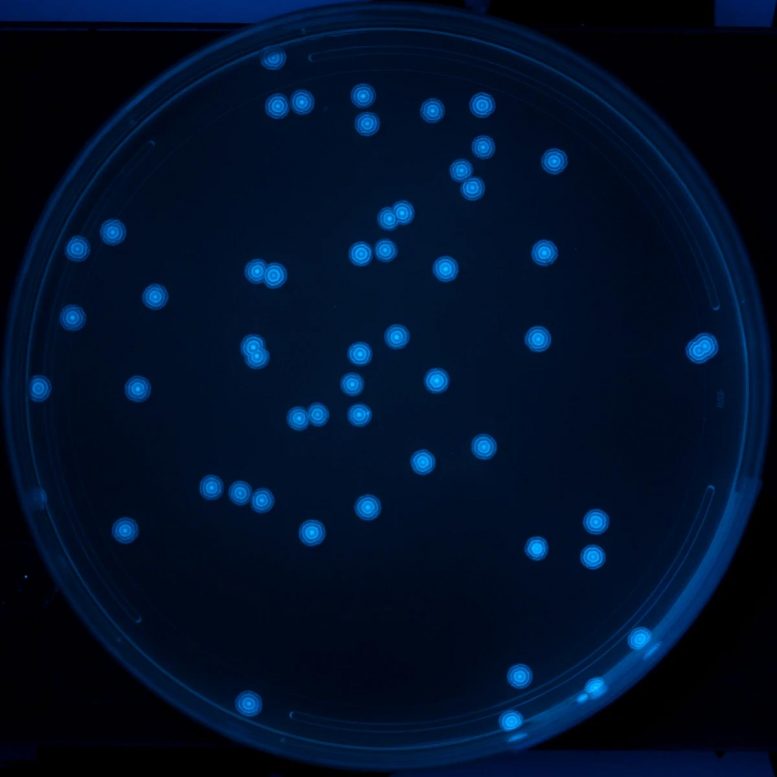
Genetic oscillator records changes in microbiome growth patterns in vivo.
For all the attention the human microbiome has been getting over the last few years, one aspect of such research rarely makes headlines: the difficulty of observing how it changes over time in response to various stimuli. The most common analysis method is extracting bacteria from fecal samples and then sequencing their genomes, but this approach, while minimally invasive, loses crucial information about where and when bacterial changes occur in the gut, providing scientists with an incomplete picture of the dynamics of the microbiome.
Now, a new tool created by researchers at the Wyss Institute for Biologically Inspired Engineering at Harvard University and Harvard Medical School (HMS) provides a solution to this problem in the form of a set of bacterial genes that have been engineered to detect and record changes in the growth of different populations of bacteria over time in the guts of living mice with single-cell precision, and can serve as a platform for complex, synthetic-biology-based diagnostics and therapeutics for a variety of applications in the gut. The study was published today, October 11, 2019, in Nature Communications.
Keeping time
The system uses an oscillating gene circuit, called a repressilator, as a kind of genetic clock to measure bacterial growth. The repressilator consists of three bacterial genes that code for three proteins (tetR, cl, and lacI), each of which blocks the expression of one of the other proteins. The genes are linked into a negative feedback loop, so that when the concentration of one of the repressor proteins falls below a certain level, the protein it had been repressing is expressed, which blocks the expression of the third protein, and the process repeats in a cyclical fashion.

When all three genes are inserted into a plasmid and introduced into bacteria, the number of negative feedback loop cycles completed can serve as a record of how many cell divisions the bacteria have undergone. Every time the bacteria divide, any repressor proteins present in their cytoplasm are diluted, so their concentration gradually falls and triggers the expression of the next protein in the repressilator cycle. Crucially, the repressilator cycle repeats after 15.5 bacterial generations regardless of how quickly or slowly the bacteria are growing. This allows it to act as an objective measurement of time, much like a clock or a watch.
“Imagine if you had two people wearing two different watches, and the second hand on one person’s watch was moving twice as fast as the other person’s,” explained first author David Riglar, Ph.D., a former postdoc at the Wyss Institute and HMS who now leads a research group as a Sir Henry Dale Fellow at Imperial College London. “If you stopped both watches after one hour, they wouldn’t agree on what time it was, because their measurement of time varies based on the rate of the second hand’s movement. In contrast, our repressilator is like a watch that always moves at the same speed, so no matter how many different people are wearing one, they will all give a consistent measurement of time. This quality allows us to more precisely study the behavior of bacteria in the gut.”
