
A neural network analyzes an extensive gut microbiome dataset to uncover insights into human health.
Gut bacteria play an important role in a wide range of health conditions. Their sheer diversity and the complexity of their interactions with both the body’s biochemistry and with one another make them difficult to study. In a new approach, researchers at the University of Tokyo applied a specialized form of artificial intelligence known as a Bayesian neural network to analyze a large dataset on gut microbes.
This method allowed them to uncover patterns and connections that traditional analytical techniques have struggled to detect reliably.
Why gut bacteria matter for health
The human body contains roughly 30 to 40 trillion cells, yet the intestines house around 100 trillion gut bacteria. In other words, microbial cells in your body outnumber your own. While these bacteria are commonly associated with digestion, they also influence a wide range of bodily functions.
They exist in vast diversity and generate or modify numerous chemical compounds known as metabolites. These metabolites act as signaling molecules, traveling through the body and impacting systems such as immunity, metabolism, brain activity, and mood. Gaining deeper insight into gut bacteria could offer significant health benefits.
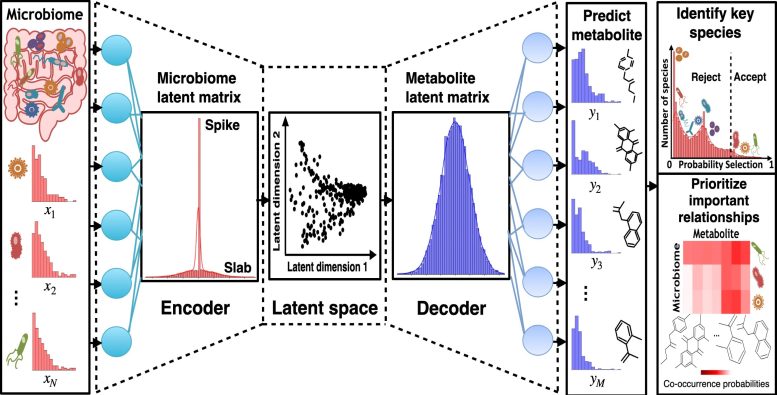
“The problem is that we’re only beginning to understand which bacteria produce which human metabolites and how these relationships change in different diseases,” said Project Researcher Tung Dang from the Tsunoda lab in the Department of Biological Sciences. “By accurately mapping these bacteria-chemical relationships, we could potentially develop personalized treatments. Imagine being able to grow a specific bacterium to produce beneficial human metabolites or designing targeted therapies that modify these metabolites to treat diseases.”
The challenge of complexity
It may sound promising, but there is a major challenge. The sheer number and diversity of both gut bacteria and metabolites make the potential relationships between them overwhelmingly complex. Collecting data is already an enormous task, but analyzing it to uncover meaningful biological patterns is even more difficult. To address this, Dang and his team turned to advanced artificial intelligence (AI) tools for deeper analysis.
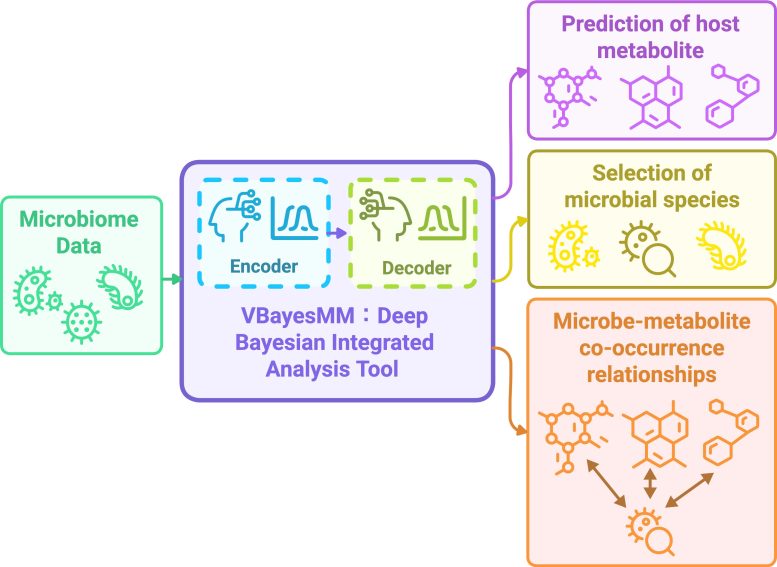
“Our system, VBayesMM, automatically distinguishes the key players that significantly influence metabolites from the vast background of less relevant microbes, while also acknowledging uncertainty about the predicted relationships, rather than providing overconfident but potentially wrong answers,” said Dang. “When tested on real data from sleep disorder, obesity, and cancer studies, our approach consistently outperformed existing methods and identified specific bacterial families that align with known biological processes, giving confidence that it discovers real biological relationships rather than meaningless statistical patterns.”
How VBayesMM tackles gut data
As VBayesMM can handle and communicate issues of uncertainty, it gives researchers more confidence than a tool that does not. Even though the system is optimized to cope with heavy analytical workloads, mining such huge datasets still comes with high computational cost; however, as time goes on, this will become less and less of a barrier to those wishing to use it.
Other limitations at present include that the system benefits from having more data about the gut bacteria than the metabolites they produce; when there’s insufficient bacteria data, the accuracy drops. Also, VBayesMM assumes the microbes act independently, but in reality, gut bacteria interact in an incredibly complex number of ways.
“We plan to work with more comprehensive chemical datasets that capture the complete range of bacterial products, though this creates new challenges in determining whether chemicals come from bacteria, the human body, or external sources like diet,” said Dang. “We also aim to make VBayesMM more robust when analyzing diverse patient populations, incorporating bacterial ‘family tree’ relationships to make better predictions, and further reducing the computational time needed for analysis. For clinical applications, the ultimate goal is identifying specific bacterial targets for treatments or dietary interventions that could actually help patients, moving from basic research toward practical medical applications.”
Reference: “VBayesMM: variational Bayesian neural network to prioritize important relationships of high-dimensional microbiome multiomics data” by Tung Dang, Artem Lysenko, Keith A Boroevich and Tatsuhiko Tsunoda, 4 July 2025, Briefings in Bioinformatics.
DOI: 10.1093/bib/bbaf300
This work was partly supported by JSPS KAKENHI Grant Numbers JP20H03240 and JP24K15175, and JST CREST Grant Number JPMJCR2231.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
2 Comments
When i see articles like this, i think of AI hallucinations and researchers not understanding how AI reaches the concessions it does.
In other words, give me independent verification before touting their achievements
Look at the possibilities rather the limitations!