
Potential implications for American chestnut restoration and blight-resistance.
The chromosomes of American and Chinese chestnuts are not so similar after all, at least in one key region of the genome – the nucleolus organizing region (NOR).
The finding, published today (January 15) in an article in Scientific Reports, has major implications for anyone with the goal of conferring blight-resistance to American chestnuts through hybridization with the Chinese chestnut.
“This is an unprecedented finding in the field of plant cytology,” says Nurul Faridi, a Forest Service geneticist and lead author of the study.
Understanding Backcross Breeding
Traditional backcross breeding, involving hybridization between two species, aims to combine an ideal mix of traits from two species without genetic engineering. Backcross breeding can only succeed when the chromosomes of both species are compatible. Because Chinese-American chestnut hybrids are viable, people have assumed that the two species are highly compatible. But the new study reveals significant differences in the NOR of the two species.

The NOR’s Role
The NOR is part of every plant and animal cell. It carries the genetic instructions for making ribosomes – the molecular machines that make the proteins essential for life.
The NOR is located near the end of the short arm of a particular chromosome. It is present on both species, but in Chinese chestnut it is packed with a type of DNA known as heterochromatin and constitutes about 25% of the chromosome. The structure and composition of this DNA surprised the researchers – it is highly condensed, lacking gene content, and transcriptionally inactive. In contrast, the American chestnut satellite is very small and appeared to be euchromatic. Euchromatic regions of DNA are transcriptionally active.
Revealing the Discovery
Faridi first noticed a small pair of Chinese chestnut chromosomes exhibiting very bright fluorescence with a specialized microscope, a UV filter, and a dye that binds to the DNA.
Faridi used a technique called fluorescent in situ hybridization (FISH) to further analyze the discovery.
“Our high-quality FISH images provide unequivocal evidence of this unique DNA arrangement,” says Faridi. “These images are not just pictures; they are a testament to the dynamic nature of genetic material.”
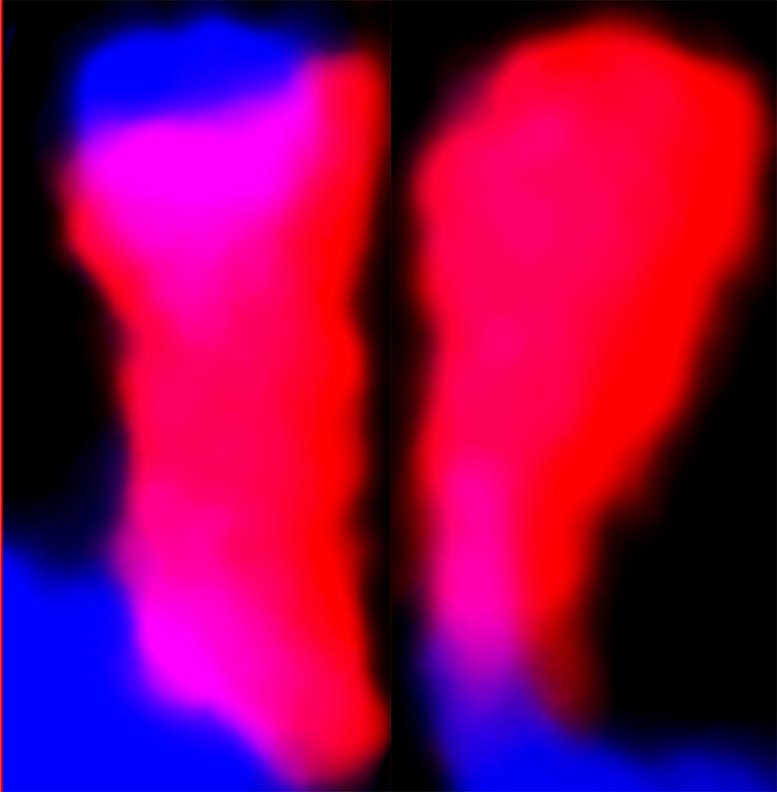
Faridi has been working with FISH since 1991 and has extensive experience preparing plant chromosomes for analyses. Well-separated chromosomes from enzymatically digested root tips that are mostly free of cell walls, nuclear membranes, and cytoplasmic debris are best for FISH.
Most FISH imagery is obtained from animal cells, as plant cells, and especially trees, are more challenging to work with. Faridi has found that chestnuts are far more difficult to work with than pine and poplar.
The Road Ahead
The researchers will use a technique called oligonucleotide FISH for further investigation. Oligo-FISH uses short specific DNA probes acquired from DNA sequencing. Since the entire genomes of American and Chinese chestnut have been sequenced, oligo-FISH will allow the researchers to make detailed genetic studies that will discern subtle genomic differences. The technique is especially useful for studying hybrids since it can indicate which parent a gene is from.
Progress Towards Blight Resistance
The progress in developing American chestnut hybrids with height of the American chestnut and the blight-resistance of the Chinese chestnut has been significant. However, the most advanced hybrids do not currently have enough blight resistance for restoration, as previous Forest Service research has shown.
Reference: “Cyto-molecular characterization of rDNA and chromatin composition in the NOR-associated satellite in Chestnut (Castanea spp.)” by Nurul Islam-Faridi, George L. Hodnett, Tetyana Zhebentyayeva, Laura L. Georgi, Paul H. Sisco, Frederick V. Hebard and C. Dana Nelson, 15 January 2024, Scientific Reports.
DOI: 10.1038/s41598-023-45879-6
In addition to Faridi and C. Dana Nelson of the USDA Forest Service Southern Research Station, the genetics team included researchers from Texas A&M University, Pennsylvania State University, University of Kentucky, and The American Chestnut Foundation.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.