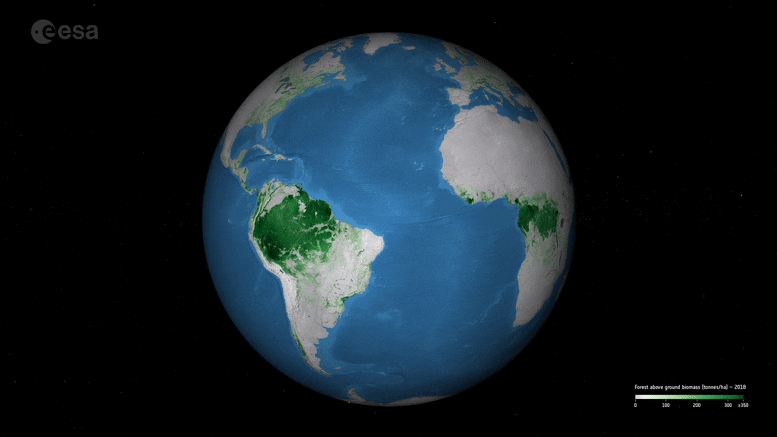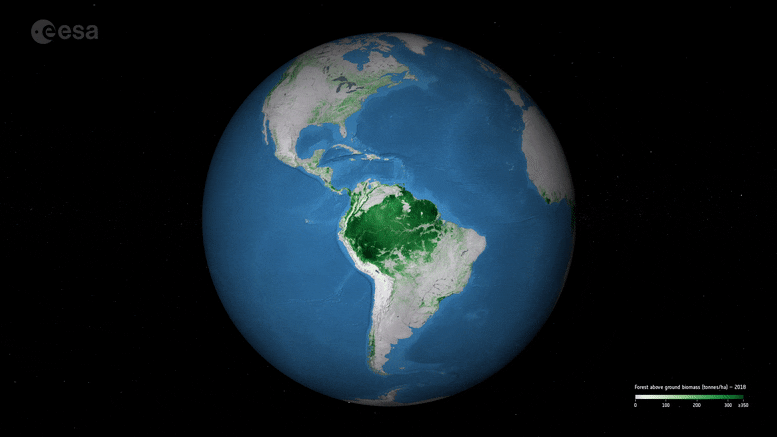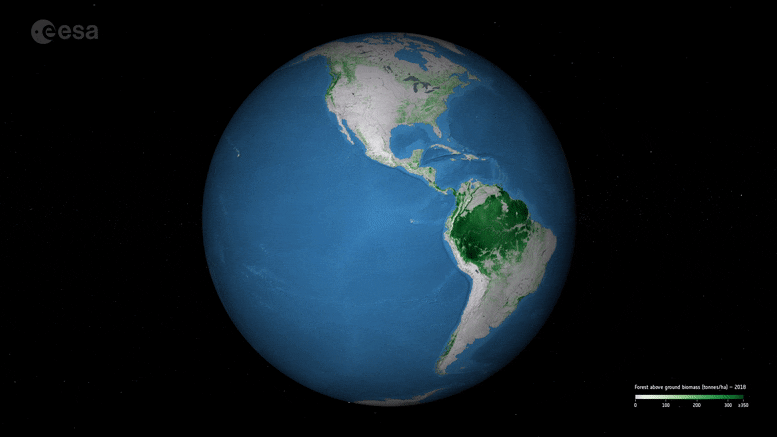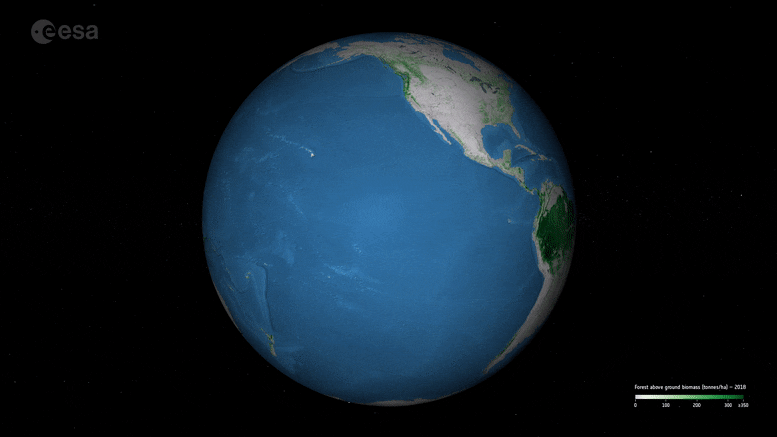
New ESA maps of global forest biomass help track carbon storage and support climate goals like the Paris Agreement.
