
The Telomere-to-Telomere consortium has fully sequenced the Y chromosome, uncovering 41 new genes and adding 30 million new bases to the human genome. This breakthrough will impact studies on reproduction, evolution, and human population changes, and correct previous misidentifications of bacterial DNA. Future endeavors aim to integrate this data into the human pangenome for global research collaborations.
For decades, the Y chromosome – one of the two human sex chromosomes – has been notoriously challenging for the genomics community to sequence due to the complexity of its structure. Now, this elusive area of the genome has been fully sequenced, a feat that finally completes the set of end-to-end human chromosomes and adds 30 million new bases to the human genome reference, mostly from challenging-to-sequence satellite DNA. These bases reveal 41 additional protein-coding genes, and provide crucial insight for those studying important questions related to reproduction, evolution, and population change.
Researchers from the Telomere-to-Telomere (T2T) consortium, which is co-led by the University of California, Santa Cruz Assistant Professor of Biomolecular Engineering Karen Miga, announced this achievement in a new paper to be published today (August 23) in the journal Nature. The complete, annotated Y chromosome reference is available for use on the UCSC Genome Browser and can be accessed via Github.
“Just a few years ago, half of the human Y chromosome was missing [from the reference] – the challenging, complex satellite areas,” said Monika Cechova, co-lead author on the paper and postdoctoral scholar in biomolecular engineering at UCSC. “Back then we didn’t even know if it could be sequenced, it was so puzzling. This is really a huge shift in what’s possible.”
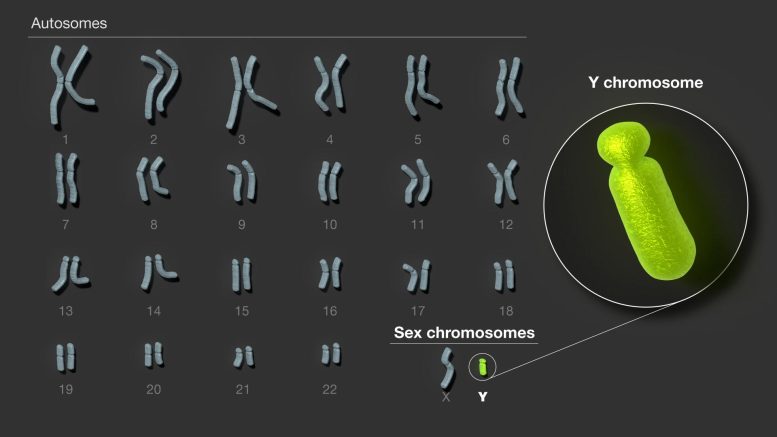
Decoding the Y Chromosome
When scientists and clinicians study an individual’s genome, they compare the individuals’ DNA to that of a standard reference to determine where there is variation. Until now, the Y chromosome portion of the human genome has contained large gaps which made it difficult to understand variation and associated disease.
The structure of the Y chromosome has been challenging to decode because some of the DNA is organized in palindromes – long sequences that are the same forward and backward – spanning up to more than a million base pairs. Moreover, a very large part of the Y chromosome that was missing from the previous version of the Y reference is satellite DNA – large, highly repetitive regions of non-protein-coding DNA. On the Y chromosome, two satellites are interlinked with each other, further complicating the sequencing process.

The researchers were able to achieve a gapless read of the Y chromosome due to advances in long-read sequencing technology and new, innovative computational assembly methods that could deal with the repetitive sequences and transform the raw data from sequencing into a usable resource. These new method assemblies allowed the team to tackle some of the particularly challenging aspects of the Y chromosome, such as pinpointing precisely where an inversion occurs in a palindromic sequence — a technique that can be used to find other inversions. The methods established in the paper will allow scientists to complete more end-to-end reads of human Y chromosomes to get a better understanding of how this genetic material affects the diverse human population.
“It was the Y chromosome that lacked the most sequences from the previous reference genome,” said Arang Rhie, a staff scientist at the National Human Genome Research Institute and the paper’s lead author. “It was always irritating knowing we were missing half the Y whenever we tried to do any reference-based analysis. I was really excited to curate the first complete Y, to see what we were actually missing, and what we can now do.”