The researchers coupled each of the three repressor proteins to a differently colored fluorescent molecule, and developed an imaging workflow called RINGS (Repressilator-based Inference of Growth at Single-cell level) to track which protein is expressed at different time points during the bacteria’s growth. “As a bacterial colony grows outwards, the repressilator circuit creates these different fluorescents, tree-ring-like signatures based on which repressor protein was active in the single bacterium that started the colony,” said Riglar. “The pattern of the fluorescent rings records how many repressilator cycles have occurred since growth began, and we can analyze that pattern to study how growth rates vary between different bacteria and in different environments.”
Using RINGS, the team was able to successfully track cell divisions in several different bacterial species grown in vitro, and observed that the length of the bacteria’s repressilator cycle remained consistent when they were grown on extracted samples of mouse intestine (to simulate a complex microenvironment) or exposed to an antibiotic (to simulate stress conditions and inconsistent growth patterns).

Tracking change
To evaluate the repressilator’s performance in vivo, the team administered E. coli containing the repressilator circuit to mice orally, then analyzed bacteria extracted from fecal samples. The repressilator remained active for up to 16 days after introduction, showing that long-term oscillatory gene expression could be maintained in gut bacteria in living mammals. The RINGS analysis successfully detected changes in bacterial growth patterns, and bacteria whose repressilator circuits were in different stages could be “synchronized” by giving the mice a compound in their drinking water that arrested the repressilator cycle at a given stage.
Finally, the researchers tested the repressilator’s ability to detect differences in bacterial growth rates that have been observed as a result of gut inflammation. Mice were given an inflammation-inducing compound, followed by repressilator-loaded bacteria. After 15 hours, RINGS analysis showed that the bacteria from mice with inflammation had repressilators in a wider range of phases compared to bacteria from control mice, suggesting that inflammation produces an environment that drives inconsistencies in bacterial growth, potentially leading to imbalances in the gut microbiome.
“This repressilator allows us to really probe the intricacies of bacterial behavior in the living gut, not only in both healthy and diseased states, but also spatially and temporally,” said corresponding author Pamela Silver, Ph.D., who is a Core Faculty member at the Wyss Institute and the Elliot T. and Onie H. Adams Professor of Biochemistry and Systems Biology at HMS. “The fact that we can re-synchronize the repressilator when it’s already in the gut, as well as maintain it without the need to administer selective antibiotics, also means that we can study the microbiome in a more natural state with minimal disruption.”
In addition to understanding the dynamics of the microbiome, the repressilator unlocks the potential for complex, synthetic-biology-based diagnostics and therapeutics for the human gut. Potential applications include the creation of a system that is programmed to initiate a gene transcription cascade at a certain point in the circadian rhythm, or a diagnostic that records how much time has elapsed following the detection of a given biomarker.
“Not only does this research solve a specific problem related to monitoring dynamic changes in microbiome physiology within the living gut, it provides a platform that could lead to entirely new types of diagnostics and even time-dependent therapeutics,” said Wyss Founding Director Donald Ingber, M.D., Ph.D., who is also the Judah Folkman Professor of Vascular Biology at HMS and the Vascular Biology Program at Boston Children’s Hospital, as well as Professor of Bioengineering at Harvard’s John A. Paulson School of Engineering and Applied Sciences.
###
Additional authors of the paper include David Richmond, Laurent Potvin-Trottier, Andrew Verdegaal, Somenath Bakshi, Emanuele Leoncini, Lorena Lyon, and Johan Paulsson from HMS, and Alexander Naydich from the Wyss Institute, HMS, and Harvard’s John A. Paulson School of Engineering and Applied Sciences. This research was supported by a Human Frontier Science Program Long-Term Fellowship, the Menzies Foundation, the Wellcome Trust, the National Science Foundation, the Defense Advanced Research Projects Agency, Harvard Medical School, and the Wyss Institute for Biologically Inspired Engineering.
Reference: “Bacterial variability in the mammalian gut captured by a single-cell synthetic oscillator” by David T. Riglar, David L. Richmond, Laurent Potvin-Trottier, Andrew A. Verdegaal, Alexander D. Naydich, Somenath Bakshi, Emanuele Leoncini, Lorena G. Lyon, Johan Paulsson and Pamela A. Silver, 11 October 2019, Nature Communications.
DOI: 10.1038/s41467-019-12638-z
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.