
Chinese scientists have developed a novel AI-guided protein engineering strategy that enables faster, cost-effective, and highly accurate protein design.
A group of scientists in China, led by Professor Caixia Gao from the Institute of Genetics and Developmental Biology (IGDB) at the Chinese Academy of Sciences, has introduced an innovative method that may significantly advance protein engineering. This new technique, known as AI-informed Constraints for protein Engineering (AiCE), accelerates the evolution of proteins by combining structural and evolutionary insights within a standard inverse folding model. Remarkably, it does so without requiring the development or training of dedicated artificial intelligence (AI) systems.
The research, published in Cell on July 7, takes aim at several long-standing limitations in conventional protein engineering practices.
Ideally, protein engineering would achieve high-performance results with minimal complexity. However, most current methods struggle with high costs, low efficiency, and limited scalability. Even though AI-based approaches offer improvements, they typically demand significant computational resources. This has created a need for solutions that are both powerful and practical, allowing for more widespread use without sacrificing accuracy.
Introducing AiCEsingle: Precision Through Structural Constraints
In this study, the researchers first developed AiCEsingle, a module designed to predict high-fitness (HF) single amino acid substitutions. It enhances prediction accuracy by extensively sampling inverse folding models—AI models that generate compatible amino acid sequences based on protein 3D structures—while incorporating structural constraints.
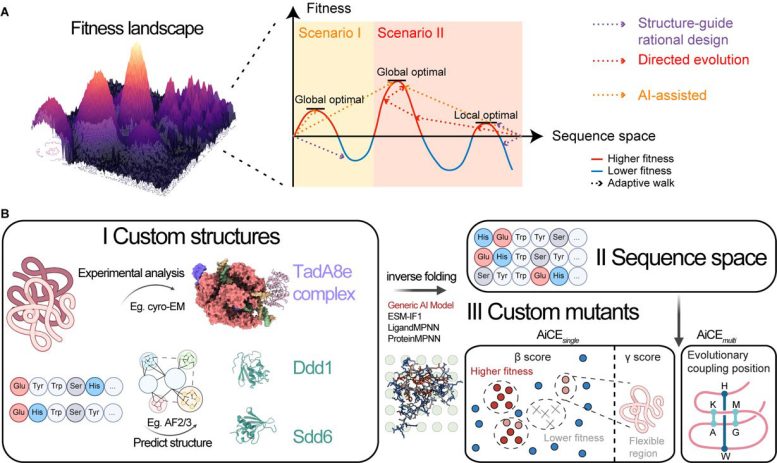
Benchmarking against 60 deep mutational scanning (DMS) datasets demonstrated that AiCEsingle outperforms other AI-based methods by 36–90%. Its effectiveness for complex proteins and protein–nucleic acid complexes was also validated. Notably, incorporating structural constraints alone yielded a 37% improvement in accuracy.
To address the challenge of negative epistatic interactions in combinatorial mutations, the researchers developed the AiCEmulti module, which integrates evolutionary coupling constraints. This allows for accurate prediction of multiple high-fitness mutations at minimal computational cost, expanding the tool’s versatility and practical utility.
Engineering a New Generation of Functional Proteins
Using the AiCE framework, the researchers successfully evolved eight proteins with diverse structures and functions, including deaminases, nuclear localization sequences, nucleases, and reverse transcriptases. These engineered proteins have enabled the creation of several next-generation base editors for applications in precision medicine and molecular breeding. These include: enABE8e, a cytosine base editor with a ~50% narrower editing window; enSdd6-CBE, an adenine base editor with 1.3× higher fidelity; and enDdd1-DdCBE, a mitochondrial base editor showing a 13× increase in activity.
AiCE represents a simple, efficient, and broadly applicable strategy for protein engineering. By unlocking the potential of existing AI models, it offers a promising new direction for the field and enhances the interpretability of AI-driven protein redesign.
Reference: “Advancing protein evolution with inverse folding models integrating structural and evolutionary constraints” by Hongyuan Fei, Yunjia Li, Yijing Liu, Jingjing Wei, Aojie Chen and Caixia Gao, 7 July 2025, Cell.
DOI: 10.1016/j.cell.2025.06.014
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
1 Comment
Anyone ever heard of Prions?