Fluctuations in the carbon-rich biomass held within the world’s forests can contribute to, or slow, climate change. A series of new maps of above ground biomass, generated using space observations, is set to help our understanding of global carbon cycling and support forest management, emissions reduction and sustainable development policy goals.
Above ground biomass refers to the stem, bark, branches, and twigs of woody components of vegetation. As photosynthesis withdraws carbon dioxide from the atmosphere, it stores carbon in vegetation in an amount comparable to that of atmospheric carbon. The vegetation has the potential to sequester more carbon in the future or to contribute as an even larger source.
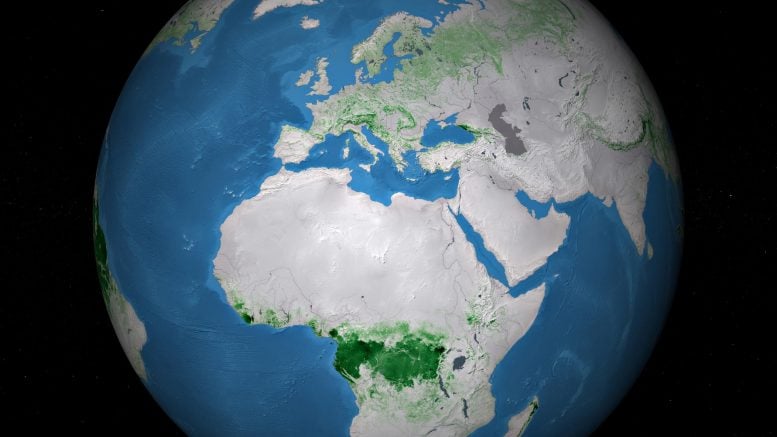
Vegetation biomass is a crucial ecological variable for understanding the evolution and potential future changes of the climate system, on a local, regional, and even global scale. For this reason, it is recognized by the Global Climate Observing System (GCOS) as one of the 54 Essential Climate Variables used to characterize climate.
New maps, generated by a research team working as part of ESA’s Climate Change Initiative, provide a global view of above ground biomass distribution and spatial density over three separate years – 2010, 2017, and 2018. The maps are derived from a combination of data, depending on the year, from the Copernicus Sentinel-1 mission, Envisat’s ASAR instrument, and JAXA’s Advanced Land Observing Satellite (ALOS-1 and ALOS-2), along with additional information from Earth observation sources.