The Path to Completion
In 2018, Miga and her colleagues released the first complete map of a human centromere on the Y chromosome. This first gap closure was credited to access to ultra-long data, which builds on nanopore sequencing technology that has its origins here at UCSC. It was clear at that point that emerging technology and high-coverage long-read datasets had the potential to complete entire chromosomes end to end, which led to the launch of the T2T Consortium, co-led by Phillippy and Miga.
Now, just five years later, the T2T consortium has filled in 30 million additional base pairs, in addition to the first fully sequenced human genome (all the autosomes and the X chromosome) that was released in 2022.

Enabling New Research and Discoveries
The Y chromosome is most commonly associated with male individuals, but may be found in others, such as intersex people. The sex characteristics regulated by DNA on the Y chromosome are also not equivalent to an individual’s gender identity. While there are relatively few genes on the Y chromosome, the ones that are present are complex and dynamic, and code for important functions such as spermatogenesis, the production of sperm. The complete Y chromosome reference will allow scientists to better study a myriad of features about this part of the human genome in a way that has never before been possible.
The complex structure of the Y chromosome has lent itself to rapid evolution within its gene families. In fact, the Y chromosome is the most rapidly changing human chromosome, and even the most rapidly changing chromosome among great apes. This means two healthy people’s Y chromosomes can look very different – for example, one person might have 40 copies of one gene, while another person has 19 copies. This evolution can now be better studied using the new reference and the established methods for sequencing Y chromosomes. This could be the future focus of in vitro fertilization clinics or other research on reproduction and infertility.
The end-to-end Y chromosome sequence is a hugely important resource for those studying human population evolution and drift. This is because the Y chromosome is inherited from generation to generation in one group of genetic material, with very little recombination outside of that group, unlike the autosomes and genes on the human X chromosome which often recombine and share genetic material with each other. Having a clearer picture of the Y chromosome makes it easier to track genes across generations of inheritance and learn how the location and content of genes has changed over time.
The 30 million new bases added to the Y chromosome reference will also be crucial for studying genome evolution. It will now be possible to study specific and unique Y chromosome sequence patterns, such as the structure of the two satellites and the location and copy numbers of the genes. Even within the Y chromosome, the genes are split into several regions, which are very different from each other in terms of content, structure, and evolutionary history. Understanding rates of change on the Y chromosome and how to interpret this change are intriguing questions that will now be possible to study using the techniques developed in this paper to completely sequence human Y chromosomes.
The richer reference that includes the full sequence of the Y chromosome satellite DNA will also allow scientists to better understand the evolutionary relationship of these sequences with satellite DNA found elsewhere on the genome.
“It is exciting to be able to finally see these sequences in heterochromatic [densely-packed] regions for the first time. Finally, we can design experiments to test the impact and function of these previously unexplored parts of the Y chromosome,” Miga said.
It’s been shown that people with Y chromosomes can lose some or all of that genetic material as they age, but scientists have never fully understood why this happens and the effects it may have. The complete Y chromosome reference may help to illuminate this mystery. It will also be easier to study conditions and disorders that are linked to the Y chromosome, such as the lack of sperm production which leads to infertility.
Contamination in Bacterial Genomes
An unexpected finding from this paper was that Y chromosome DNA has been repeatedly mistaken to be bacterial DNA in past studies due to the incomplete removal of human contamination in bacterial DNA. This discovery promises to improve the study of bacterial species’ genomes.
Human DNA can appear as a contaminant in the genomic samples of bacterial species because the bacterial DNA is often taken from swipes off of human skin. Scientists use the current human genome reference to identify which sequences come from human contamination and remove those, leaving just the bacterial DNA for their study. But, because large parts of the human Y chromosome were missing from the past human reference, scientists were not able to identify them as human and thus mistook them to be part of the DNA of the species they were studying.
This paper finds evidence that about 5,000 bacterial genomes in a common database likely contained contamination matching human Y sequences. The groups studying these bacterial species can use the updated Y reference to correctly remove all human contamination from their reference genomes and get a clearer understanding of the bacterial genome.
