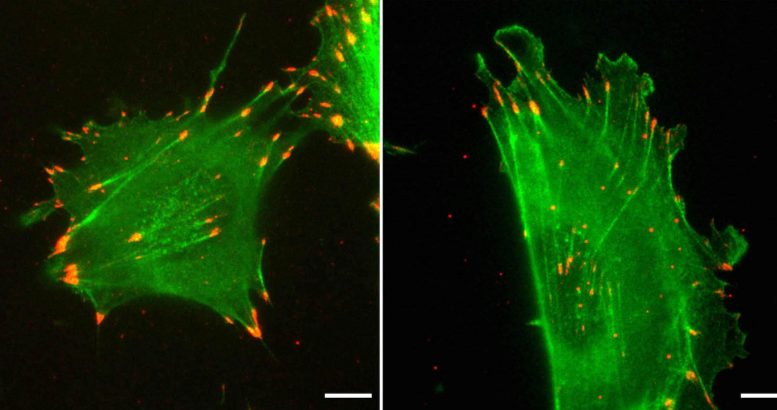
A University of Pennsylvania-led study shows how, despite having nearly identical amino acid sequences, two forms of the protein actin differ in function due to their distinct nucleotide sequences.
The protein actin is ubiquitous and essential for life. In mammals, every cell expresses two of its forms, beta-actin, and gamma-nonmuscle-actin. Despite having distinct roles, the two forms are nearly identical, sharing 99% of their amino acid sequence.
Research by Anna Kashina of Penn’s School of Veterinary Medicine and colleagues has shown that, contrary to scientific dogma, it’s not the slight differences in amino acid sequence that govern these proteins’ discrete functions in the cell. Rather, their nucleotide sequences — the “letters” that make up their DNA coding sequence, which differ by roughly 13% between the two forms — are responsible for their individual roles in organisms’ survival and cell migration.
And in a new study the researchers offer an explanation for why: Beta-actin mRNA is translated into protein faster than gamma-actin. Both forms help cells move, but beta-actin’s faster rate seems to cause cell to affix to a substrate more strongly, slowing down cell movement.
“On a global, philosophical level, this expands our understanding of genetic code,” says Kashina, a professor of biochemistry at Penn Vet and the senior author on the study, which was published in the journal eLife. “We used to believe that the role of the nucleotides was to encode amino acids, but now we see that, actually, proteins with the same amino acid sequence have different translation rates, and that makes a difference in their function.”
Kashina uses the term “silent code” to refer to the influence of these nucleotide differences. In earlier work, her team showed that, in mice, editing the amino acid sequence but maintaining the silent nucleotide code could cause gamma-actin to behave like beta-actin in the body. Normally, mice lacking beta-actin would die before birth, but the researchers showed that performing gene editing to the beta-actin gene so it had the same amino acid sequence as gamma-actin kept mice alive thanks to the nucleotide differences.
A finding from an earlier paper, also published in eLife, motivated the new work. In that earlier study the researchers found that beta-actin RNA had a much higher density of ribosomes than that of gamma-actin. Ribosomes are crucial to the synthesis of protein from RNA, leading the scientists to hypothesize that this difference in protein translation rate could be responsible for the different functions between gamma and beta-actin.
To test their idea, they used cell lines to express only the coding sections of beta- and gamma-actin in mouse cells, as well as their edited versions: the beta-actin that had been edited to have the same amino acid sequence as gamma-actin and vice versa for gamma-actin.
When put to the test in a wound-healing experiment, the researchers found that the nucleotide sequence was paramount in determining the speed at which actin facilitated cell movement. Cells expressing only the typical beta-actin migrated at typical rates, but gamma-actin-expressing cells moved twice as fast. The cells containing edited versions of actin proved that this difference is nucleotide-sequence dependent. Beta-actin edited to have the amino acid sequence of gamma moved like gamma-actin-expressing cells, and those with gamma-actin edited to have the amino acid sequence of beta-actin moved at the rate of beta-actin-expressing cells.
These results surprised the researchers since they expected the higher density of ribosomes in beta-actin mRNA could support faster translation and thus faster movement. And, indeed, when they measured the rate of translation on a single-molecule level, they found that translation occurs about twice as fast for beta-actin as for gamma-actin.
“We expected that faster translation would mean faster movement,” Kashina says, “and that’s not what we found. It took us a long time to explain why.”
What they eventually discovered, was that although the subunits of beta-actin could be supplied faster than those of gamma-actin, that speed worked to the detriment of cell migration speed.
“We found that the faster you supply it, the better the cell attaches to the substrate,” Kashina says. “It creates proper traction, which is essential for normal migration. And if you don’t supply it fast enough the cell can’t attach properly and starts sliding. So that explained our seemingly counterintuitive results.”
Kashina and colleagues plan to continue to probe the role of the nucleotide sequence, including why evolutionary forces led to the production of such similar forms of actin and whether the “silent code” is at work in other proteins.
“We think this is part of a bigger story,” Kashina says. “We believe actins are not the only proteins that behave this way. There are a number of protein families in the human genome that contain highly similar proteins encoded by different genes. This silent code could be at play in those families as well.”
Reference: “Different translation dynamics of β-and γ-actin regulates cell migration” by Pavan Vedula, Satoshi Kurosaka, Brittany MacTaggart, Qin Ni, Garegin Papoian, Yi Jiang, Dawei Dong and Anna Kashina, 24 June 2021, eLife.
DOI: 10.7554/eLife.68712
Anna Kashina is a professor of biochemistry at the University of Pennsylvania School of Veterinary Medicine.
Kashina’s coauthors on the study were Penn Vet’s Pavan Vedula, Brittany MacTaggart, and Dawei Dong; Satoshi Kurosaka of Japan’s Kindai University’s; the University of Maryland’s Qin Ni and Garegin A. Papoian; and Georgia State University’s Yi Jiang.
The study was supported by the National Institutes of Health (grants GM122505, CA201340, and EY028450) and National Science Foundation (grants 1800418 and 1806903).
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.