
Researchers uncovered four distinct, shifting layers of neuron types inside the hippocampal CA1 region—an area central to memory, navigation, and emotion.
Using high-resolution RNA imaging, they created a remarkably clear cellular map that reveals a hidden structural order previously missed. These layered “stripes” of specialized neurons help explain why different parts of CA1 support different behaviors and why certain cells are more vulnerable in diseases like Alzheimer’s and epilepsy.
Hidden Structure in the Brain’s Memory Hub
Researchers at the Mark and Mary Stevens Neuroimaging and Informatics Institute (Stevens INI) at the Keck School of Medicine of USC have identified a previously unknown pattern of organization in one of the brain’s most important areas for learning and memory.
The study, published today (December 3) in Nature Communications, reveals that the CA1 region of a mouse’s hippocampus, a structure vital for memory formation, spatial navigation, and emotions, has four distinct layers of specialized cell types. This discovery changes our understanding of how information is processed in the brain and could explain why certain cells are more vulnerable in diseases like Alzheimer’s and epilepsy.
“Researchers have long suspected that different parts of the hippocampus’ CA1 region handle different aspects of learning and memory, but it wasn’t clear how the underlying cells were arranged,” said Michael S. Bienkowski, PhD, senior author of the study and assistant professor of physiology and neuroscience and of biomedical engineering.
“Our study shows that CA1 neurons are organized into four thin, continuous bands, each representing a different neuron type defined by a unique molecular signature. These layers aren’t fixed in place; instead, they subtly shift and change in thickness along the length of the hippocampus. This shifting pattern means that each part of CA1 contains its own mix of neuron types, which helps explain why different regions support different behaviors. This may also clarify why certain CA1 neurons are more vulnerable in conditions like Alzheimer’s disease and epilepsy: if a disease targets one layer’s cell type, the effects will vary depending on where in CA1 that layer is most prominent.”
Mapping Neuron Types With High-Resolution RNA Imaging
To reveal this hidden organization, the scientists used an RNA labeling technique called RNAscope together with high-resolution microscopy. This combination allowed them to see individual molecules of gene activity inside mouse CA1 cells and to distinguish different neuron types based on their genetic signatures. In a total of 58,065 CA1 pyramidal cells, they identified more than 330,000 RNA molecules, which are the genetic messages that indicate when and where genes are active. By tracking these patterns across the tissue, the team produced a detailed map that outlines the boundaries between different nerve cell types throughout the CA1 region of the hippocampus.
The analysis showed that CA1 is not a loose mixture of cells but is instead made up of four continuous layers of nerve cells, each defined by its own set of active genes. Viewed in three dimensions, the layers form extended sheets that vary slightly in thickness and structure as they run along the length of the hippocampus. This clear, banded arrangement helps reconcile earlier studies that suggested CA1 was organized more like a gradual blend or mosaic of cell types.

“Stripes” of Neurons Reveal Hidden Brain Architecture
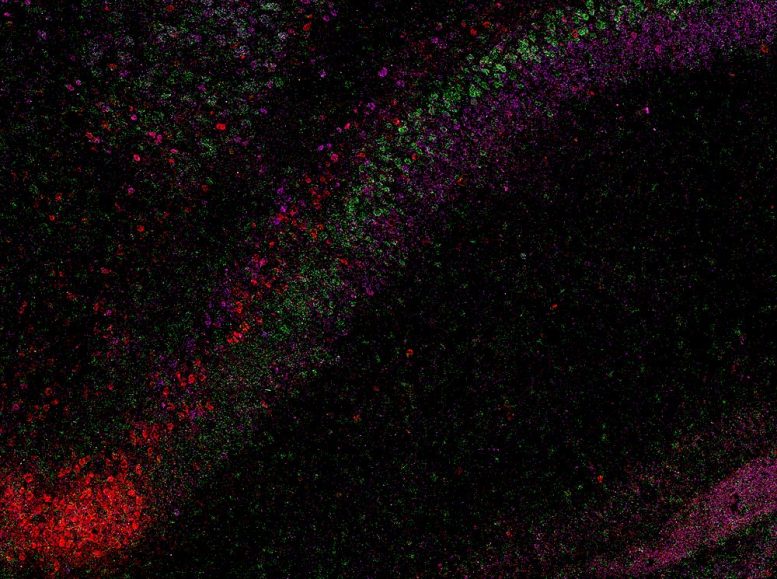
“When we visualized gene RNA patterns at single-cell resolution, we could see clear stripes, like geological layers in rock, each representing a distinct neuron type,” said Maricarmen Pachicano, doctoral researcher at the Stevens INI’s Center for Integrative Connectomics and co–first author of the paper. “It’s like lifting a veil on the brain’s internal architecture. These hidden layers may explain differences in how hippocampal circuits support learning and memory.”
The hippocampus is one of the first brain regions to deteriorate in Alzheimer’s disease and is also involved in epilepsy, depression, and several other neurological and psychiatric conditions. By showing that CA1 is arranged in four specific layers, the new work offers a guide for pinpointing which neuron types are most at risk in each disorder.
Linking Layered CA1 Structure to Brain Disease
“Discoveries like this exemplify how modern imaging and data science can transform our view of brain anatomy,” said Arthur W. Toga, PhD, director of the Stevens INI and the Ghada Irani Chair in Neuroscience at the Keck School of Medicine of USC. “This work builds on the Stevens INI’s long tradition of mapping the brain at every scale, from molecules to whole networks, and will inform both basic neuroscience and translational studies targeting memory and cognition.”
A 3D Hippocampal Atlas for the Research Community
The researchers compiled their findings into a CA1 cell-type atlas based on data from the Hippocampus Gene Expression Atlas (HGEA). This resource is freely available to scientists worldwide. It includes interactive 3D visualizations accessible through the Schol-AR augmented-reality app, developed at the Stevens INI, which allows users to explore the layered hippocampal structure in fine detail.
Because the layered pattern seen in mice resembles what has been observed in primate and human brains, including similar changes in CA1 thickness, the team believes this organization may be common across many mammalian species. Although further studies are needed to confirm the same structure in humans, the discovery provides a strong starting point for comparative and translational research on how hippocampal architecture supports memory and other mental functions.
“Understanding how these layers connect to behavior is the next frontier,” Bienkowski said. “We now have a framework to study how specific neuron layers contribute to such different functions like memory, navigation, and emotion, and how their disruption may lead to disease.”
Reference: “Laminar organization of pyramidal neuron cell types defines distinct CA1 hippocampal subregions” by Maricarmen Pachicano, Shrey Mehta, Angela Hurtado, Tyler Ard, Jim Stanis, Bayla Breningstall and Michael S. Bienkowski, 3 December 2025, Nature Communications.
DOI: 10.1038/s41467-025-66613-y
In addition to Bienkowski and Pachicano, the study’s other authors include Shrey Mehta, Angela Hurtado, Tyler Ard, Jim Stanis, and Bayla Breningstall.
This work was supported by the National Institutes of Health/National Institute of Aging (K01AG066847, R36AG087310-01, supplement P30-AG066530-03S1), National Science Foundation (grant 2121164), and funding from the USC Center for Neuronal Longevity. Research data reported in this publication was supported by the Office of the Director, National Institutes of Health under award number S10OD032285.
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.