A mysterious RNA molecule found only in breast cancer led researchers on a six-year hunt that uncovered an entire hidden layer of cancer biology.
The story began with T3p, a tiny RNA molecule found in breast cancer tumors but not in healthy tissue. First described in 2018, T3p quickly stood out as something unusual. What followed was a six-year effort to systematically search for similar orphan non-coding RNAs (oncRNAs) across many cancers, determine which ones actively influence disease, and test whether they could be used to track cancer through simple blood tests.
In a paper published today, the researchers describe how this work evolved from large-scale analysis of cancer genome data to building machine learning tools, running functional experiments in mice, and ultimately confirming the clinical potential of these RNAs in nearly 200 breast cancer patients using blood samples.
Cancer-Specific RNAs Are Widespread
One of the first surprises was that T3p was not unique to breast cancer. By examining small RNA sequencing data from The Cancer Genome Atlas across 32 cancer types, the team identified roughly 260,000 cancer-specific small RNAs distributed across every cancer they studied. These RNAs, known as oncRNAs, appeared consistently in tumors but not in normal tissue.
Their presence followed clear patterns. Each cancer type produced its own distinct set of oncRNAs. Lung cancers showed different RNA profiles than breast cancers, for example. When these patterns were analyzed using machine learning, the models correctly identified cancer types with 90.9% accuracy. The results held up in a separate group of 938 tumors, where classification accuracy reached 82.1%.
RNA Patterns Reveal Cancer Identity
Differences were also found within individual cancer types. In breast cancer, basal tumors produced oncRNA patterns that differed from luminal tumors. This suggests that oncRNAs capture subtle differences in cancer cell state, including divisions that may not yet be fully recognized.
Taken together, the presence or absence of specific oncRNAs functions like a set of “digital molecular barcodes.” These barcodes reflect cancer identity at multiple levels, from broad tumor type to subtype and even underlying cellular behavior.
Some OncRNAs Actively Drive Cancer
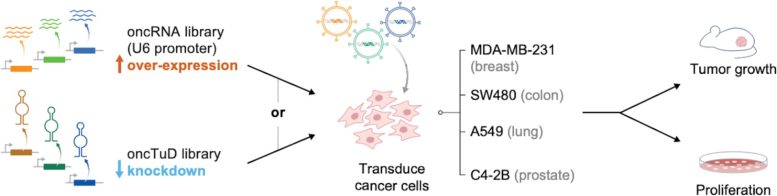
Beyond their value as markers, the researchers wanted to know whether oncRNAs could also influence how cancers grow. To test this, they built large screening libraries containing about 400 oncRNAs drawn from breast, colon, lung, and prostate cancers.
These oncRNAs were introduced into cancer cells using lentiviral vectors. Some were overexpressed, while others were reduced using “Tough Decoy” constructs. The modified cells were then implanted into mice to see which oncRNAs gave tumors a growth advantage.
About 5% of the oncRNAs tested produced clear effects in mouse models. Two breast cancer oncRNAs were studied in depth. One triggered epithelial-mesenchymal transition, a key step in cancer spread. The other activated E2F target genes, increasing cell division. Both led to faster tumor growth and greater metastatic spread in multiple cell line models.
When patient tumor data were examined, tumors expressing these same oncRNAs showed matching biological pathway changes. Observing the same effects in TCGA samples and experimental models strengthened confidence that these RNAs play real functional roles in cancer.
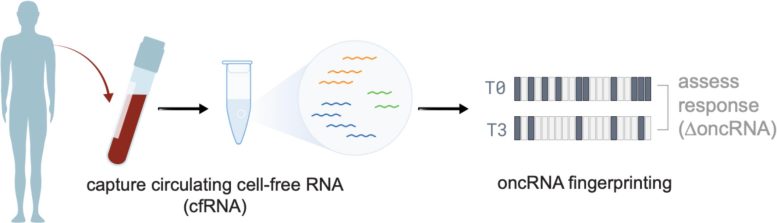
Cancer Cells Release OncRNAs Into Blood
The most clinically promising finding was that cancer cells release many of these RNAs into the bloodstream. Measuring them can provide insight into how patients are responding to treatment.
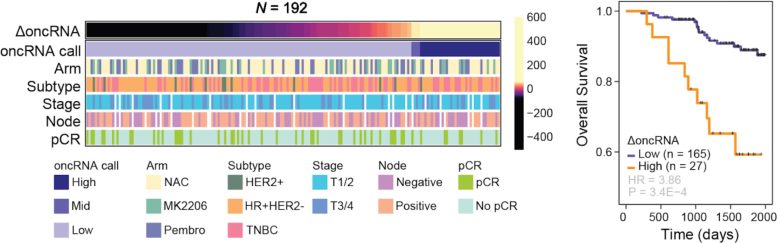
The team analyzed cell-free RNA from 25 cancer cell lines representing 9 tissue types and found that about 30% of oncRNAs are actively secreted. To test whether this mattered in patients, they studied blood serum from 192 breast cancer patients enrolled in the I-SPY 2 neoadjuvant chemotherapy trial. Blood was collected before and after treatment, and researchers calculated changes in total oncRNA burden (ΔoncRNA below).
That single measurement proved highly informative. Patients with high levels of oncRNA remaining after chemotherapy had nearly four times worse overall survival. This relationship remained strong even after accounting for standard clinical indicators such as pathologic complete response and residual cancer burden.
A New Path for Cancer Monitoring
This result was the researchers’ most ambitious goal. While oncRNAs were known to appear in blood, it was unclear whether they would reliably reflect patient outcomes. Detecting a strong signal using just 1 milliliter of serum exceeded expectations.
The finding addresses a long-standing challenge in breast cancer care. Monitoring minimal residual disease using cell-free DNA is difficult because many tumors shed very little DNA, particularly in early-stage disease. RNA-based monitoring may offer an advantage, since cancer cells actively release RNA rather than shedding DNA passively.
What Comes Next
Many important questions remain. Researchers still need to understand how functional oncRNAs work at the molecular level, including whether they interact with proteins or other RNAs. It is also unknown whether tracking oncRNA changes in real time could help guide treatment decisions, detect recurrence earlier, or improve patient stratification. Answering these questions will require larger studies and prospective clinical trials.
At the same time, translation is already underway. The finding that oncRNAs provide cancer-specific signals in blood is being developed further through collaboration with the biotech company Exai Bio (Hani is a co-founder), which is building AI-driven models and large datasets for cancer detection and classification.
Behind every data point is a patient who contributed a blood sample and took part in research with the hope of helping others. Keeping that human context in mind is a central motivation for the team.
The researchers believe oncRNAs represent a newly recognized class of cancer-emergent molecules that function both as disease drivers and as biomarkers. By making this resource open source, they hope it will enable new discoveries and push the field in unexpected directions.
Reference: “Systematic annotation of orphan RNAs reveals blood-accessible molecular barcodes of cancer identity and cancer-emergent oncogenic drivers” by Jeffrey Wang, Jung Min Suh, Brian J. Woo, Albertas Navickas, Kristle Garcia, Keyi Yin, Lisa Fish, Taylor Cavazos, Benjamin Hänisch, Daniel Markett, Gillian L. Hirst, Lamorna Brown-Swigart, Laura J. Esserman, Laura J. van ‘t Veer and Hani Goodarzi, 23 January 2026, Cell Reports Medicine.
DOI: 10.1016/j.xcrm.2025.102577
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.