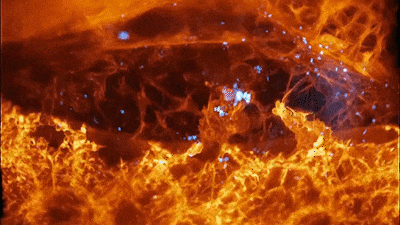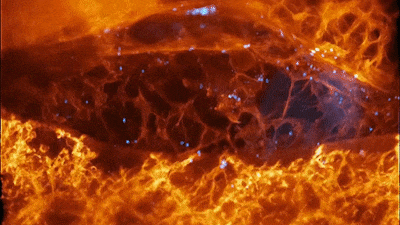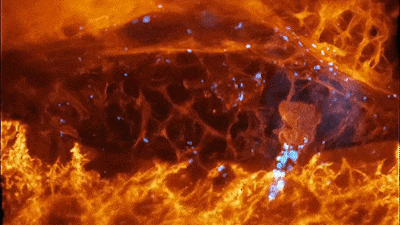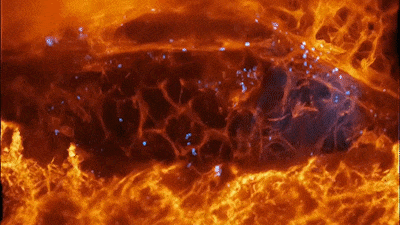
Cells of a zebrafish eye, computationally “exploded.” Video by Liu et al
More than 350 years ago, the English natural philosopher Robert Hooke looked through a microscope at a thin slice of cork and discovered that it was made of small, box-like compartments, which he named “cells.”
From that moment on, Hooke and countless inquisitive minds after him strove for a better view of these fundamental building blocks of life. And now, the window into the cellular world has become a lot clearer.
In a new study in the April 20 issue of Science, researchers from Howard Hughes Medical Institute’s (HHMI) Janelia Research Campus, Harvard Medical School, and collaborating institutions report the development of a microscope capable of capturing, in unprecedented detail, 3-D images and videos of cells inside living organisms.
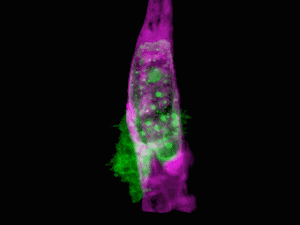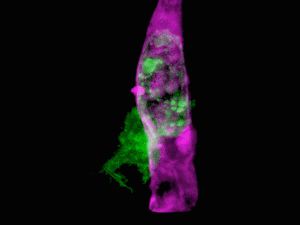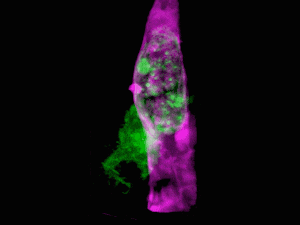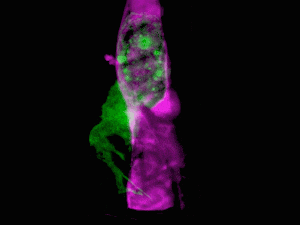
Adapting a technique used by astronomers to study distant stars, the research team, led by Nobel laureate and Janelia group leader Eric Betzig, showcased the new technology by generating a series of stunning movies: cancer cells crawling through blood vessels, spinal nerve cells wiring up into circuits, immune cells cruising through a zebrafish’s inner ear, and much more.
The resolution of the microscope is so powerful it can even capture subcellular details such as the dynamics of minuscule bubbles known as vesicles, which transport molecular cargo through to the cell.
“This is the miracle of being able to see what we have never been able to see before. It’s simply incredible,” said study co-author Tomas Kirchhausen, HMS professor of cell biology, and the Springer Family Chair of pediatrics and a senior investigator at Boston Children’s Hospital.
“Every time we’ve done an experiment with this microscope, we’ve observed something novel — and generated new ideas and hypotheses to test,” Kirchhausen said. “It can be used to study almost any problem in a biological system or organism I can think of.”
While scientists have used microscopes to look at cells for centuries, the clearest views thus far have come from cells isolated on glass slides. Visualizing living cells in real time inside live organisms has remained far more challenging.
