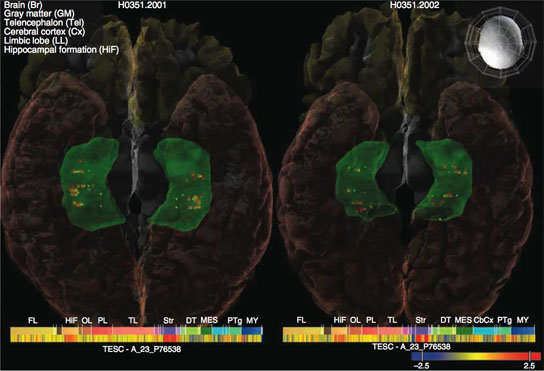
Based on a genetic analysis of more than 9,000 people, a new study shows that four genes may speed shrinkage of the hippocampus in the brain and could lead to an increased risk of Alzheimer’s disease. This study, and another on genetic variants of intracranial volume, may lead to new findings that help doctors better understand brain development.
Two research studies, co-led by UC Davis neurologist Charles DeCarli and conducted by an international team that included more than 80 scientists at 71 institutions in eight countries, has advanced understanding of the genetic components of Alzheimer’s disease and of brain development. Both studies appear in the April 15 edition of the journal Nature Genetics.
The first study, based on a genetic analysis of more than 9,000 people, has found that certain versions of four genes may speed shrinkage of a brain region involved in making new memories. The brain area, known as the hippocampus, normally shrinks with age, but if the process speeds up, it could increase vulnerability to Alzheimer’s disease, the research suggests.
The second paper identifies two genes associated with intracranial volume — the space within the skull occupied by the brain when the brain is fully developed in a person’s lifespan, usually around age 20.
DeCarli is an internationally renowned pioneer in the field of neuroimaging of the aging brain who has been at the forefront of developing and using quantifiable imaging techniques to define the relationship between structure and function in the healthy aging brain and to characterize the changes associated with vascular and Alzheimer’s dementias. He is a professor of neurology and director of the UC Davis Alzheimer’s Disease Center and the UC Davis Imaging of Dementia and Aging Laboratory.
Genetic variants of hippocampus study
The gene variants identified in the first study do not cause Alzheimer’s, but they may rob the hippocampus of a kind of “reserve” against the disease, which is known to cause cell destruction and dramatic shrinkage of this key brain site. The result is severe loss of memory and cognitive ability.
Scientists calculated that hippocampus shrinkage in people with these gene variants accelerates by about four years on average. The risk of Alzheimer’s doubles every five years beginning at age 65, so a person of that age would face almost twice the Alzheimer’s risk if he or she had these versions of the gene.
Looked at another way, if a person with one of these variants did get Alzheimer’s, the disease would attack an already compromised hippocampus and so would lead to a more severe condition at a younger age than otherwise, the research suggests.
“This is definitely a case of ‘bigger is better,'” said DeCarli. “We already know that Alzheimer’s disease causes much of its damage by shrinking hippocampus volume. If someone loses a greater-than-average amount of volume due to the gene variants we’ve identified, the hippocampus is more vulnerable to Alzheimer’s.”
Why the aging hippocampus normally decreases in volume is unclear. The new research shows that the genes most strongly linked to shrinkage are involved in maturation of the hippocampus and in apoptosis, or programmed cell death – a continual process by which older cells are removed from active duty.
The scientists suggest that if the gene variants they identified do affect either maturation or the rate at which cells die, this could underlie at least some of the increased rates of hippocampus shrinkage.
“Either by making more or healthier hippocampal neurons or preventing them from dying with advancing age, the healthy versions of these genes influence how people remember as they get older,” said DeCarli. “The alternate versions of the genes may not fully provide these benefits.”
The researchers hope that they can find ways to protect the hippocampus from premature shrinkage or slow its decline by studying the normal regulation of the proteins coded by these genes.
The genetic analysis draws on what is known as a genome-wide association study — research aimed at finding the common genetic variants associated with specific diseases or other conditions. Different versions of a gene usually come down to changes in just one of the tens of thousands of DNA “letters” that make up genes. These one-letter differences are known as single-nucleotide polymorphisms, or SNPs.
