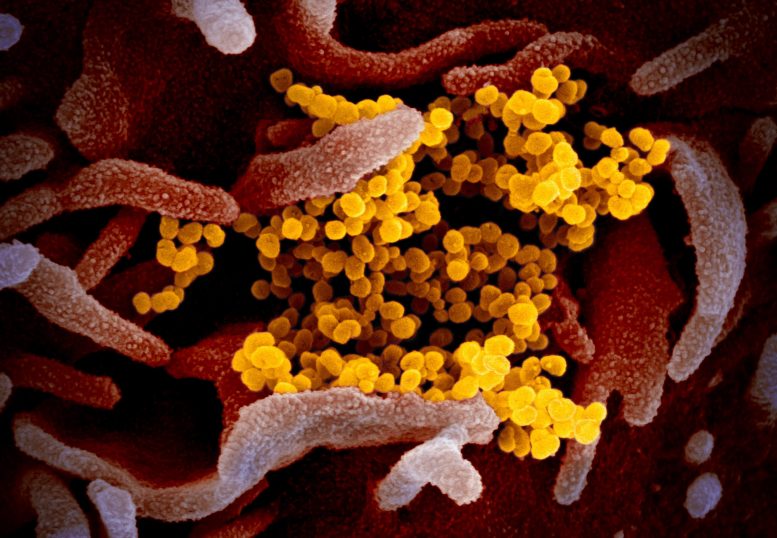
Study finds specific cells in the lungs, nasal passages, and intestines that are more susceptible to infection.
Researchers at MIT; the Ragon Institute of MGH, MIT, and Harvard; and the Broad Institute of MIT and Harvard; along with colleagues from around the world have identified specific types of cells that appear to be targets of the coronavirus that is causing the COVID-19 pandemic.
Using existing data on the RNA found in different types of cells, the researchers were able to search for cells that express the two proteins that help the SARS-CoV-19 virus enter human cells. They found subsets of cells in the lung, the nasal passages, and the intestine that express RNA for both of these proteins much more than other cells.
The researchers hope that their findings will help guide scientists who are working on developing new drug treatments or testing existing drugs that could be repurposed for treating COVID-19.
“Our goal is to get information out to the community and to share data as soon as is humanly possible, so that we can help accelerate ongoing efforts in the scientific and medical communities,” says Alex K. Shalek, the Pfizer-Laubach Career Development Associate Professor of Chemistry, a core member of MIT’s Institute for Medical Engineering and Science (IMES), an extramural member of the Koch Institute for Integrative Cancer Research, an associate member of the Ragon Institute, and an institute member at the Broad Institute.
Shalek and Jose Ordovas-Montanes, a former MIT postdoc who now runs his own lab at Boston Children’s Hospital, are the senior authors of the study, which appears today in Cell. The paper’s lead authors are MIT graduate students Carly Ziegler, Samuel Allon, and Sarah Nyquist; and Ian Mbano, a researcher at the Africa Health Research Institute in Durban, South Africa.
Digging into data
Not long after the SARS-CoV-2 outbreak began, scientists discovered that the viral “spike” protein binds to a receptor on human cells known as angiotensin-converting enzyme 2 (ACE2). Another human protein, an enzyme called TMPRSS2, helps to activate the coronavirus spike protein, to allow for cell entry. The combined binding and activation allows the virus to get into host cells.
“As soon as we realized that the role of these proteins had been biochemically confirmed, we started looking to see where those genes were in our existing datasets,” Ordovas-Montanes says. “We were really in a good position to start to investigate which are the cells that this virus might actually target.”
Shalek’s lab, and many other labs around the world, have performed large-scale studies of tens of thousands of human, nonhuman primate, and mouse cells, in which they use single-cell RNA sequencing technology to determine which genes are turned on in a given cell type. Since last year, Nyquist has been building a database with partners at the Broad Institute to store a huge collection of these datasets in one place, allowing researchers to study potential roles for particular cells in a variety of infectious diseases.
