
Scientists have traced the origins of all complex life to Asgard archaea, ancient microbes that include our closest evolutionary relatives. Genetic evidence suggests that these microorganisms developed traits that paved the way for eukaryotic life, revealing a crucial step in life’s evolution.
The mythological Norse god Thor hails from the celestial city of Asgard, and according to revolutionary research published in the scientific journal, Nature, he’s not the only Asgardian. This new research suggests that we humans — along with eagles, starfish, daisies, and every complex organism on Earth — are, in a sense, Asgardians.
The research team at The University of Texas at Austin, along with collaborators from different institutions, conducted a genomic analysis of several hundreds of microorganisms known as archaea. Their findings revealed that eukaryotes – complex life forms with nuclei in their cells, including all flora, fauna, insects, and fungi across the globe – can trace their origins back to a common Asgard archaean ancestor.
A Common Ancestor for Complex Life
That means eukaryotes are, in the parlance of evolutionary biologists, a “well-nested clade” within Asgard archaea, similar to how birds are one of several groups within a larger group called dinosaurs, sharing a common ancestor. The team has found that all eukaryotes share a common ancestor among the Asgards.
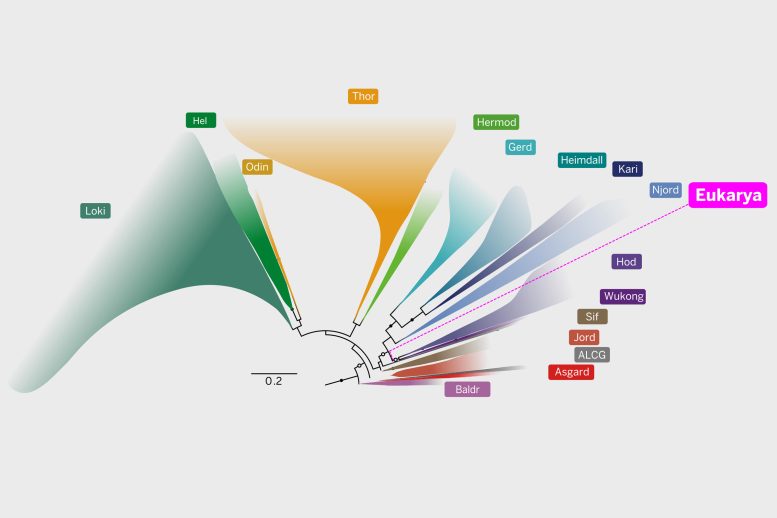
No fossils of eukaryotes have been found from farther back than about 2 billion years ago, suggesting that before that, only various types of microbes existed.
“So, what events led microbes to evolve into eukaryotes?” said Brett Baker, UT Austin associate professor of integrative biology and marine science. “That’s a big question. Having this common ancestor is a big step in understanding that.”
Led by Thijs Ettema of Wageningen University in the Netherlands, the research team identified the closest microbial relative to all complex life forms on the tree of life as a newly described order called the Hodarchaeales (or Hods for short). The Hods, found in marine sediments, are one of several subgroups within the larger group of Asgard archaea.
The Asgard archaea evolved more than 2 billion years ago, and their descendants are still living. Some have been discovered in deep-sea sediments and hot springs around the world, but so far only two strains have been successfully grown in the lab. To identify them, scientists collect their genetic material from the environment and then piece together their genomes. Based on genetic similarities with other organisms that can be grown in the lab and studied, the scientists can infer metabolism and other features of the Asgards.
Reconstructing the Dawn of Complex Life
“Imagine a time machine, not to explore the realms of dinosaurs or ancient civilizations, but to journey deep into the potential metabolic reactions that could have sparked the dawn of complex life,” said Valerie De Anda, a researcher in Baker’s lab. “Instead of fossils or ancient artifacts, we look at the genetic blueprints of modern microbes to reconstruct their past.”
