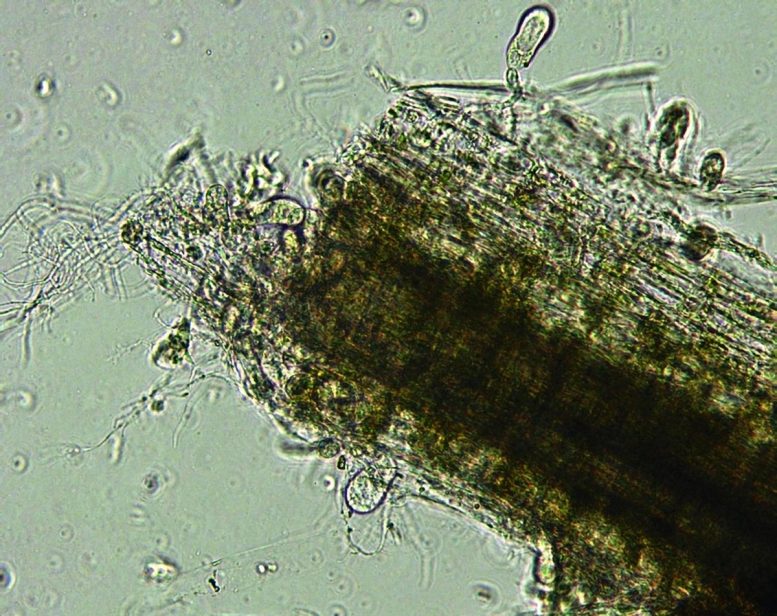

In an effort to replicate the cellulose-destroying capabilities of anaerobic fungi in yeast and create a potentially cost-effective single-organism process for manufacturing cellulosic biofuels, MIT biologists and genetic-sequencing experts have developed a novel analytical approach that will enable them to determine which enzymes in gut fungi are key players in plant digestion and how those enzymes are made and assembled.
MIT researchers are genetically engineering yeast to break down stubborn plant fibers into sugars that it can then ferment — a first step toward a potentially cost-effective single-organism process for manufacturing cellulosic biofuels. The source of the genetic material: anaerobic fungi from the digestive system of a horse on a hay-only diet. By teaming up with genetic-sequencing experts at MIT, the researchers have developed a novel analytical approach that will enable them to determine which enzymes in gut fungi are key players in plant digestion and how those enzymes are made and assembled — information that will help them replicate the fungi’s cellulose-destroying capabilities in yeast. Their approach could reveal new enzymes that are champion cellulose degraders but naturally occur only in unknown organisms that cannot be grown in a lab.
One promising source of renewable fuels is biomass from agricultural residues, grasses, and other dedicated energy crops. Methods exist to convert such cellulosic biomass into energy-rich fuels that could substitute for petroleum, particularly in transportation systems. In general, those methods use enzymes and chemicals to break plant biomass down into sugars such as glucose, which are then fed to a microorganism to ferment the sugars to make ethanol, butanol and other end products. But such methods are not economically viable for widespread use, largely due to the expense associated with enzyme production and activity and the costs associated with separating cellulosic processing into so many steps. Chris Kaiser, MIT’s provost and the MacVicar Professor of Biology, believes that a better approach is to genetically engineer a single organism that can both break down the cellulose and ferment the product — and he believes that the best organism for the job is yeast. “We already know how to genetically modify yeast to control its metabolic pathways, and we have vast experience with using yeast to conduct fermentation on an industrial scale,” Kaiser says. The steps needed to implement his idea are known: Select an organism that naturally digests cellulose, identify the key enzymes involved in that process, determine which genes express those enzymes, retrieve those genes, and insert them into yeast. Such engineered yeast could readily be put into an industrial process and scaled up for biofuel production.
Researchers in Kaiser’s lab know yeast: They use it as a model organism for studying many fundamental cellular processes. But genetically engineering yeast to process cellulosic biomass was a novel concept and a new direction for them. Says Kaiser, the idea “was sparked by MIT Energy Initiative seed funding and has blossomed into a broader MIT research endeavor and a new way of thinking about finding improved organisms and enzymes for use in manufacturing biofuels.”
