
Once chemically adjusted for use in animals, cilagicin consistently and safely eliminated Gram-positive bacteria in the lab, did not damage human cells, and successfully cured bacterial infections in mice.
Antibiotic-resistant pathogens could be defeated with the assistance of a synthetic antibiotic
A brand-new antibiotic that was developed at The Rockefeller University using computational models of bacterial gene products appears to kill even bacteria that are resistant to other antibiotics. According to a study published in the journal Science, the drug, known as cilagicin, is effective in mice and employs a novel mechanism to combat MRSA, C. diff, and numerous other dangerous infections.
The findings imply that computer models may be used to develop a new class of antibiotics. “This isn’t just a cool new molecule, it’s a validation of a novel approach to drug discovery,” says Rockefeller’s Sean F. Brady. “This study is an example of computational biology, genetic sequencing, and synthetic chemistry coming together to unlock the secrets of bacterial evolution.”
Acting on eons of bacterial warfare
Bacteria have spent billions of years inventing novel methods to kill one another, so it’s not surprising that many of our most potent antibiotics originated from bacteria. With the exception of penicillin and a few other prominent antibiotics originating from fungus, the majority of antibiotics were first used as weapons by bacteria to combat other bacteria.
“Eons of evolution have given bacteria unique ways of engaging in warfare and killing other bacteria without their foes developing resistance,” says Brady, the Evnin Professor and head of the Laboratory of Genetically Encoded Small Molecules. Antibiotic drug discovery once largely consisted of scientists growing streptomyces or bacillus in the lab and bottling their secrets to treat human diseases.
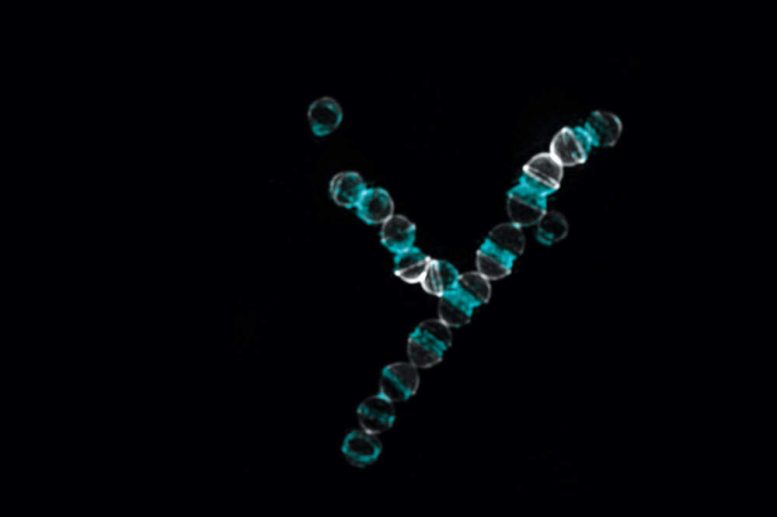
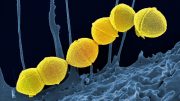
The synthetic antibiotic cilagicin was particularly active against Gram-positive bacteria such as Streptococcus pyogenes, depicted above. Credit: Rockefeller University
But with the rise of antibiotic-resistant bacteria, there is an urgent need for new active compounds—and we may be running out of bacteria that are easy to exploit. Untold numbers of antibiotics, however, are likely hidden within the genomes of stubborn bacteria that are tricky or impossible to study in the lab. “Many antibiotics come from bacteria, but most bacteria can’t be grown in the lab,” Brady says. “It follows that we’re probably missing out on most antibiotics.”
Finding antibacterial genes in soil and cultivating them inside more lab-friendly bacteria is an alternate strategy that has been championed by the Brady lab for the last fifteen years. But even this approach has certain drawbacks. The majority of antibiotics come from genetic sequences that are locked within bacterial gene clusters named “biosynthetic gene clusters,” which work together to collectively code for a number of proteins. But with present technology, such clusters are often inaccessible.
“Bacteria are complicated, and just because we can sequence a gene doesn’t mean we know how the bacteria would turn it on to produce proteins,” Brady says. “There are thousands and thousands of uncharacterized gene clusters, and we have only ever figured out how to activate a fraction of them.”
A new pool of antibiotics
Frustrated with their inability to unlock many bacterial gene clusters, Brady and colleagues turned to algorithms. By teasing apart the genetic instructions within a DNA sequence, modern algorithms can predict the structure of the antibiotic-like compounds that a bacterium with these sequences would produce. Organic chemists can then take that data and synthesize the predicted structure in the lab.
It may not always be a perfect prediction. “The molecule that we end up with is presumably, but not necessarily, what those genes would produce in nature,” Brady says. “We aren’t concerned if it is not exactly right—we only need the synthetic molecule to be close enough that it acts similarly to the compound that evolved in nature.”
Postdoctoral associates Zonggiang Wang and Bimal Koirala from the Brady lab began by searching through an enormous genetic-sequence database for promising bacterial genes that were predicted to be involved in killing other bacteria and hadn’t been examined previously. The “cil” gene cluster, which had not yet been explored in this context, stood out for its proximity to other genes involved in making antibiotics. The researchers duly fed its relevant sequences into an algorithm, which proposed a handful of compounds that cil likely produces. One compound, aptly dubbed cilagicin, turned out to be an active antibiotic.