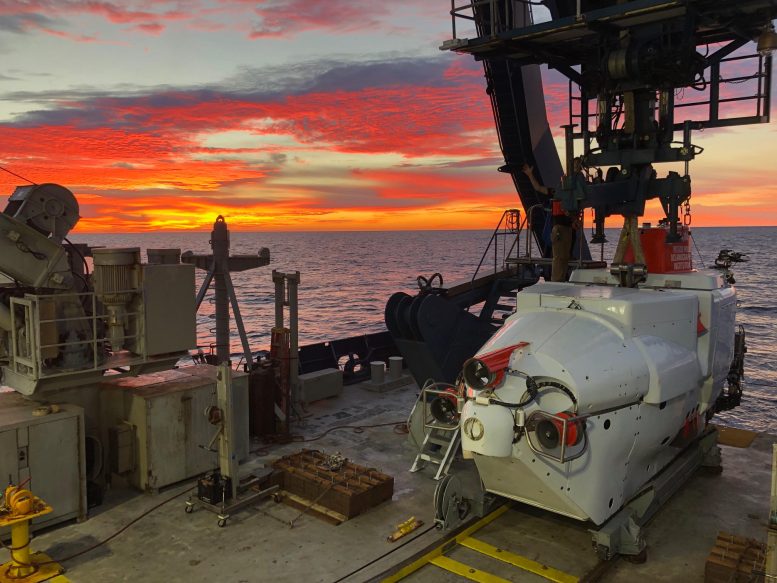

The researchers expanded the known Asgard genomic diversity, adding more than 50 undescribed Asgard genomes as input for their modeling. Their analysis indicates that the ancestor of all modern Asgards appears to have been living in hot environments, consuming CO2 and chemicals to live. Meanwhile, Hods, which are more closely related to eukaryotes, are metabolically more similar to us, eating carbon and living in cooler environments.
“This is really exciting because we are looking for the first time at the molecular blueprints of the ancestor that gave rise to the first eukaryotic cells,” De Anda said.
In Norse mythology, Hod (also spelled Höd, Höðr or Hoder) is a god, the blind son of Odin and Frigg, who is tricked into killing his own brother Baldr.
“I keep joking in my talks that ‘We are all Asgardian’,” Baker said. “Now that’s probably going to be on my tombstone.”
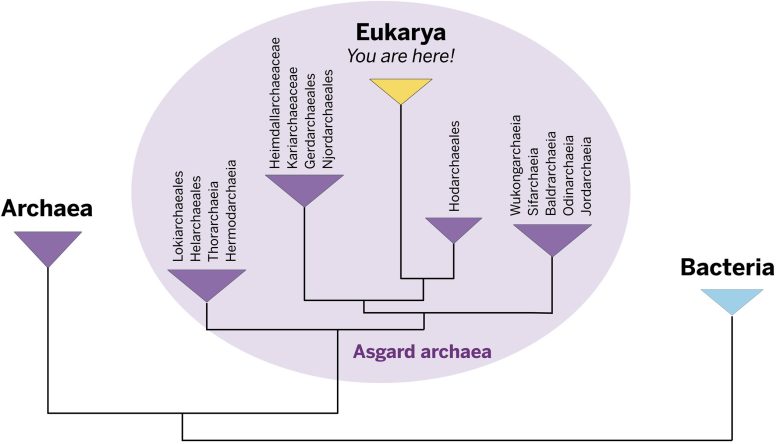
“To me, the most exciting thing is that we’re starting to see the transition from what biologists think is an archaeon to this organism Hodarchaeales that is more like a eukaryote,” Baker explained. “Another way to put it is that these Hods are our sister group in the archaeal world.”
How Gene Duplication Drove Evolution
Baker said it makes sense that of all the archaea, the Asgards are the ones that spawned eukaryotes. Like eukaryotes, members of the Asgard archaea have many genes with multiple copies in their genomes. In eukaryotes, when genes became duplicated, the new copies often took on new functions, giving organisms new abilities. It was one of the big drivers of evolution.
