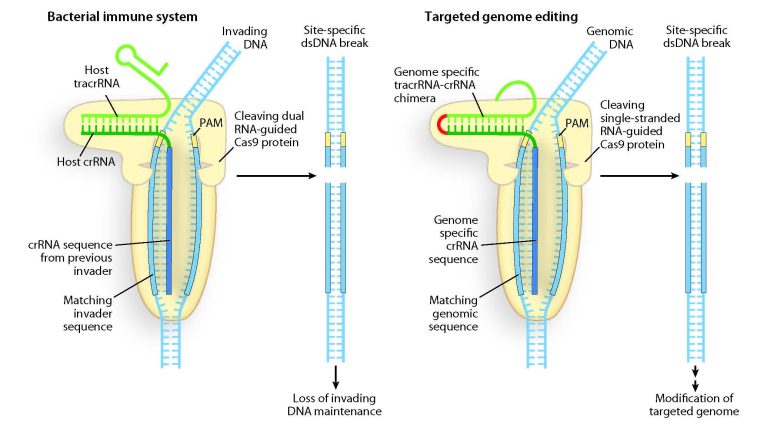
A team of scientists at the Berkeley Lab has discovered the mechanism behind the RNA-guided cleavage of double-stranded DNA that is central to the bacterial acquired immunity system, possible providing genetic engineers with a promising alternative to artificial enzymes for gene targeting and genome editing in bacteria and other cell types.
Genetic engineers and genomics researchers should welcome the news from the Lawrence Berkeley National Laboratory (Berkeley Lab) where an international team of scientists has discovered a new and possibly more effective means of editing genomes. This discovery holds potentially big implications for advanced biofuels and therapeutic drugs, as genetically modified microorganisms, such as bacteria and fungi, are expected to play a key role in the green chemistry production of these and other valuable chemical products.
Jennifer Doudna, a biochemist with Berkeley Lab’s Physical Biosciences Division and professor at the University of California (UC) Berkeley, helped lead the team that identified a double-RNA structure responsible for directing a bacterial protein to cleave foreign DNA at specific nucleotide sequences. Furthermore, the research team found that it is possible to program the protein with a single RNA to enable cleavage of essentially any DNA sequence.
“We’ve discovered the mechanism behind the RNA-guided cleavage of double-stranded DNA that is central to the bacterial acquired immunity system,” says Doudna, who holds appointments with UC Berkeley’s Department of Molecular and Cell Biology and Department of Chemistry, and is an investigator with the Howard Hughes Medical Institute (HHMI). “Our results could provide genetic engineers with a new and promising alternative to artificial enzymes for gene targeting and genome editing in bacteria and other cell types.”
Doudna is one of two corresponding authors of a paper in the journal Science describing this work titled “A programmable dual RNA-guided DNA endonuclease in adaptive bacterial immunity.” The second corresponding author is Emmanuelle Charpentier of the Laboratory for Molecular Infection Medicine at Sweden’s Umeå University. Other co-authors of the paper were Martin Jinek, Krzysztof Chylinski, Ines Fonfara and Michael Hauer.
Bacterial and archaeon microbes face a never-ending onslaught from viruses and invading circles of nucleic acid known as plasmids. To survive, the microbes deploy an adaptive-type nucleic acid-based immune system that revolves around a genetic element known as CRISPR, which stands for Clustered Regularly Interspaced Short Palindromic Repeats. Through the combination of CRISPRs and associated endonucleases, called CRISPR-associated – “Cas” – proteins, bacteria and archaeons are able to utilize small customized crRNA molecules (for CRISPR-derived RNA) to target and destroy the DNA of invading viruses and plasmids.
There are three distinct types of CRISPR/Cas immunity systems. Doudna and her colleagues studied the Type II system which relies exclusively upon one family of endonucleases for the targeting and cleaving of foreign DNA, the Cas9 proteins.
“For the Type II CRISPR/Cas system, we found that crRNA connects via base-pairs with a trans-activating RNA (tracrRNA), to form a two-RNA structure,” Doudna says. “These dual RNA molecules (tracrRNA:crRNA) direct Cas9 proteins to introduce double-stranded DNA breaks at specific sites targeted by the crRNA-guide sequence.”

Doudna and her colleagues demonstrated that the dual tracrRNA:crRNA molecules can be engineered as a single RNA chimera for site-specific DNA cleavage, opening the door to RNA-programmable genome editing.
“Cas9 binds to the tracrRNA:crRNA complex which in turn directs it to a specific DNA sequence through base-pairing between the crRNA and the target DNA,” Doudna says. “Microbes use this elegant mechanism to cleave and destroy viruses and plasmids, but for genome editing, the system could be used to introduce targeted DNA changes into the genome.
Doudna notes that the “beauty of CRISPR loci” is that they can be moved around on plasmids.
“It is well-established that CRISPR systems can be transplanted into heterologous bacterial strains,” she says. “Also, there is evidence to suggest that CRISPR loci are horizontally transferred in nature.”
Doudna and her colleagues are now in the process of gathering more details on how the RNA-guided cleavage reaction works and testing whether the system will work in eukaryotic organisms including fungi, worms, plants, and human cells.
“Although we’ve not yet demonstrated genome editing, given the mechanism we describe it is now a very real possibility,” Doudna says.
Reference: “A Programmable Dual-RNA–Guided DNA Endonuclease in Adaptive Bacterial Immunity” by Martin Jinek, Krzysztof Chylinski, Ines Fonfara, Michael Hauer, Jennifer A. Doudna and Emmanuelle Charpentier, 28 June 2012, Science.
DOI: 10.1126/science.1225829
This work was funded primarily by the Howard Hughes Medical Institute, the Austrian Science Fund, and the Swedish Research Council.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
1 Comment
This brilliant team is one strong beacon of hope for the families of children afflicted by complicated congenital heart defects, and an inspiration to those who open their minds to the possibilities. Thank you for what you do, Jennifer and team. Where can I get an autograph for “A Crack in Creation”, the book?
your admirer in time, Kyram.