Cells of interest are surrounded by tissues and other biological structures that scramble light coming from and returning to a microscope objective, which blurs and obscures important details. Light powerful enough to penetrate biological structures and yield a crisper view of cells, on the other hand, can damage tissues.
“This raises the nagging doubt that we are not seeing cells in their native state, happily ensconced in the organism in which they evolved,” said Betzig, who is corresponding author on the study. “It’s often said that seeing is believing, but when it comes to cell biology, I think the more appropriate question is, ‘When can we believe what we see?’”
A new microscope technology allows us to see cells and tissues like we’ve never seen before.
Betzig and colleagues applied this principle to the microscopic world, using a two-photon laser to create an adaptive optics system that maintains the thin illumination of a lattice light sheet as it penetrates an organism to generate distortion-free images of their target.
The team validated the new adaptive optics/lattice light-sheet microscope on a variety of biological samples, carrying out much of the work through the laboratories of Kirchhausen and Sean Megason, HMS associate professor of systems biology.
To make sense of the data they generated, the team created customized software and computational and visualization workflows, spearheaded by study co-lead authors Gokul Upadhyayula, HMS instructor in pediatrics at Boston Children’s and research associate in the Kirchhausen lab, and Tsung-Li Liu, formerly a research scientist in the Betzig lab at HHMI.
“For the types of data we generated, there’s no one commercial software that we can use to create interpretable movies and extract biologically meaningful information, so we built the necessary tools,” Upadhyayula said. “This allowed us to understand what we acquired and visualize the data in meaningful ways, including, in the near future, fully interactive 3-D movies.”
Into focus
The results were remarkable. In one movie, a fiery orange immune cell wriggles madly through a zebrafish’s ear while scooping up blue sugar particles along the way. In another, a migrating cancer cell trails sticky appendages as it rolls through a blood vessel and attempts to squeeze through the vessel wall.
The team captured movies of the behavior of organelles as they remodel themselves inside cells during cell division, and even visualized in real-time and at near-molecular detail the process of clathrin-mediated endocytosis, which cells use to capture materials from their exterior environment.
“I work on understanding how cells ‘eat’ using machinery based on vesicular carriers, and all my life I’ve dreamed of seeing this in a live organism,” Kirchhausen said. “We have finally achieved this.”
The complexity of the 3-D multicellular environment can be overwhelming, Betzig said, but the clarity of the imaging allows the researchers to computationally “explode” apart the individual cells in the tissue to focus on the dynamics within any particular one.
“It’s like ‘Star Trek.’ It’s the age of exploration again,” Upadhyayula said. “We don’t even know what questions to ask yet because we’ve never even seen some of these biologies at this level of detail.”
All this detail is hard to see without adaptive optics, Betzig said. “It’s just too damn fuzzy.” In his view, adaptive optics is one of the most important areas in microscopy research today, and the lattice light-sheet microscope, which excels at 3-D live imaging, is the perfect platform to showcase its power.
The next step is making the technology affordable and user-friendly. The current microscope takes up a 10-foot-long table. In collaboration with Kirchhausen and Upadhyayula, Betzig’s team is working on a next-generation version that should fit on a small desk at a cost within the reach of individual labs.
The first such instrument will go to Janelia’s Advanced Imaging Center, where scientists from around the world can apply to use it. A second instrument built at the same time will be located in the Kirchhausen laboratory at the HMS Quad in Boston. Plans to build the instrument will also be made available.
Reference: “Observing the cell in its native state: Imaging subcellular dynamics in multicellular organisms” by Tsung-Li Liu, Srigokul Upadhyayula, Daniel E. Milkie, Ved Singh, Kai Wang, Ian A. Swinburne, Kishore R. Mosaliganti, Zach M. Collins, Tom W. Hiscock, Jamien Shea, Abraham Q. Kohrman, Taylor N. Medwig, Daphne Dambournet, Ryan Forster, Brian Cunniff, Yuan Ruan, Hanako Yashiro, Steffen Scholpp, Elliot M. Meyerowitz, Dirk Hockemeyer, David G. Drubin, Benjamin L. Martin, David Q. Matus, Minoru Koyama, Sean G. Megason, Tom Kirchhausen and Eric Betzig, 20 April 2018, Science.
DOI: 10.1126/science.aaq1392
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
3 Comments
Oculus Rift, next stop.
6
Keep up the good work. Your stories are the best.