“We don’t know, in these Asgards specifically, what the gene duplications led to,” Baker said. “But we know in eukaryotes that gene duplications led to new functions and an increasing of cellular complexity. So, we think that that’s one of the ways that Asgards led to the innovations that define eukaryotes.”
Scientists studying archaea have found many proteins that were once thought to be exclusive to eukaryotes. Baker said that raises the question: What functions are these eukaryotic proteins serving in the archaea?
“I think studying these simpler forms of life and their eukaryotic characteristics is going to tell us a lot about ourselves,” Baker said.
Reference: “Inference and reconstruction of the heimdallarchaeial ancestry of eukaryotes” by Laura Eme, Daniel Tamarit, Eva F. Caceres, Courtney W. Stairs, Valerie De Anda, Max E. Schön, Kiley W. Seitz, Nina Dombrowski, William H. Lewis, Felix Homa, Jimmy H. Saw, Jonathan Lombard, Takuro Nunoura, Wen-Jun Li, Zheng-Shuang Hua, Lin-Xing Chen, Jillian F. Banfield, Emily St John, Anna-Louise Reysenbach, Matthew B. Stott, Andreas Schramm, Kasper U. Kjeldsen, Andreas P. Teske, Brett J. Baker and Thijs J. G. Ettema, 14 June 2023, Nature.
DOI: 10.1038/s41586-023-06186-2
Support for this research was provided by the Origin of Eukaryotes program at the Moore and Simons Foundations, U.S. National Science Foundation, the Wellcome Trust Foundation, the European Research Council, the Swedish Research Council, the Dutch Research Council, the National Natural Science Foundation of China, the Wenner-Gren Foundation, the Science for Life Laboratory (Sweden) and the European Commission’s Marie Skłodowska-Curie Actions.
Other authors from UT Austin are Kiley W. Seitz and Nina Dombrowski. In addition to Ettema, authors from other institutions are Laura Eme, Daniel Tamarit, Eva Caceres, Courtney Stairs, Max Schön, William Lewis, Felix Homa, Jimmy Saw, Jonathan Lombard, Takuro Nunoura, Wen-Jun Li, Zheng-Shuang Hua, Lin-Xing Chen, Jillian Banfield, Emily St. John, Anna-Louise Reysenbach, Matthew Stott, Andreas Schramm, Kasper Kjeldsen and Andreas Teske.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
34 Comments
Complex life is more than just eukaryotic. Connective tissues made higher forms of life possible. And that required the evolution of two amino acids not in the genetic code for collagens. Any non-metazoan ancestor would have to have had the ability to synthesize at least one. Do any of these asgardians? If not they are not viable.
Posttranslational modifications to non-proteogenic amino acids outnumber the 22 (or in metazoans, 21) proteogenic amino acids by many order of magnitudes. The two hydrolytic conversions of proline and lysine that are seen in collagen are key metabolites for building respectively destruction of amino acids, and collagens evolved in many eukaryotes and prokaryotes alike.
But what has later metazoan evolution to do with stem eukaryote evolution!?
To me where evolution falls apart is this . You take the newest great ape ancestor . The paranthropus boisei . Wich has sagital crest ..on skull that remotely looks human ..and take the oldest known human in the homo genus the homo eregastor that looks remotely human and could be the evil twin of the astraulopithsenes or paranthropus great ape crowd.
You stand them together they look like twins so evolutionists say yeah hey one led to another there verry close in looks . But in reality . These two look alikes phyosiplogy wise could never be futher apart . One is phyisiology is human ancestor and body is built like a human the orher is greeat ape and built like an ancient ape ..different skull different eyes different mussle type different vocal system different skeleton different skin and digestive tract . Completely different inside .so evolution telling me that in one evolution stage great apes went from apes to human phyisioligy …..in one generation if you believe that I gotta bridge to sell you I don’t care how close chimpanzees are DNA wise to people. They bodily function and material are not anything close to human anatomy yeah were smarter but in every other way primate modern apes would out class us in everything else strength speed healing digestive track night vision .five times our strength big fangs body hair aborial .lighting fast instincts .reflexes we git a lot weaker
The topic is about the origin of complex life, not evolution in general.
