“Does the flap of a butterfly’s wings in Brazil set off a tornado in Texas?” — Edward Lorenz, at the 139th meeting of the American Association for the Advancement of Science
Scientists have recently discovered the mechanism by which a minuscule change in 3 atoms in a protein molecule can affect immune signaling in cells. This ‘butterfly effect’ is used by the bacterium, Shigella flexneri, to survive within the host cells that it infects.
Ranabir Das’ team at the National Center for Biological Sciences (NCBS), Bangalore, has found that a tiny change in the protein UBC13, caused by a bacterial enzyme, creates a cascade of small atomic alterations that add up until they prevent UBC13 from binding to a partner protein, TRAF6. Without the UBC13-TRAF6 complex, the host cell is unable to begin signaling for an immune response against the bacteria. This study examines the mechanics of a subtle alteration—involving the loss of just 3 atoms—and investigates how it can have far-reaching consequences.
According to the chaos theory in mathematics, a minute change such as the ‘flap of a butterfly’s wing’ could cause huge changes elsewhere. This seems to hold true at much smaller scales too—for example, within a cell. Scientists from the National Center for Biological Sciences (NCBS), Bangalore, have now found how a minuscule atomic change in a protein molecule enables the bacterium to shut down its host’s immune signaling system.
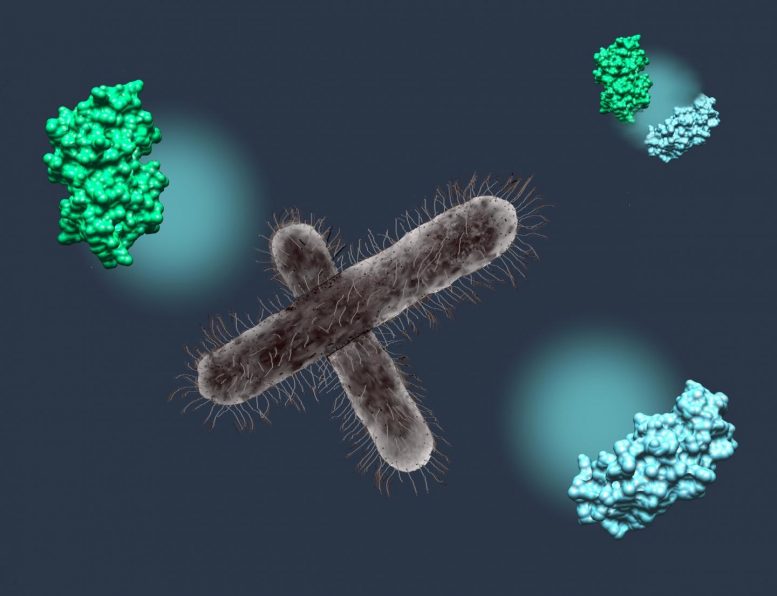
Shigella flexneri is a sneaky and highly infective bacteria. The organisms, which cause diarrhea in humans, first attach to the cells lining the host’s gut. Then using a needle-like apparatus, S. flexneri begin pumping in their secret weapon—an enzyme that alters a single amino acid in the host cell protein UBC13 (UBiquitin Conjugating enzyme E2 13). This change renders UBC13 unable to bind to a partner protein TRAF6 (TumouR necrosis factor Associated Factor 6) and effectively blocks the host cell from signaling for an inflammatory response. Subsequently, Shigella penetrates into the host cells to multiply.
Before now, little was known about how such a subtle effect—a change in just 3 atoms in a protein molecule containing roughly 3000 atoms—could shut down a host’s immune response.
A new study by Ranabir Das’ team at NCBS, however, has shown that when an amine group (formed of one nitrogen and two hydrogen atoms) is removed from a single amino acid in the host protein UBC13, a series of relatively small atomic changes pile up to block a key step in the immune response pathway. The work, which received funding support from the Tata Institute of Fundamental Research and the Department of Biotechnology, India, has been published as a paper in the journal eLife.
“We used a combination of structural studies, computational modeling, and enzymatic experiments to find that the atomic changes in UBC13 do not alter its structure. Rather, these changes completely disrupt its ability to bind its partner protein, TRAF6,” says Priyesh Mohanty, who is part of Das’ team, and the first author in the paper that describes these results.
When an amine group (formed of one nitrogen and two hydrogen atoms) is removed from a single amino acid in the host protein UBC13, a series of relatively small atomic changes pile up to block a key step in the immune response pathway.
The 14th amino acid in UBC13, an arginine residue (Arginine14), is critical in forming a ‘salt bridge’—a bond between oppositely charged ions—with TRAF6. This salt bridge between the surfaces of the two proteins is necessary to stabilize and maintain the UBC13-TRAF6 complex, which in turn, plays a pivotal role in immune response signaling. When the bacterial enzyme, named Ospl, deamidates a glutamine residue (Glutamine100), a neutral amine group is replaced by a negatively charged hydroxyl group. Now, this group literally steals away Arginine14 from the UBC13-TRAF6 inter-molecular salt bridge, to form an intra-molecular salt bridge with itself. This, in turn, compromises the transient interactions between UBC13 and TRAF6, which are necessary in forming the final complex. Finally, the new negative charge on the UBC13 surface creates a transient repulsive force between the UBC13 and TRAF6 proteins.
The salt bridge and the surface attractive forces collectively create a force powerful enough to steady (or hold) the UBC13-TRAF6 complex. With the loss of these forces, the complex falls apart, and without the complex, the immune signal against the bacteria is blocked.
“This investigation has really helped us understand the role of individual amino acids in the associations between proteins and how proteins function within the cell,” say Mohanty and Das. “To the best of our knowledge, our study is the first to provide a mechanism by which glutamine deamidation of a target protein hinders its function inside the host cell. Interestingly, this mechanism allows the bacteria to attenuate the inflammatory response and promotes its ability to survive in a human host,” they add.
Currently, Shigella infections cause almost 0.2 million deaths due to diarrhea every year, of which one-third occur in infants. Several multidrug-resistant strains of these bacteria have also appeared, and yet there are no vaccines or drugs to prevent or limit the spread of Shigella outbreaks.
“It is, therefore, very important to study what mechanism Shigella uses to attenuate inflammatory responses in human hosts. Such studies may help researchers to identify new drugs that could control Shigella outbreaks in humans,” say Mohanty and Das.
Reference: “Deamidation disrupts native and transient contacts to weaken the interaction between UBC13 and RING-finger E3 ligases” by Priyesh Mohanty, Rashmi Agrata, Batul Ismail Habibullah, Arun Geetha Surendran and Ranabir Das, 22 October 2019, eLife.
DOI: 10.7554/eLife.49223
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.