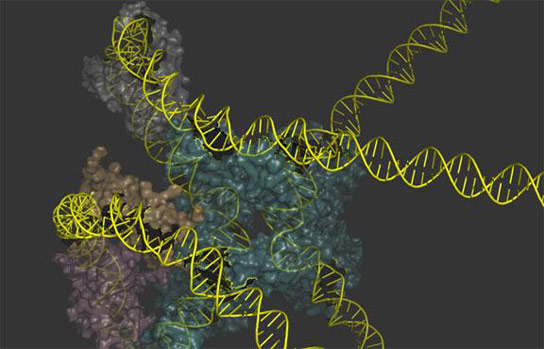
After more than 50 years of research, biologists now have a picture of the molecular machines that allow the lambda virus to insert or remove DNA from the cells that it infects.
For half a century biologists have studied the way that the lambda virus parks DNA in the chromosome of a host E. coli bacterium and later extracts it as a model reaction of genetic recombination. But for all that time, they could never produce an overall depiction of the protein-DNA machines that carry out the work. In a pair of back-to-back papers in the Proceedings of the National Academy of Sciences, scientists produce those long-sought renderings and describe how they figured out what they should look like.
The team of researchers at Brown University and the University of Pennsylvania mapped the specific ways that the recombination proteins in the machines bridge separate sites along the DNA of the host and the virus. To understand what that means is to appreciate the evolved “ingenuity” of the lambda virus that has made it the subject of 50 years of intriguing study. Thankfully lambda is benign, although it has medically nasty cousins that work much the same way.
Lambda, said co-corresponding author Arthur Landy, professor emeritus of biology at Brown, is not one of those devil-may-care viruses that just smashes and grabs a host cell for immediate reproduction. Instead, it senses the physiology of its E. coli host and waits for the cell to be healthy before striking, perhaps several generations of cell division later. To execute this more refined strategy it assembles an ensemble of proteins — a “machine” —to insert its DNA in a precise location in the DNA of the host (to ensure that its DNA will persist for many generations of host cells), and another machine to extract the DNA when the chosen moment of ideal host health arrives.
“The viruses make a ‘decision’ when they infect a cell as to whether or not this is a good time to lyse the cell and make more virus or whether it would be more propitious to integrate their chromosome into the E. coli chromosome, turn off their genes and sit there for generations,” Landy said. “Then when things look good again they use a different but related pathway to excise their chromosome, in order to make more virus and kill the cell.”
The way lambda senses the health status of the host is built right into the machines. The machines incorporate key DNA-bending proteins that are made by E. coli to regulate the expression of its own genes. Therefore the levels of these proteins also reflect the physiological state of the cell, Landy said. While some recombination proteins bind to just one site in each DNA, lambda and its ilk go through the trouble of latching on to two distant sites in the DNA. The reason is to make the whole reaction depend on the presence of these key DNA-bending proteins to bend the sites into closer proximity. Without those proteins, the machines can’t work.
“It makes the system gratuitously dependent on the proteins of the cell which serve as reporters of how well the cell is doing and where it is in its life cycle,” Landy said. “This makes it exquisitely sensitive to the physiology of the cell.”
Mapping bridges
Biologists already knew all this, but they had never quite figured out how the recombination proteins in the machines bridge the two DNA sites, (i.e., which sites were bound or bridged by the same recombination protein). Therefore they couldn’t really figure out what the whole machines looked like. Direct imaging tools such as crystallography never worked on the whole machines because it has too many forms, Landy said, and nuclear magnetic resonance never worked because it was too hard to make enough of the machines in a high enough concentration.
So the team did a couple of other experiments to figure out the bridging. One used chemicals and the other used genetics, but in each case they were essentially tagging different pairs of locations on the proteins and the DNA to see if any of those pairs, upon being connected, would produce a tell-tale effect: In the case of the chemistry the tell-tale was a change in the structural properties of the complex; in the case of the genetics it was a successful integration or excision reaction.
It’s a bit like trying to wire a battery into a light bulb circuit where there are many pairs of loose wires, but only two that actually connect to the bulb. When one finally connects (or “bridges”) the two correct loose wires with the battery’s electrodes, the light bulb lights and confirms where those correct two wires are.
They combined their new mappings, including distance measurements within the protein-DNA complexes using fluorescent tags, with all of the other biochemical and structural information biologists have learned over the last 50 years, into a computer model to render the overall depictions of the machines.
They confirmed in further research that the models they generated explained and agreed with much of the observed behavior of the machines.
So now, long after they began to understand how worthwhile the lambda virus was to study, researchers can finally see what these model DNA insertion and removal machines look like. That information will serve to make lambda an even more powerful model for research and teaching, Landy said.
The lead authors of the paper that focused on mapping the bridging are Wenjun Tong and David Warren. Other Brown authors, in addition to Landy are Nicole Seah and Gurunathan Laxmikanthan. The co-corresponding author at Penn is Gregory Van Duyne.
Seah is the lead author of the paper that resolves the overall architecture of the machines. Warren, Tong, and Laxmikanthan are co-authors and Van Duyne is again a co-corresponding author.
References:
“Mapping the λ Integrase bridges in the nucleoprotein Holliday junction intermediates of viral integrative and excisive recombination” by Wenjun Tong, David Warren, Nicole E. Seah, Gurunathan Laxmikanthan, Gregory D. Van Duyne and Arthur Landy, 11 August 2014, PNAS.
DOI: 10.1073/pnas.1413007111
“Nucleoprotein architectures regulating the directionality of viral integration and excision” by Nicole E. Seah, David Warren, Wenjun Tong, Gurunathan Laxmikanthan, Gregory D. Van Duyne and Arthur Landy, 11 August 2014, PNAS.
DOI: 10.1073/pnas.1413019111
The National Institutes of Health supported the research (grants: GM062723, GM033928, and GM108751).
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.