted to predict what mutations are likely to occur next—what could become the zeta, eta or theta variant?
“When the pandemic started, we saw the opportunity to apply techniques mastered by our lab to make a contribution,” said Irene Francino Urdaniz, co-author on the paper, graduate student in chemical and biological engineering and a Balsells fellow. “When a new variant was detected, based on my research, I could most of the times guess which mutations were present. I am very excited to have contributed with my work not only to this pandemic but possibly to future vaccines.”

Francino-Urdaniz developed a genetically engineered strain of common baker’s yeast, which could display different portions of the viral spike protein on its surface. She then discovered how to screen through thousands of mutations in a single test tube to find the ones that evaded neutralizing antibodies.
As some home-bound bakers discovered in 2020 while experimenting with sourdough starters, yeast grows quite quickly. This means that the researchers can see a wide variety of mutations develop at the same speed at which the yeast can grow—leaps and bounds faster than the rate at which mutations will emerge in real time. This could give scientists an invaluable head start.
A Universal Vaccine
The researchers have already found some of the same mutations now circulating the globe, as well as identified more mutations with the potential to evade our immune systems. They will also provide all their libraries of information, methods and software as an openly available community resource to accelerate new therapeutic strategies against SARS-CoV-2.
This means the next COVID-19 vaccine or booster shot produced for the public could have the ability to pack the most punch possible. It also gives hope for those who are immunocompromised or remain at a higher risk of contracting a bad case, as this research can be applied to proactively prepare antibody cocktails for specific mutations, giving them a better chance at survival and recovery.
But the promise doesn’t stop there. Due to the adaptability of new mRNA vaccines which work with spike proteins, the applications of this research are not limited to one virus, said Whitehead.
“You can use it for mapping trajectories for influenza and for HIV potentially; for other viral diseases that are known, and also potentially emerging pandemic ones,” he said.
Reference: “One-shot identification of SARS-CoV-2 S RBD escape mutants using yeast screening” by Irene M. Francino-Urdaniz, Paul J. Steiner, Monica B. Kirby, Fangzhu Zhao, Cyrus M. Haas, Shawn Barman, Emily R. Rhodes, Alison C. Leonard, Linghang Peng, Kayla G. Sprenger, Joseph G. Jardine and Timothy A. Whitehead, 9 August 2021, Cell Reports.
DOI: 10.1016/j.celrep.2021.109627
Other authors on this publication include Paul Steiner, Monica Kirby, Cyrus Haas, Emily Rhodes, Alison Leonard and Kayla Sprenger in Chemical and Biological Engineering at CU Boulder; Fangzhu Zhao, Shawn Barman and Linghang Peng of The Scripps Research Institute; and Joseph Jardine of the International AIDS Vaccine Initiative.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.