Evolution is an observed fact. It takes generations for populations to speciate, often geological times (i.e. millions of years). And that underlies the new result, again attesting to that fact.
Your many erroneous claims about apes, such as humans, are not relevant here.
Almost like we came from a pure form of creation when we were first designed like Adam and Eve.
The paper adds to the evidence for evolution.
Ok its about a discovery of 2 billion years old stuff. Nothing more nothing less👩🦰👩🏻🏫🤦🏼♀️🤷🏼♀️👣
When they say we are all Asgardian it is including the evolution of humans not just complex life.
Asgaard was actually areal place it was the centre of odinma in todays Finland. And it was the origin of mankind more than hundred million years ago. Source bocksaga
Every study I have seen says humans evolved in Africa then migrated outward across the globe.So why are “all Asgardians”and not all Africans.
Because you evidently did not actually read the article – if you did, I don’t think you understood it at all. The point went by MILES above your head.
Correct
Just because we may know how something came to be doesn’t mean we know why it came to be. The preconditions for protein formation and the actual encoding of a protein are so finely tuned, that the probability of one randomly coming into existence, given the entire age of the universe, is impossible without citing a supernatural cause.
In the beginning,God created the Heavens and the Earth. Genesis 1:1
The claims that protein formation and evolved protein metabolism is fine tuned is not in evidence. Chemists have now models that show that heavily hydrogenated acidic hydrothermal vents that produce amino acids are also capable of producing a modicum of proteins. And the evolution of genetically encoded proteins show that it is the opposite of finely tuned but capable of producing a plethora of proteins.
And all that are results of observed natural processes.
But please, refrain from pushing your magical and irrelevant rites onto an uninterested public.
If we all just believed that we are all gods one form or another and spent more time spreading love and joy to each and everyone I about bet that it doesn’t matter about the rest of the stuff at all.
Archeological discovery of Ardi (ARA-VP-6/500) a fossilized skeletal remains of an Ardipithecus ramidus, an early human-like female anthropoid, and upon examination the skeletal remains has been dated to be 4.4 million years old. There is so much-needed information being uncovered.
Here is where Paganism starts standing up tall. You start this writing off with the story of a pagan god, which most people don’t realize are false entities backed by evil spirits. And because people give credence to these evil spirits and call them gods while denying the one and only true living God, our creator whose name is YHWH, God of Abraham, Isaac and Jacob, who sent his son Jesus the embodiment of God himself, this is when our land suffers and the people of the earth experience all types of lies and disgusting abominable practices…too much to write about here. So how then can your “research” and your “discoveries” even be trusted when you start out with a credence or reverence to a lie? Let light shine brighter than darkness and give credit where credit is due, which is to our heavenly Father, who created all things, including this organism you claim to have discovered. Science knows that God exist because science coincides with His creation. Express Truth because our world is suffering….from lies.
This is about observed evolution and not about the purloined myths that went into naming the specific archaea. The first Asgard archaea was discovered by metagenomic methods around the Loki’s Castle hydrothermal vent, and it was also humoring the complex evolution observed.
“In the summer of 2010, sediments were analysed from a gravity core taken in the rift valley on the Knipovich ridge in the Arctic Ocean, near the Loki’s Castle hydrothermal vent site. Specific sediment horizons previously shown to contain high abundances of novel archaeal lineages were subjected to metagenomic analysis.[8][9] In 2015, an Uppsala University-led team proposed the Lokiarchaeota phylum based on phylogenetic analyses using a set of highly conserved protein-coding genes.[10] The group was named for the shape-shifting Norse god Loki, in an allusion to the hydrothermal vent complex from which the first genome sample originated.[11] The Loki of mythology has been described as “a staggeringly complex, confusing, and ambivalent figure who has been the catalyst of countless unresolved scholarly controversies”,[12] analogous to the role of Lokiarchaeota in the debates about the origin of eukaryotes.”