Looking for enzymes
The first task was to find cellulose-degrading, or “cellulolytic,” enzymes that would work well in yeast. A good place to look is in the digestive tracts of large herbivores such as cows and horses. Such animals consume large quantities of grass and hay, and parts of their digestive tract contain a mixture of bacteria, fungi and protozoa optimized to digest those difficult materials.
The idea of seeking cellulolytic enzymes in the rumen or hindgut is not new, but usually the focus is on bacteria — the workhorse of the industrial microbiology world. However, genes from bacteria generally will not express enzymes efficiently in yeast. Genes from fungi would work better, but relatively little is known about the fungi in large mammalian herbivores. A widely accepted view is that they are present in very low concentrations in the digestive tract of large herbivores, so they cannot be critical players in cellulose digestion. In addition, they are extremely difficult to isolate and culture in a lab since they require carefully controlled growing conditions, including the absence of oxygen. Expose this “obligate anaerobe” to oxygen and it dies. The result: anaerobic fungi have largely been ignored.

Kaiser and his colleague Michelle O’Malley, USDA-NIFA Postdoctoral Fellow in biology and now an assistant professor of chemical engineering at the University of California, Santa Barbara, argue that these fungi should not be dismissed so readily. For one thing, the analyses that detect low concentrations of gut fungi test only liquid samples. “But the anaerobic fungi we’re interested in are almost always found burrowed into the cellulosic fibers they’re digesting,” says O’Malley. Indeed, analyses that include both liquid and solid fractions show that fungi can be up to 20% of the microbial population.
Moreover, those fungi are thought to be among the most efficient and robust known digesters of “lignocellulosic” material — that is, not only cellulose but also other materials that occur with it, notably hemicellulose and lignin, polymers in plant cell walls that are extremely tough and can block access to the cellulose itself, further complicating its breakdown. In anaerobic fungi, the activity of key enzymes is aided by the action of mycelium—branching, thread-like filaments that can penetrate the plant material and break it up mechanically. While fungi may not be the dominant microbial population in the digestive tract, they clearly play a major role in degrading cellulose in the gut.
Given anaerobic fungi’s remarkable capabilities and their potential compatibility with yeast, Kaiser and O’Malley decided that they should learn to work with them. In winter 2011, O’Malley spent five weeks at a laboratory in the United Kingdom studying with Professor Michael Theodorou, now of Durham University and the Center for Process Innovation, UK — one of a “handful of people in the world” who could teach her the skills and techniques needed to isolate, classify, and culture these challenging organisms.
First steps, new challenges
To start out, the MIT team aimed simply to determine whether genes from fungi would express cellulolytic enzymes in yeast. For the screening, O’Malley inserted known cellulolytic genes from the best-understood type of ruminal fungus, Piromyces, into yeast, where they grew and expressed the enzymes of interest at high levels. However, some of the expressed enzymes were not reactive toward cellulose, and it was difficult to determine why not. O’Malley and Kaiser agreed that they needed to better understand how the cellulolytic enzymes are made and operate in their native environment — that is, inside Piromyces.
In fact, for the best outcome, they needed to understand that process in detail. It takes at least three types of enzymes to cut cellulose polymers into shorter segments. If those enzymes acted by random collision with the cellulosic material, the process would be extremely inefficient. To speed things up, over millions of years, gut bacteria and fungi learned to assemble groups of enzyme-producing genes on a supporting molecule called a scaffoldin. Kaiser likens this overall structure — the cellulosome — to a workbench on which the genes are tethered and spaced so that the expressed enzymes can degrade all components of the cellulose fibers synergistically.
Ideally, the researchers would replicate the assembly and operation of such fungal cellulosomes inside yeast. But first they needed to define the composition and structure of complete cellulosomes in Piromyces — a task best accomplished in a pure sample of the fungus.
To prepare such a sample, O’Malley collected feces from a horse called Finn at Verrill Farm Stables in Concord, Mass.. Finn eats only grass and hay, so his digestive system is well-tuned for cellulose destruction and his feces are high in cellulose-degrading microbes. To begin, O’Malley grew samples on grass plus antibiotics to suppress the growth of both bacteria and bacteria-consuming protozoa. She then withdrew a small sample from the mixture and grew it in the presence of cellulosic substrates. Performing that process several times over two months yielded an “isolate” — a single type of fungus that appears to be a new species of Piromyces, never before characterized or named.
