
The New Technique Is “Like Upgrading From the Hubble to the James Webb”
A new imaging technique captures the three-dimensional architecture of the human genome with unprecedented detail, showing how individual genes fold at the level of nucleosomes, the fundamental units constituting the genome’s three-dimensional architecture.
The technology, which was created by Barcelona-based researchers at the Centre for Genomic Regulation (CRG) and the Institute for Research in Biomedicine (IRB Barcelona), combines high-resolution microscopy with sophisticated computer modeling. It is the most comprehensive technique to date for studying the shape of genes.
The new technique allows researchers to create and digitally navigate three-dimensional models of genes, seeing not just their architecture but also information on how they move or how flexible they are. Understanding how genes function might help us better understand how they influence the human body in both health and disease since almost every human disease has some genetic basis.
Scientists will eventually be able to utilize this knowledge to predict what happens to genes when things go wrong, such as cataloging differences in the structure of genes that cause disease. The method might potentially be used to test drugs that change the shape of an aberrant gene, helping in the development of novel treatments for various diseases.
The technology is the next evolution of imaging techniques used to study living organisms, which first started more than four hundred years ago with the creation of microscopes. These played a crucial role in advancing medicine and human health, for example, used by Robert Hooke to describe cells for the first time and later used by Santiago Ramón y Cajal to identify neurons. Despite great advances, the limitations of optical microscopes were clear as far back as 1873, with researchers stipulating that their maximum resolution could not surpass 0.2 micrometers.
This physical limit was overcome in the 21st century with the creation of super-resolution microscopy, a breakthrough that was awarded the Nobel Prize in Chemistry in 2014. Using fluorescence, researchers extended the limits of light microscopy and captured events at 20 nanometres, a feat that revealed how life works at an unprecedented molecular scale.
Super-resolution microscopy changed the course of biomedical research, enabling scientists to track proteins in a variety of diseases. It also enabled researchers to study molecular events that regulate gene expression. Scientists now want to build on the technology and take it one step further by adding more layers of information.
MiOS: A New Era of Super-Resolution Imaging
Researchers hypothesized that taking super-resolution microscopy and merging it with advanced computational tools could be a way of imaging genes at the level of detail necessary to study their shape and function. An interdisciplinary team of scientists shared their expertise and created a new technique called Modeling immuno-OligoSTORM – or MiOS for short.
The two research groups teamed up as part of the Barcelona Institute of Science and Technology’s (BIST) Ignite Call, which facilitates the exchange of knowledge among different scientific fields and exploring new approaches to address complex questions.