Much of the data came from labs that belong to the Human Cell Atlas project, whose goal is to catalog the distinctive patterns of gene activity for every cell type in the human body. The datasets that the MIT team used for this study included hundreds of cell types from the lungs, nasal passages, and intestine. The researchers chose those organs for the COVID-19 study because previous evidence had indicated that the virus can infect each of them. They then compared their results to cell types from unaffected organs.
“Because we have this incredible repository of information, we were able to begin to look at what would be likely target cells for infection,” Shalek says. “Even though these datasets weren’t designed specifically to study COVID, it’s hopefully given us a jump start on identifying some of the things that might be relevant there.”
In the nasal passages, the researchers found that goblet secretory cells, which produce mucus, express RNAs for both of the proteins that SARS-CoV-2 uses to infect cells. In the lungs, they found the RNAs for these proteins mainly in cells called type II pneumocytes. These cells line the alveoli (air sacs) of the lungs and are responsible for keeping them open.
In the intestine, they found that cells called absorptive enterocytes, which are responsible for the absorption of some nutrients, express the RNAs for these two proteins more than any other intestinal cell type.
“This may not be the full story, but it definitely paints a much more precise picture than where the field stood before,” Ordovas-Montanes says. “Now we can say with some level of confidence that these receptors are expressed on these specific cells in these tissues.”
Fighting infection
In their data, the researchers also saw a surprising phenomenon — expression of the ACE2 gene appeared to be correlated with activation of genes that are known to be turned on by interferon, a protein that the body produces in response to viral infection. To explore this further, the researchers performed new experiments in which they treated cells that line the airway with interferon, and they discovered that the treatment did indeed turn on the ACE2 gene.
Interferon helps to fight off infection by interfering with viral replication and helping to activate immune cells. It also turns on a distinctive set of genes that help cells fight off infection. Previous studies have suggested that ACE2 plays a role in helping lung cells to tolerate damage, but this is the first time that ACE2 has been connected with the interferon response.
The finding suggests that coronaviruses may have evolved to take advantage of host cells’ natural defenses, hijacking some proteins for their own use.
“This isn’t the only example of that,” Ordovas-Montanes says. “There are other examples of coronaviruses and other viruses that actually target interferon-stimulated genes as ways of getting into cells. In a way, it’s the most reliable response of the host.”
Because interferon has so many beneficial effects against viral infection, it is sometimes used to treat infections such as hepatitis B and hepatitis C. The findings of the MIT team suggest that interferon’s potential role in fighting COVID-19 may be complex. On one hand, it can stimulate genes that fight off infection or help cells survive damage, but on the other hand, it may provide extra targets that help the virus infect more cells.
“It’s hard to make any broad conclusions about the role of interferon against this virus. The only way we’ll begin to understand that is through carefully controlled clinical trials,” Shalek says. “What we are trying to do is put information out there, because there are so many rapid clinical responses that people are making. We’re trying to make them aware of things that might be relevant.”
Shalek now hopes to work with collaborators to profile tissue models that incorporate the cells identified in this study. Such models could be used to test existing antiviral drugs and predict how they might affect SARS-CoV-2 infection.
The MIT team and their collaborators have made all the data they used in this study available to other labs who want to use it. Much of the data used in this study was generated in collaboration with researchers around the world, who were very willing to share it, Shalek says.
“There’s been an incredible outpouring of information from the scientific community with a number of different parties interested in contributing to the battle against COVID in any way possible,” he says. “It’s been incredible to see a large number of labs from around the world come together to try and collaboratively tackle this.”
The research was funded by the Searle Scholars Program, the Beckman Young Investigator Program, the Pew-Stewart Scholars Program for Cancer Research, a Sloan Fellowship in Chemistry, the National Institutes of Health, the Aeras Foundation, the Bill and Melinda Gates Foundation, the Richard and Susan Smith Family Foundation, the National Institute of General Medical Sciences, the UMass Center for Clinical and Translational Science Project Pilot Program, the MIT Stem Cell Initiative, Fondation MIT, and the Office of the Assistant Secretary of Defense for Health Affairs.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.