The research involved more than 80 scientists at 71 institutions in 8 countries. Many researchers are needed for such a study in order to put together the large samples, or cohorts, of people whose genetic makeup is to be investigated, to measure the hippocampus from magnetic resonance pictures of the brain and for the labor-intensive statistical analysis of the findings.
The study used a very large assemblage of genetic and disease data called the Cohorts for Heart and Aging Research in Genomic Epidemiology Consortium, or CHARGE. The consortium brings together several population-based cohorts in the United States and Europe.
The cohort was made up of 9,232 dementia-free volunteers with an average age of 67. The study identified four different gene variants associated with hippocampus volume decline. One, known as rs7294919, showed a particularly strong link to a reduced hippocampus volume, suggesting that this gene is very important to hippocampus development or health.
The findings were then assessed in two other cohorts. One, including both normal and cognitively compromised people with an average age of 40, showed that three of the suspect SNPs were linked to reduced hippocampus volume. Analysis of results from the third group, comprised primarily of older people, showed a significant association between one of the SNPs and accelerated memory loss.
“With this study, we have new evidence that aging, the hippocampus, and memory are influenced by specific genes,” DeCarli said. “Understanding how these genes affect the development and aging of the hippocampus may give us new tools to delay memory loss with advanced age and possibly reduce the impact of such diseases as Alzheimer’s disease.”
Genetic variants of Intracranial-volume study
While the first study deals with the genetic associations with brain shrinkage, the second deals with associations impacting intracranial volume, which is an indirect measure of the size of the brain at full development.
Though brain volume and intracranial volume are both highly heritable, the genetic influences on these measures may differ. To assess the genetic influence on these two measures, researchers in the second study performed a genome-wide association study on cross-sectional measures of intracranial volume and brain volume in 8,175 elderly in the CHARGE consortium.
They found no associations for brain volume, but they did discover that intracranial volume was significantly associated with two loci: rs4273712, a known height locus on chromosome 6q22, and rs9915547, tagging the inversion on chromosome 17q21.
“Since geneticists are already familiar with the other functions of these same genes, associating these particular genes with intracranial volume may help us better understand brain development in general,” said DeCarli. “For instance, we know that one of these genes has played a unique evolutionary role in human development, and perhaps we as a species are selecting this gene as a way of providing further advances in brain development.”
Both studies involved international teams representing scores of institutions, funded by a variety of NIH grants as well as grants from agencies around the world. Please refer to the papers for complete lists of authors, affiliations, and funding sources.
References:
“Common variants at 12q14 and 12q24 are associated with hippocampal volume” by Enhancing Neuro Imaging Genetics through Meta-Analysis (ENIGMA) Consortium and the Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium, 15 April 2012, Nature Genetics.
DOI: 10.1038/ng.2237
“Common variants at 6q22 and 17q21 are associated with intracranial volume” by the Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium and Early Growth Genetics (EGG) Consortium, 15 April 2012, Nature Genetics.