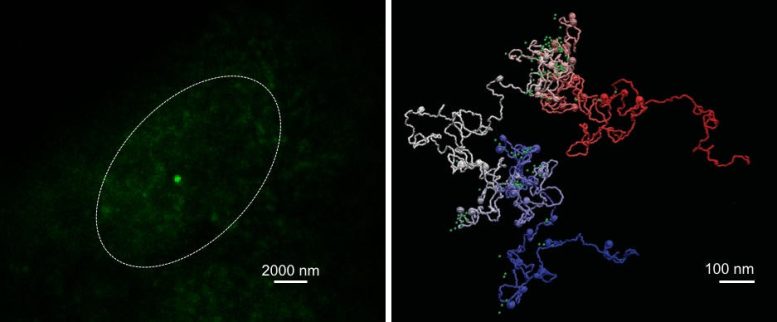
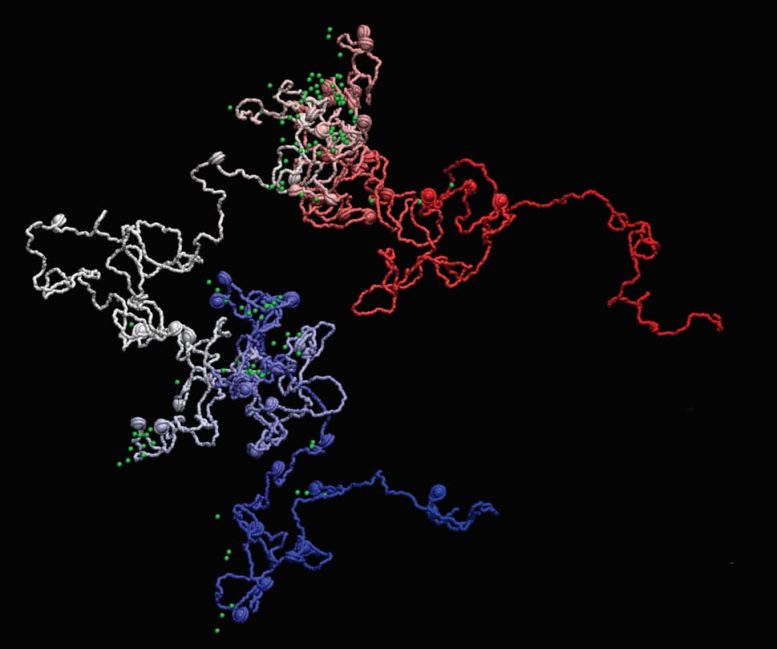
“Our computational modeling strategy integrates data from DNA sequencing techniques and super-resolution microscopy to provide an essential picture (or movie) of the 3D shape of genes at resolutions beyond the size of nucleosomes, reaching the scales needed to understand in detail the interaction between chromatin and other cell factors,” says Dr. Juan Pablo Arcon, co-first author of the work and postdoctoral researcher at IRB Barcelona.
Unprecedented Insights Into Gene Structure and Function
As proof of concept, the research team used MiOS to provide new insights on the position, shape, and compaction of key housekeeping and pluripotency genes, revealing new structures and details that are not captured using conventional techniques alone. The findings are published in the journal Nature Structural & Molecular Biology. The study’s corresponding authors include ICREA Research Professor Pia Cosma at the CRG and Professor Modesto Orozco at IRB Barcelona, as well as Pablo Dans, previously a researcher at IRB Barcelona and now at University of the Republic (Uruguay) and the Institut Pasteur of Montevideo.
“We show that MiOS provides unprecedented detail by helping researchers virtually navigate inside genes, revealing how they are organized at a completely new scale. It is like upgrading from the Hubble Space Telescope to the James Webb, but instead of seeing distant stars we’ll be exploring the farthest reaches inside a human nucleus,” says Dr. Vicky Neguembor, co-first and also a co-corresponding author of the study and researcher at the CRG.
While a lot of genome-based research is already changing how we diagnose, treat, or prevent diseases, the impact of MiOS is more long-term. By shedding light on how genes work and how they are regulated at the nanoscale, the technique will enable new discoveries in the scientific laboratory, some of which might eventually translate into clinical practice.
The research team is already putting MiOS to use by exploring genes that are important for human development. The team will also keep developing MiOS further, adding additional functionality that can for example detect how transcription factors – proteins involved in the process of converting or transcribing, DNA into RNA – bind to DNA.
Reference: “MiOS, an integrated imaging and computational strategy to model gene folding with nucleosome resolution” by Maria Victoria Neguembor, Juan Pablo Arcon, Diana Buitrago, Rafael Lema, Jürgen Walther, Ximena Garate, Laura Martin, Pablo Romero, Jumana AlHaj Abed, Marta Gut, Julie Blanc, Melike Lakadamyali, Chao-ting Wu, Isabelle Brun Heath, Modesto Orozco, Pablo D. Dans and Maria Pia Cosma, 11 October 2022, Nature Structural & Molecular Biology.
DOI: 10.1038/s41594-022-00839-y
The study was funded by the Barcelona Institute of Science and Technology, the Horizon 2020 Framework Programme, the Spanish Ministry of Science, Innovation, and Universities, and the Government of Catalonia.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.