“That was a surprising thing,” Rhie said. “People were guessing at it, but no one could prove that this was happening until now.”
Pangenome Y and Future Directions
While the complete human Y chromosome will open the door to many new discoveries, the researchers plan to further improve the study of this region by including the Y chromosome in future versions of the human pangenome. The pangenome is a new reference for genomics that combines the genomic information of multiple people from various ancestral backgrounds to ultimately enable more equitable research and clinical discoveries such as helping to diagnose disease, predict medical outcomes, and guide treatments.
In collaboration with the Human Pangenome Reference Consortium, the researchers plan to incorporate complete Y chromosome sequences into the individual genomes that make up the pangenome. This will help scientists understand how the Y chromosome varies among people of different ancestral backgrounds and provide a better point of reference for understanding the Y across the diversity of the human population.
The researchers hope to be able to collaborate with scientists around the world to enable others to complete Y chromosome sequencing.
“We aim to make these data widely accessible,” Miga said. “By creating and sharing these important catalogs of genetic differences on the Y chromosome, we can expand genetic studies of human disease and provide new insights into basic biology.”
For more on this breakthrough, see Complete Human Y Chromosome Sequence Assembled for the First Time.
Reference: “The complete sequence of a human Y chromosome” by Arang Rhie, Sergey Nurk, Monika Cechova, Savannah J. Hoyt, Dylan J. Taylor, Nicolas Altemose, Paul W. Hook, Sergey Koren, Mikko Rautiainen, Ivan A. Alexandrov, Jamie Allen, Mobin Asri, Andrey V. Bzikadze, Nae-Chyun Chen, Chen-Shan Chin, Mark Diekhans, Paul Flicek, Giulio Formenti, Arkarachai Fungtammasan, Carlos Garcia Giron, Erik Garrison, Ariel Gershman, Jennifer L. Gerton, Patrick G. S. Grady, Andrea Guarracino, Leanne Haggerty, Reza Halabian, Nancy F. Hansen, Robert Harris, Gabrielle A. Hartley, William T. Harvey, Marina Haukness, Jakob Heinz, Thibaut Hourlier, Robert M. Hubley, Sarah E. Hunt, Stephen Hwang, Miten Jain, Rupesh K. Kesharwani, Alexandra P. Lewis, Heng Li, Glennis A. Logsdon, Julian K. Lucas, Wojciech Makalowski, Christopher Markovic, Fergal J. Martin, Ann M. Mc Cartney, Rajiv C. McCoy, Jennifer McDaniel, Brandy M. McNulty, Paul Medvedev, Alla Mikheenko, Katherine M. Munson, Terence D. Murphy, Hugh E. Olsen, Nathan D. Olson, Luis F. Paulin, David Porubsky, Tamara Potapova, Fedor Ryabov, Steven L. Salzberg, Michael E. G. Sauria, Fritz J. Sedlazeck, Kishwar Shafin, Valery A. Shepelev, Alaina Shumate, Jessica M. Storer, Likhitha Surapaneni, Angela M. Taravella Oill, Françoise Thibaud-Nissen, Winston Timp, Marta Tomaszkiewicz, Mitchell R. Vollger, Brian P. Walenz, Allison C. Watwood, Matthias H. Weissensteiner, Aaron M. Wenger, Melissa A. Wilson, Samantha Zarate, Yiming Zhu, Justin M. Zook, Evan E. Eichler, Rachel J. O’Neill, Michael C. Schatz, Karen H. Miga, Kateryna D. Makova and Adam M. Phillippy, 23 August 2023, Nature.
DOI: 10.1038/s41586-023-06457-y
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
1 Comment
“The Y chromosome is most commonly associated with male individuals, but may be found in others, such as intersex people. The sex characteristics regulated by DNA on the Y chromosome are also not equivalent to an individual’s gender identity.”
Gotta love “Science” these days.
Wokeism injected at every possible moment.
There is no such thing as “Gender identity”, its a symptom of mental illness.