DOI: 10.1038/ng.2245
“Identification of common variants associated with human hippocampal and intracranial volumes” by Jason L Stein, Sarah E Medland, Alejandro Arias Vasquez, Derrek P Hibar, Rudy E Senstad, Anderson M Winkler, Roberto Toro, Katja Appel, Richard Bartecek, Ørjan Bergmann, Manon Bernard, Andrew A Brown, Dara M Cannon, M Mallar Chakravarty, Andrea Christoforou, Martin Domin, Oliver Grimm, Marisa Hollinshead, Avram J Holmes, Georg Homuth, Jouke-Jan Hottenga, Camilla Langan, Lorna M Lopez, Narelle K Hansell, Kristy S Hwang, Sungeun Kim, Gonzalo Laje, Phil H Lee, Xinmin Liu, Eva Loth, Anbarasu Lourdusamy, Morten Mattingsdal, Sebastian Mohnke, Susana Muñoz Maniega, Kwangsik Nho, Allison C Nugent, Carol O’Brien, Martina Papmeyer, Benno Pütz, Adaikalavan Ramasamy, Jerod Rasmussen, Mark Rijpkema, Shannon L Risacher, J Cooper Roddey, Emma J Rose, Mina Ryten, Li Shen, Emma Sprooten, Eric Strengman, Alexander Teumer, Daniah Trabzuni, Jessica Turner, Kristel van Eijk, Theo G M van Erp, Marie-Jose van Tol, Katharina Wittfeld, Christiane Wolf, Saskia Woudstra, Andre Aleman, Saud Alhusaini, Laura Almasy, Elisabeth B Binder, David G Brohawn, Rita M Cantor, Melanie A Carless, Aiden Corvin, Michael Czisch, Joanne E Curran, Gail Davies, Marcio A A de Almeida, Norman Delanty, Chantal Depondt, Ravi Duggirala, Thomas D Dyer, Susanne Erk, Jesen Fagerness, Peter T Fox, Nelson B Freimer, Michael Gill, Harald H H Göring, Donald J Hagler, David Hoehn, Florian Holsboer, Martine Hoogman, Norbert Hosten, Neda Jahanshad, Matthew P Johnson, Dalia Kasperaviciute, Jack W Kent Jr, Peter Kochunov, Jack L Lancaster, Stephen M Lawrie, David C Liewald, René Mandl, Mar Matarin, Manuel Mattheisen, Eva Meisenzahl, Ingrid Melle, Eric K Moses, Thomas W Mühleisen, Matthias Nauck, Markus M Nöthen, Rene L Olvera, Massimo Pandolfo, G Bruce Pike, Ralf Puls, Ivar Reinvang, Miguel E Rentería, Marcella Rietschel, Joshua L Roffman, Natalie A Royle, Dan Rujescu, Jonathan Savitz, Hugo G Schnack, Knut Schnell, Nina Seiferth, Colin Smith, Vidar M Steen, Maria C Valdés Hernández, Martijn Van den Heuvel, Nic J van der Wee, Neeltje E M Van Haren, Joris A Veltman, Henry Völzke, Robert Walker, Lars T Westlye, Christopher D Whelan, Ingrid Agartz, Dorret I Boomsma, Gianpiero L Cavalleri, Anders M Dale, Srdjan Djurovic, Wayne C Drevets, Peter Hagoort, Jeremy Hall, Andreas Heinz, Clifford R Jack Jr, Tatiana M Foroud, Stephanie Le Hellard, Fabio Macciardi, Grant W Montgomery, Jean Baptiste Poline, David J Porteous, Sanjay M Sisodiya, John M Starr, Jessika Sussmann, Arthur W Toga, Dick J Veltman, Henrik Walter, Michael W Weiner, the Alzheimer’s Disease Neuroimaging Initiative (ADNI), EPIGEN Consortium, IMAGEN Consortium, Saguenay Youth Study Group (SYS), Joshua C Bis, M Arfan Ikram, Albert V Smith, Vilmundur Gudnason, Christophe Tzourio, Meike W Vernooij, Lenore J Launer, Charles DeCarli, Sudha Seshadri, Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium, Ole A Andreassen, Liana G Apostolova, Mark E Bastin, John Blangero, Han G Brunner, Randy L Buckner, Sven Cichon, Giovanni Coppola, Greig I de Zubicaray, Ian J Deary, Gary Donohoe, Eco J C de Geus, Thomas Espeseth, Guillén Fernández, David C Glahn, Hans J Grabe, John Hardy, Hilleke E Hulshoff Pol, Mark Jenkinson, René S Kahn, Colm McDonald, Andrew M McIntosh, Francis J McMahon, Katie L McMahon, Andreas Meyer-Lindenberg, Derek W Morris, Bertram Müller-Myhsok, Thomas E Nichols, Roel A Ophoff, Tomas Paus, Zdenka Pausova, Brenda W Penninx, Steven G Potkin, Philipp G Sämann, Andrew J Saykin, Gunter Schumann, Jordan W Smoller, Joanna M Wardlaw, Michael E Weale, Nicholas G Martin, Barbara Franke, Margaret J Wright and Paul M Thompson for the Enhancing Neuro Imaging Genetics through Meta-Analysis (ENIGMA) Consortium, 15 April 2012, Nature Genetics.
DOI: 10.1038/ng.2250
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.