Enhancing Model Accuracy and Reducing Uncertainty
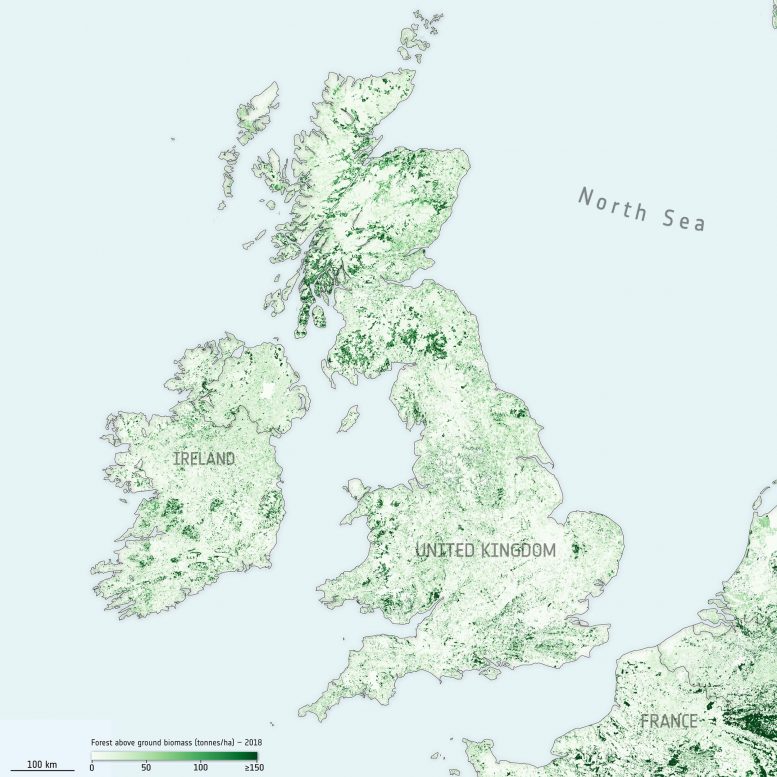
Earth observation data are routinely used to validate the accuracy, or identify biases, in climate models. The new maps, provided at 100 m resolution, have trimmed uncertainty estimates and will help to further constrain models.
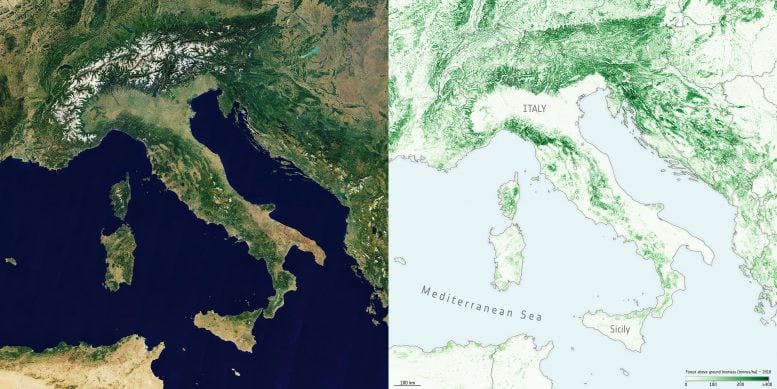
Crucially, and according to the team’s science leader, Shaun Quegan, the new maps capture the higher biomass levels in high-density forest areas, such as in the tropics, due to major improvements to the algorithm.
By using a globally consistent retrieval methodology, the advent of multi-year biomass maps brings the prospect of monitoring changes a step closer to reality. However, users are currently discouraged from quantifying biomass changes just by subtracting the current maps, since the retrieval procedure is still being tuned to account for the different mission and sensor observations used in their generation.
The team is currently developing a map for 2020 while also addressing temporal consistency between the different years, with the integration of additional low geometric resolution data streams under consideration, namely L-band vegetation optical depth from ESA’s Soil Moisture and Ocean Salinity (SMOS) satellite and scatterometer data from the ASCAT on board Eumetsat’s Metop satellites.
Shaun Quegan explains, “Combining these new data is anticipated to increase the consistency of these high-resolution maps, and move a step closer towards tracking changes and direct estimation of gross gains and losses of above ground biomass at scale.” Alternative approaches to correct for bias are also under investigation.
Tracking Change and Supporting Policy
With a decade of global biomass estimates on the horizon, the maps are set to allow scientists to undertake trend analyses, allowing, for example, the impact of regional climate phenomena such as El Niño on biomass dynamics to be better understood.
Significantly, an ability to track global biomass change is set to support global and national policy aimed at meeting emission reduction commitments to limit global warming. Biomass estimates provide critical support for both reporting of national greenhouse emissions under the Paris Agreement and for forest management through the United Nations’ Reducing Emissions from Deforestation and Degradation-plus (REDD+) initiative.
Tracking biomass change is also becoming increasingly important as national governments work towards reporting for the Global Stocktake – an aspect of the global Paris climate deal – that will periodically check international progress towards meeting emissions reduction commitments to limit global warming.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.