A well named group of archaea, if you ask me.
God did not create man. Man created “gods” in an effort to explain things primitive man did not understand.When you think about it, we as children all had beliefs in mythological beings. Then we evolved and grew, and realized that things much more believable than god, like Santa Clause, the Easter Bunny, and The Tooth Fairy, were all stories just to influence children to behave “properly” much like the stories in the greatest work of fiction the world has ever known, THE BIBLE. Some day you Bible thumpers will also evolve.
The more i hear billions of years the more dumb i think scientist are. We have nothing that can date anything that old because natural disasters destroy the fabric to date. To say billions of years is laughable. Nothing here has been around that long.
The whole Earth has been around that long, and many rocks with fossils as well as life with genetic fossils shared by common ancestors. Those two humongous data sets, in combination with biogeographical evidence – species radiations from locales out – are the three pillars of evolution science.
FWIW, as a common study in bioinformatics you can pull out the genetic transcription factors of bacteria, archaea and eukaryotes and – consistent with the article – show that eukaryote genetic machinery evolved from the archaeal version. (With the bacterial version a distant relative.) That is data covering 1-2 billion years of evolution, which anyone can pull from open access databases such as NCBI and observe such old molecules (and test evolution at the same time). There are no excuses to laugh at what you are too lazy to see for yourself, these sites teach you how to do it within hours.
The asgard archaeology The Nature article states that two strains have been produced in lab conditions.Is this living cells?
Yes.
“The God of peace will soon crush Satan under your feet. May the grace of our Lord Jesus be with you.”
Romans 16:20 NLT
🤣
Another scenario I’d like to bring the attention is crude oil‘s just said to come from fossil fuels. That’s the biggest joke scam I’ve ever heard just about my life gotcha. It’s coming from the government clothes come from the core of the planet naturally things like swamps in in Meadows can overtime Create truth oils it’s pretty simple and logical. They got everybody buffalo didn’t believe in true laurels come from fossil fuels. It’s the biggest scam I’ve ever heard just like the greenhouse effect. Government says there’s depleting ozone air from things like the suns rays but more importantly, we humans here on earth or killing off the ozone layer with carbon dioxide carbon dioxide, aerosol cans, carbon carbon oxide exhaling too much and cows farting. That’s the biggest scam I’ve ever heard it’s another way from the taxes. The only thing depleting the ozone layer is the suns rays and that’s it , I could go on and on about things like going to the moon man man stepping foot on the moon and man, not even being able to get to the depths of our deepest waters here on the planet and they said we’ve been another planet I find that very hard to believe
Conspiracy theory is the opposite of fact.
This is rather old news.. many publications since 2017… Pretty sure PBS Eons aired an episode all about it just last year?
It’s a deeper analysis verifying and extending earlier results, yes.
There was another scientist who had said about origin of species much more clearly before more than 2500 years.that is Lord Buddha.If you study Thipitaka you will find more than this.believe me.
🤣
Last year in 2022, one night in my bedroom, I saw a light figure with a glow, shape of a man just like the Asgard, it had the same look, like a glowing light. I couldn’t take my eyes off it, it was stunning. I couldn’t move, I was terrified. It just stood there until it moved towards me. I closed my eyes quickly then it disappeared. I wondered what it was until now after reading this article. It is a light being before we become human form, our beginning. I now understand what it was. There’s another form of life in our universe that most of us cannot see or that form will not allow us to see. It will terrify us or maybe we will not accept the truth. Scientists have been searching for the longest for our origin, now you have found it and there is more you will find. It’s all here on earth. The earth that God created. The light being, our origin. Everything is here. It’s not on another planet. All the answers are right here on this planet and moving around us in our existence.
Your experience is not verified fact.
The article describe verified fact.