
Research Could Provide Clues to Extraterrestrial Life
A team of Rutgers University scientists dedicated to pinpointing the primordial origins of metabolism – a set of core chemical reactions that first powered life on Earth – has identified part of a protein that could provide scientists clues to detecting planets on the verge of producing life.
The research, published on March 10 in the journal Science Advances, has important implications in the search for extraterrestrial life because it gives researchers a new clue to look for, said Vikas Nanda, a researcher at the Center for Advanced Biotechnology and Medicine (CABM) at Rutgers.
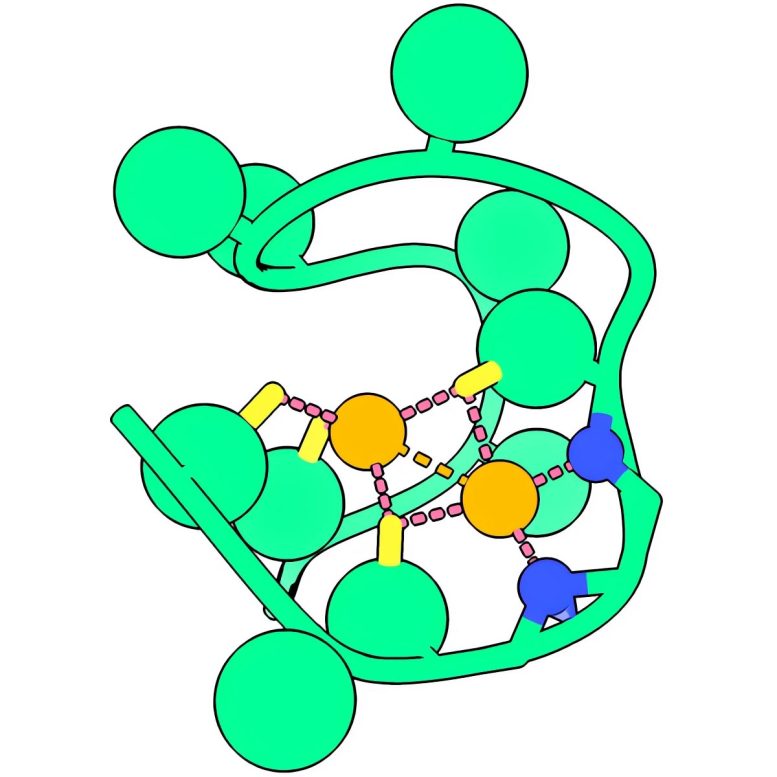
Based on laboratory studies, Rutgers scientists say one of the most likely chemical candidates that kickstarted life was a simple peptide with two nickel atoms they are calling “Nickelback” not because it has anything to do with the Canadian rock band, but because its backbone nitrogen atoms bond two critical nickel atoms. A peptide is a constituent of a protein made up of a few elemental building blocks known as amino acids.
“Scientists believe that sometime between 3.5 and 3.8 billion years ago there was a tipping point, something that kickstarted the change from prebiotic chemistry – molecules before life – to living, biological systems,” Nanda said. “We believe the change was sparked by a few small precursor proteins that performed key steps in an ancient metabolic reaction. And we think we’ve found one of these ‘pioneer peptides’.”
NASA’s Quest for Biosignatures
The scientists conducting the study are part of a Rutgers-led team called Evolution of Nanomachines in Geospheres and Microbial Ancestors (ENIGMA), which is part of the Astrobiology program at NASA. The researchers are seeking to understand how proteins evolved to become the predominant catalyst of life on Earth.
When scouring the universe with telescopes and probes for signs of past, present or emerging life, NASA scientists look for specific “biosignatures” known to be harbingers of life. Peptides like nickelback could become the latest biosignature employed by NASA to detect planets on the verge of producing life, Nanda said.
An original instigating chemical, the researchers reasoned, would need to be simple enough to be able to assemble spontaneously in a prebiotic soup. But it would have to be sufficiently chemically active to possess the potential to take energy from the environment to drive a biochemical process.
To do so, the researchers adopted a “reductionist” approach: They started by examining existing contemporary proteins known to be associated with metabolic processes. Knowing the proteins were too complex to have emerged early on, they pared them down to their basic structure.
After sequences of experiments, researchers concluded the best candidate was Nickelback. The peptide is made of 13 amino acids and binds two nickel ions.
Hydrogen and Nickel: The Spark of Life
Nickel, they reasoned, was an abundant metal in early oceans. When bound to the peptide, the nickel atoms become potent catalysts, attracting additional protons and electrons and producing hydrogen gas. Hydrogen, the researchers reasoned, was also more abundant on early Earth and would have been a critical source of energy to power metabolism.
“This is important because, while there are many theories about the origins of life, there are very few actual laboratory tests of these ideas,” Nanda said. “This work shows that, not only are simple protein metabolic enzymes possible, but that they are very stable and very active – making them a plausible starting point for life.”
Reference: “Design of a minimal di-nickel hydrogenase peptide” by Jennifer Timm, Douglas H. Pike, Joshua A. Mancini, Alexei M. Tyryshkin, Saroj Poudel, Jan A. Siess, Paul M. Molinaro, James J. McCann, Kate M. Waldie, Ronald L. Koder, Paul G. Falkowski and Vikas Nanda, 10 March 2023, Science Advances.
DOI: 10.1126/sciadv.abq1990
Other Rutgers researchers on the study include: Distinguished Professor Paul Falkowski and Jennifer Timm, a postdoctoral associate, in the Environmental Biophysics and Molecular Ecology Program, Department of Marine and Coastal Sciences at the School of Environmental and Biological Sciences; Joshua Mancini, Douglas Pike, Saroj Poudel and Alexei Tyryshkin, postdoctoral associates, and doctoral student Jan Siess at the Center for Advanced Biotechnology and Medicine and in the Department of Biochemistry and Molecular Biology at Robert Wood Johnson Medical School; and Kate Waldie, an assistant professor of the Department of Chemistry and Chemical Biology at the School of Arts and Sciences.
Researchers from the City College of New York also participated in the study.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
4 Comments
Where did the peptide come from?
It formed in the oceans from simpler constituents.
Yes, an advance 2021 was the discovery that Hadean acidic vents should have been able to produce random proteins, long thought to be too energetic demanding. Earlier work had shown that the first 10 or so amino acids that the genetic code evolved for should have been naturally produced, but this time they got many more – and a smidgen of polypeptides at the end.
[The Release of Energy During Protein Synthesis at Ultramafic-Hosted Submarine Hydrothermal Ecosystems,” was published in the journal JGR Biogeosciences.
https://agupubs.onlinelibrary.wiley.com/doi/10.1029/2021JG006436 ; the paper is targeting a vent community but IIRC the thermodynamical model is abiotic – I don’t have access right now.]
The simplest peptides likely evolved as RNA co-factors, where 1-2 amino acids suffice, and if having a prolongation machinery may detach as peptide strands. Nickelbacks are semi-ordered enzymes with a 4 amino acid core of the size as unordered catalyst “cages” of 4-5 amino acid strands, so there is a promising correspondence in the putative evolutionary results.