To search for cellulolytic enzymes and cellulosomes in the new species, the researchers are exploiting a natural process called catabolic regulation. In general, organisms will not expend the energy to produce enzymes for digesting difficult materials like cellulose if they have available something easier to digest, such as glucose. “So we can get the organism to tell us what its most valuable enzymes are for breaking down cellulose,” says O’Malley. Accordingly, she is growing some fungi on glucose — a setup that should turn off the genes required for cellulose destruction. Other samples get only cellulose, which should switch on all those genes for maximum expression. The cultures that result will give her a “logical, focused way to hunt for the actual enzymes and cellulosomes that are important to the organism for degrading cellulose,” says O’Malley — perhaps including some that are new and radically different from those known to date.
Genetic analysis
The next step — now ongoing — is to define the important genes and groups of genes and where they occur on the DNA of the organism. The good news, says Kaiser, is that he has MIT colleagues nearby who are among the world’s leading experts at accomplishing those tasks. Researchers in the MIT BioMicro Center are performing next-generation genome sequencing of O’Malley’s samples, specifically, of the messenger RNA (mRNA), which are copies of the sections of the DNA responsible for making enzymes. However, the genetic readings will represent isolated snippets of mRNA rather than full genes, so another task is to figure out which snippets belong together. At the Broad Institute of MIT and Harvard, Aviv Regev, associate professor of biology, and her team have developed new “assembly algorithms” that are ideally suited to performing that reconstruction. “Taken together, these methods will give us a concise inventory of what’s there because the mRNA copies only the part of the DNA needed to make the enzyme,” says Kaiser. “And the number of copies present tells you something about the level of expression. There should be many, many more copies of the mRNA corresponding to the cellulolytic gene in the sample growing on cellulose than in the sample growing on glucose. It’s a very powerful approach.”
One more technique being brought to bear is cluster analysis. A computer can be programmed to examine the millions of data points from genetic studies and look for correlations in expression levels. With no regard for other factors, it simply groups genes that are expressing at the same level — an indication that those genes are being “co-regulated” by the organism and are therefore likely to function together on some level. Based on that assumption, a group including known cellulolytic genes could also include unknown ones, among them perhaps some components of the cellulosome.
While O’Malley is studying the genes in a single pure fungus, others in Kaiser’s lab are using similar methods to quantify the expression of specific gene families in complex mixtures that are good at digesting cellulose. From there, they can extract novel genetic sequences of interest, insert them into yeast, and examine the enzymatic activity that results — a method of discovering promising new genes without having to identify or culture their parent organisms.
“The beauty of this approach is that we could find cellulolytic genes that come from organisms we can’t grow in the laboratory,” says Kaiser. “This is one of the exciting new areas in microbiology. It’s a way of circumventing the limitation of the classical approach in which you can only study things you can culture — and that’s just a small fraction of the living things that are out there.” Taken together, the new MIT activities should expand the list of known organisms and enzymes that degrade plant material into fermentable products — whether in the digestive system of a horse, in the muck on the floor of a pond, or in a biofuels manufacturing plant.
Reference: “Evaluating expression and catalytic activity of anaerobic fungal fibrolytic enzymes native topiromyces sp E2 inSaccharomyces cerevisiae” by Michelle A. O’Malley, Michael K. Theodorou and Chris A. Kaiser, 30 December 2011, Environmental Progress and Sustainable Energy.
DOI: 10.1002/ep.10614
This research was supported initially by a seed grant from the MIT Energy Initiative. Postdoctoral fellow Michelle O’Malley was subsequently supported by a Whiting Foundation Fellowship and a US Department of Agriculture–National Institute of Food and Agriculture Postdoctoral Fellowship. A travel fellowship from the Company of Biologists supported her work with Professor Michael Theodorou at Durham University and the Center for Process Innovation in the United Kingdom.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
1 Comment
I hope you remember that teaching yeast to digest cellulose could create a nasty new plant pathogen, or worse, an organism capable living in soil and producing ethanol, deadly to plant roots or their symbiotic partners.