Cilagicin reliably killed Gram-positive bacteria in the lab, did not harm human cells, and (once chemically optimized for use in animals) successfully treated bacterial infections in mice. Of particular interest, cilagicin was potent against several drug-resistant bacteria and, even when pitted against bacteria grown specifically to resist cilagicin, the synthetic compound prevailed.
Brady, Wang, Koirala, and colleagues determined that cilagicin works by binding two molecules, C55-P and C55-PP, both of which help maintain bacterial cell walls. Existing antibiotics such as bacitracin bind one of those two molecules but never both, and bacteria can often resist such drugs by cobbling together a cell wall with the remaining molecule. The team suspects that cilagicin’s ability to take both molecules offline may present an insurmountable barrier that prevents resistance.
Cilagicin is still far from human trials. In follow-up studies, the Brady lab will perform further syntheses to optimize the compound and test it in animal models against more diverse pathogens to determine which diseases it may be most effective in treating.
Beyond the clinical implications of cilagicin, however, the study demonstrates a scalable method that researchers could use to discover and develop new antibiotics. “This work is a prime example of what could be found hidden within a gene cluster,” Brady says. “We think that we can now unlock large numbers of novel natural compounds with this strategy, which we hope will provide an exciting new pool of drug candidates.”
Reference: “Bioinformatic prospecting and synthesis of a bifunctional lipopeptide antibiotic that evades resistance” by Zongqiang Wang, Bimal Koirala, Yozen Hernandez, Matthew Zimmerman and Sean F. Brady, 26 May 2022, Science.
DOI: 10.1126/science.abn4213




Great people of sciences arts and technologies.”Mon chapeau” gentlemen my deepest respect to most of you. These are the people that should Admn., gov., issues, economy, etc., because these politicians, lawmakers, religious pigs, are nothing but garbage, delinquency, crime opportunism, thievery, abuse.
We must do smthng to get ready of these monkeys with delirium of grandeurs…
If our country wasn’t so capitalistic, this discovery would likely mean the patent royalties due to Rockefeller University would be great enough that no student would ever need to pay tuition again! Potentially every student in New York would never have to pay tuition again due to the supposed ongoing patent royalties.
Is that how it really works? Nope, surely they will take this discovery made by an educational institution and make sure that some private drug company and a few billionaires receive all the riches. All the while, the politicians will have us deceived and start class warfare over topics like education funding. (While they charge us 6-7% recapitalized interest on our student loans.)
How are politicians (from both parties) any better than a deadly bacteria?
Um. OK
Gee Wiz is obviously a pea brain not an egg head.
Oh good, so now our kids/grandkids can grow up with a new generation of drug-immune bacteria because of this. Until the idiots that stop taking antibiotics when they start feeling better smarten up (or hopefully die off), people should not be allowed to have meds.
I couldn’t agree more !
The time for Nicola Tesla’s healthcare technologies is finally dawning .
Replacing the root cause of most of our chronic diseases that stem from Round up and all those other pesticides that are banned in Europe ….. is possible with his applications in the field of bio energetics .
This is the future perfect we have been praying for .
Looks like our wishes were granted .
I think this title is misleading, “A New Antibiotic Can Kill Even Drug-Resistant Bacteria” the “drug-resistant” bacteria are resistant to existing antibiotics. When new antibiotics are introduced we will have new bacteria that are “drug-resistant” to the new antibiotics thanks to natural selection.
The awesome part of this research is finding new ways to quickly develop antibiotics with bioinformatics.
That’s what the umbrella Corp. Was doing and look at what happened to raccoon city.
What effect does this antibiotic has on the gut microbiome. It is not only finding antibiotics to kill drug resistance bacteria but friendly antibiotics that will not cause neurotoxicity, Causing chemical imbalances etc., and kill all the important gut bacteria, leaving the patients sick after treatment, like some of us are going through now.
Marvelous, but to echo Sanyo, it will cause disbacteriosis and inevitably c diff infection in the gut, since c diff is likely to survive anyway. C diff will kill fast, much faster than the original infection, I tell that from experience
拼错了 Zonggiang Wang 应该是 Zongqiang
This definitely sounds like the start of a zombie flick.
Bacteria and Viruses only job is to stay alive. Then learn they change and they replicate. Somehow some way this drug will not be effective as described. You can’t beat the science in this. Just like PCN. The wonder drug of the future. It failed and became resistant. It just took time.
This publication routinely publishes preliminary data as if it means that a new drug is available. This is incorrect and misleading.
It isn’t “A New Antibiotic” until it goes through 3 phases of clinical trials and gets approved by the FDA if it will be used in the USA.
Right now it is an experimental molecule.
Be aware, it’s going to kill all the germs , viruses and bacteria in your body only problem your going to die from it as well.
I need this
This is truly incredible!!!
Great news but as mentioned this drug may fail during testing on humans because of toxicity! Did they study histological changes that occurred in mice? This is just a hit and it’s road to become a lead is still long.
Anyways we have AMR not only in bacteria but also in viruses but you guys are focusing on bacteria while most ARVs no longer work on HIV. HIV poses a major threat to Public health than the c.difficele you are most scared off
I would request we turn research into more important topics.
Reading these comments is often more entertaining, just as intelligent, & everybit as valuable as reading an initial report, so keep thinking & keep sounding off, you all!